Advancing sleep detection by modelling weak label sets: A novel weakly supervised learning approach
Feb 27, 2024Matthias Boeker, Vajira Thambawita, Michael Riegler, Pål Halvorsen, Hugo L. Hammer
Understanding sleep and activity patterns plays a crucial role in physical and mental health. This study introduces a novel approach for sleep detection using weakly supervised learning for scenarios where reliable ground truth labels are unavailable. The proposed method relies on a set of weak labels, derived from the predictions generated by conventional sleep detection algorithms. Introducing a novel approach, we suggest a novel generalised non-linear statistical model in which the number of weak sleep labels is modelled as outcome of a binomial distribution. The probability of sleep in the binomial distribution is linked to the outcomes of neural networks trained to detect sleep based on actigraphy. We show that maximizing the likelihood function of the model, is equivalent to minimizing the soft cross-entropy loss. Additionally, we explored the use of the Brier score as a loss function for weak labels. The efficacy of the suggested modelling framework was demonstrated using the Multi-Ethnic Study of Atherosclerosis dataset. A \gls{lstm} trained on the soft cross-entropy outperformed conventional sleep detection algorithms, other neural network architectures and loss functions in accuracy and model calibration. This research not only advances sleep detection techniques in scenarios where ground truth data is scarce but also contributes to the broader field of weakly supervised learning by introducing innovative approach in modelling sets of weak labels.
An objective validation of polyp and instrument segmentation methods in colonoscopy through Medico 2020 polyp segmentation and MedAI 2021 transparency challenges
Jul 30, 2023Debesh Jha, Vanshali Sharma, Debapriya Banik, Debayan Bhattacharya, Kaushiki Roy, Steven A. Hicks, Nikhil Kumar Tomar, Vajira Thambawita, Adrian Krenzer, Ge-Peng Ji, Sahadev Poudel, George Batchkala, Saruar Alam, Awadelrahman M. A. Ahmed, Quoc-Huy Trinh, Zeshan Khan, Tien-Phat Nguyen, Shruti Shrestha, Sabari Nathan, Jeonghwan Gwak, Ritika K. Jha, Zheyuan Zhang, Alexander Schlaefer, Debotosh Bhattacharjee, M. K. Bhuyan, Pradip K. Das, Sravanthi Parsa, Sharib Ali, Michael A. Riegler, Pål Halvorsen, Ulas Bagci, Thomas De Lange
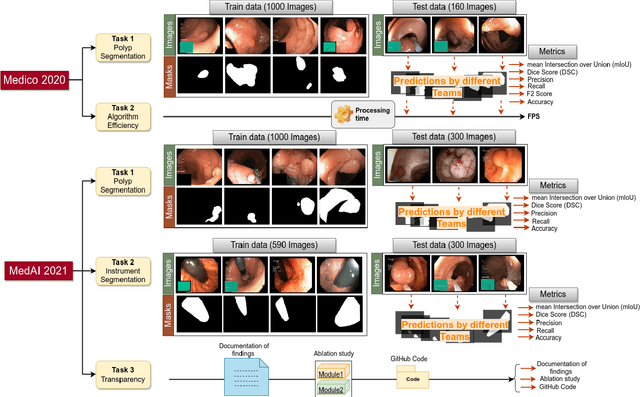
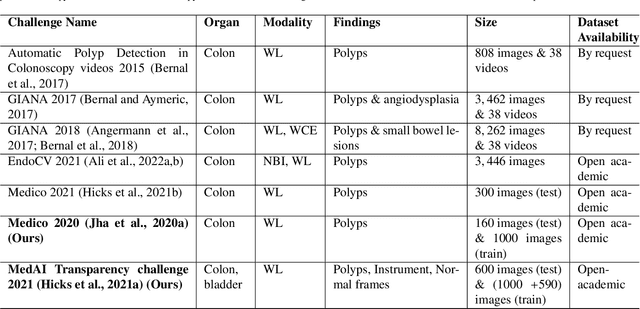
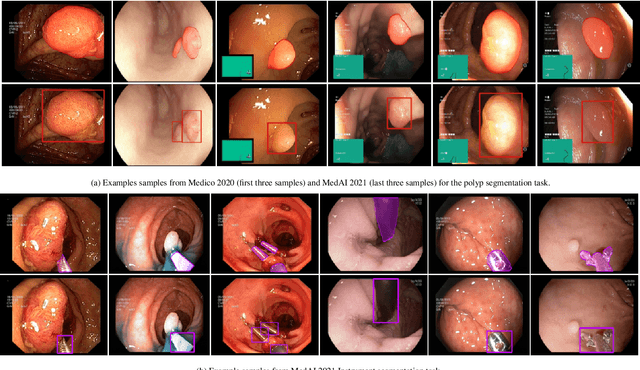
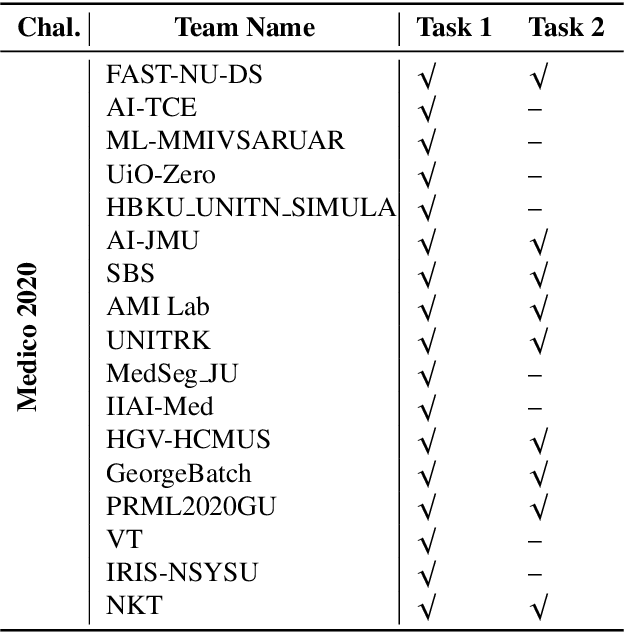
Automatic analysis of colonoscopy images has been an active field of research motivated by the importance of early detection of precancerous polyps. However, detecting polyps during the live examination can be challenging due to various factors such as variation of skills and experience among the endoscopists, lack of attentiveness, and fatigue leading to a high polyp miss-rate. Deep learning has emerged as a promising solution to this challenge as it can assist endoscopists in detecting and classifying overlooked polyps and abnormalities in real time. In addition to the algorithm's accuracy, transparency and interpretability are crucial to explaining the whys and hows of the algorithm's prediction. Further, most algorithms are developed in private data, closed source, or proprietary software, and methods lack reproducibility. Therefore, to promote the development of efficient and transparent methods, we have organized the "Medico automatic polyp segmentation (Medico 2020)" and "MedAI: Transparency in Medical Image Segmentation (MedAI 2021)" competitions. We present a comprehensive summary and analyze each contribution, highlight the strength of the best-performing methods, and discuss the possibility of clinical translations of such methods into the clinic. For the transparency task, a multi-disciplinary team, including expert gastroenterologists, accessed each submission and evaluated the team based on open-source practices, failure case analysis, ablation studies, usability and understandability of evaluations to gain a deeper understanding of the models' credibility for clinical deployment. Through the comprehensive analysis of the challenge, we not only highlight the advancements in polyp and surgical instrument segmentation but also encourage qualitative evaluation for building more transparent and understandable AI-based colonoscopy systems.
Mask-conditioned latent diffusion for generating gastrointestinal polyp images
Apr 11, 2023Roman Macháček, Leila Mozaffari, Zahra Sepasdar, Sravanthi Parasa, Pål Halvorsen, Michael A. Riegler, Vajira Thambawita
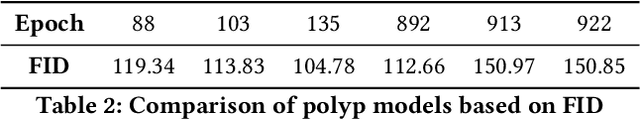
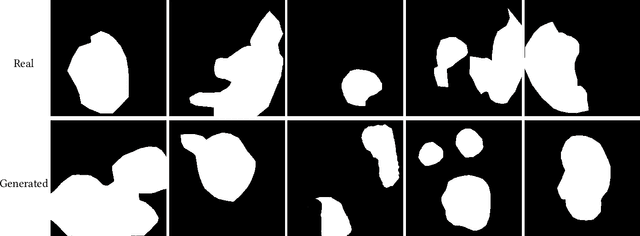
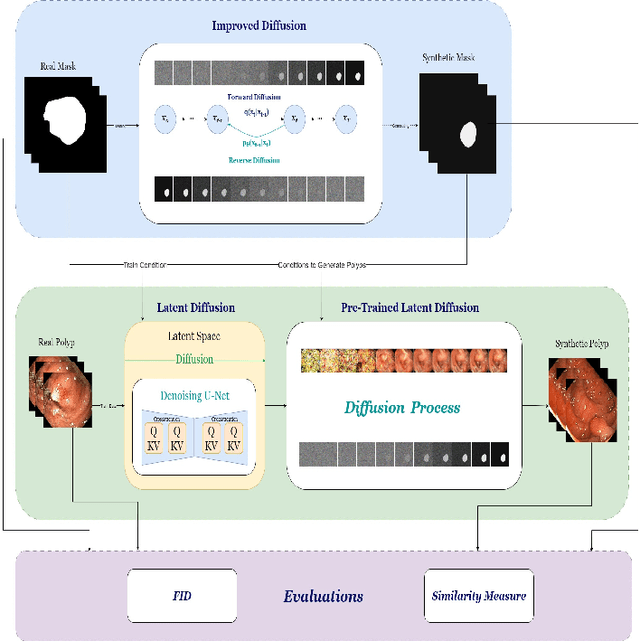
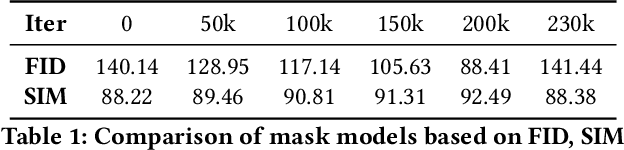


In order to take advantage of AI solutions in endoscopy diagnostics, we must overcome the issue of limited annotations. These limitations are caused by the high privacy concerns in the medical field and the requirement of getting aid from experts for the time-consuming and costly medical data annotation process. In computer vision, image synthesis has made a significant contribution in recent years as a result of the progress of generative adversarial networks (GANs) and diffusion probabilistic models (DPM). Novel DPMs have outperformed GANs in text, image, and video generation tasks. Therefore, this study proposes a conditional DPM framework to generate synthetic GI polyp images conditioned on given generated segmentation masks. Our experimental results show that our system can generate an unlimited number of high-fidelity synthetic polyp images with the corresponding ground truth masks of polyps. To test the usefulness of the generated data, we trained binary image segmentation models to study the effect of using synthetic data. Results show that the best micro-imagewise IOU of 0.7751 was achieved from DeepLabv3+ when the training data consists of both real data and synthetic data. However, the results reflect that achieving good segmentation performance with synthetic data heavily depends on model architectures.
VISEM-Tracking: Human Spermatozoa Tracking Dataset
Dec 23, 2022Vajira Thambawita, Steven A. Hicks, Andrea M. Storås, Thu Nguyen, Jorunn M. Andersen, Oliwia Witczak, Trine B. Haugen, Hugo L. Hammer, Pål Halvorsen, Michael A. Riegler
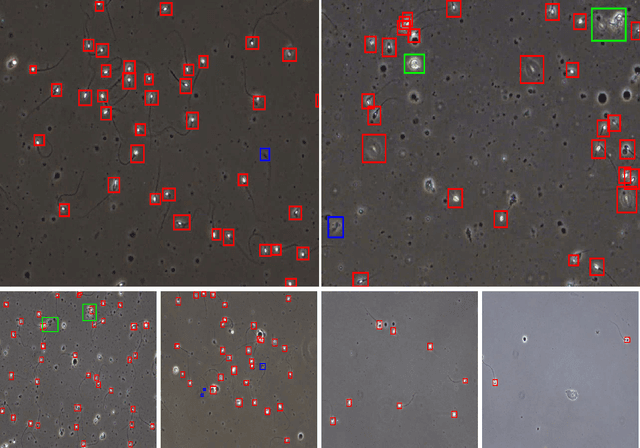
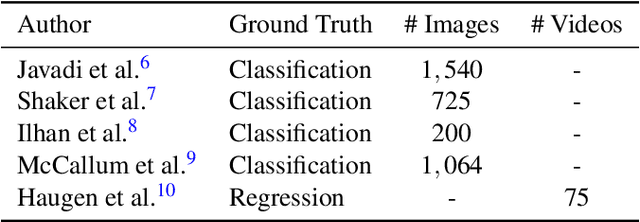

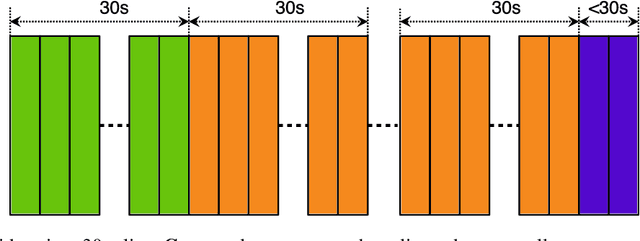
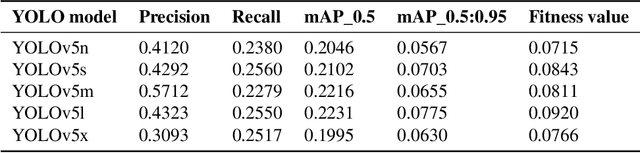
A manual assessment of sperm motility requires microscopy observation, which is challenging due to the fast-moving spermatozoa in the field of view. To obtain correct results, manual evaluation requires extensive training. Therefore, computer-assisted sperm analysis (CASA) has become increasingly used in clinics. Despite this, more data is needed to train supervised machine learning approaches in order to improve accuracy and reliability in the assessment of sperm motility and kinematics. In this regard, we provide a dataset called VISEM-Tracking with 20 video recordings of 30 seconds of wet sperm preparations with manually annotated bounding-box coordinates and a set of sperm characteristics analyzed by experts in the domain. In addition to the annotated data, we provide unlabeled video clips for easy-to-use access and analysis of the data via methods such as self- or unsupervised learning. As part of this paper, we present baseline sperm detection performances using the YOLOv5 deep learning model trained on the VISEM-Tracking dataset. As a result, we show that the dataset can be used to train complex deep learning models to analyze spermatozoa. The dataset is publicly available at https://zenodo.org/record/7293726.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Patrick Godau, Veronika Cheplygina, Michal Kozubek, Sharib Ali, Anubha Gupta, Jan Kybic, Alison Noble, Carlos Ortiz de Solórzano, Samiksha Pachade, Caroline Petitjean, Daniel Sage, Donglai Wei, Elizabeth Wilden, Deepak Alapatt, Vincent Andrearczyk, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Vivek Singh Bawa, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Jinwook Choi, Olivier Commowick, Marie Daum, Adrien Depeursinge, Reuben Dorent, Jan Egger, Hannah Eichhorn, Sandy Engelhardt, Melanie Ganz, Gabriel Girard, Lasse Hansen, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Hyunjeong Kim, Bennett Landman, Hongwei Bran Li, Jianning Li, Jun Ma, Anne Martel, Carlos Martín-Isla, Bjoern Menze, Chinedu Innocent Nwoye, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Carole Sudre, Kimberlin van Wijnen, Armine Vardazaryan, Tom Vercauteren, Martin Wagner, Chuanbo Wang, Moi Hoon Yap, Zeyun Yu, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Rina Bao, Chanyeol Choi, Andrew Cohen, Oleh Dzyubachyk, Adrian Galdran, Tianyuan Gan, Tianqi Guo, Pradyumna Gupta, Mahmood Haithami, Edward Ho, Ikbeom Jang, Zhili Li, Zhengbo Luo, Filip Lux, Sokratis Makrogiannis, Dominik Müller, Young-tack Oh, Subeen Pang, Constantin Pape, Gorkem Polat, Charlotte Rosalie Reed, Kanghyun Ryu, Tim Scherr, Vajira Thambawita, Haoyu Wang, Xinliang Wang, Kele Xu, Hung Yeh, Doyeob Yeo, Yixuan Yuan, Yan Zeng, Xin Zhao, Julian Abbing, Jannes Adam, Nagesh Adluru, Niklas Agethen, Salman Ahmed, Yasmina Al Khalil, Mireia Alenyà, Esa Alhoniemi, Chengyang An, Talha Anwar, Tewodros Weldebirhan Arega, Netanell Avisdris, Dogu Baran Aydogan, Yingbin Bai, Maria Baldeon Calisto, Berke Doga Basaran, Marcel Beetz, Cheng Bian, Hao Bian, Kevin Blansit, Louise Bloch, Robert Bohnsack, Sara Bosticardo, Jack Breen, Mikael Brudfors, Raphael Brüngel, Mariano Cabezas, Alberto Cacciola, Zhiwei Chen, Yucong Chen, Daniel Tianming Chen, Minjeong Cho, Min-Kook Choi, Chuantao Xie Chuantao Xie, Dana Cobzas, Julien Cohen-Adad, Jorge Corral Acero, Sujit Kumar Das, Marcela de Oliveira, Hanqiu Deng, Guiming Dong, Lars Doorenbos, Cory Efird, Di Fan, Mehdi Fatan Serj, Alexandre Fenneteau, Lucas Fidon, Patryk Filipiak, René Finzel, Nuno R. Freitas, Christoph M. Friedrich, Mitchell Fulton, Finn Gaida, Francesco Galati, Christoforos Galazis, Chang Hee Gan, Zheyao Gao, Shengbo Gao, Matej Gazda, Beerend Gerats, Neil Getty, Adam Gibicar, Ryan Gifford, Sajan Gohil, Maria Grammatikopoulou, Daniel Grzech, Orhun Güley, Timo Günnemann, Chunxu Guo, Sylvain Guy, Heonjin Ha, Luyi Han, Il Song Han, Ali Hatamizadeh, Tian He, Jimin Heo, Sebastian Hitziger, SeulGi Hong, SeungBum Hong, Rian Huang, Ziyan Huang, Markus Huellebrand, Stephan Huschauer, Mustaffa Hussain, Tomoo Inubushi, Ece Isik Polat, Mojtaba Jafaritadi, SeongHun Jeong, Bailiang Jian, Yuanhong Jiang, Zhifan Jiang, Yueming Jin, Smriti Joshi, Abdolrahim Kadkhodamohammadi, Reda Abdellah Kamraoui, Inha Kang, Junghwa Kang, Davood Karimi, April Khademi, Muhammad Irfan Khan, Suleiman A. Khan, Rishab Khantwal, Kwang-Ju Kim, Timothy Kline, Satoshi Kondo, Elina Kontio, Adrian Krenzer, Artem Kroviakov, Hugo Kuijf, Satyadwyoom Kumar, Francesco La Rosa, Abhi Lad, Doohee Lee, Minho Lee, Chiara Lena, Hao Li, Ling Li, Xingyu Li, Fuyuan Liao, KuanLun Liao, Arlindo Limede Oliveira, Chaonan Lin, Shan Lin, Akis Linardos, Marius George Linguraru, Han Liu, Tao Liu, Di Liu, Yanling Liu, João Lourenço-Silva, Jingpei Lu, Jiangshan Lu, Imanol Luengo, Christina B. Lund, Huan Minh Luu, Yi Lv, Yi Lv, Uzay Macar, Leon Maechler, Sina Mansour L., Kenji Marshall, Moona Mazher, Richard McKinley, Alfonso Medela, Felix Meissen, Mingyuan Meng, Dylan Miller, Seyed Hossein Mirjahanmardi, Arnab Mishra, Samir Mitha, Hassan Mohy-ud-Din, Tony Chi Wing Mok, Gowtham Krishnan Murugesan, Enamundram Naga Karthik, Sahil Nalawade, Jakub Nalepa, Mohamed Naser, Ramin Nateghi, Hammad Naveed, Quang-Minh Nguyen, Cuong Nguyen Quoc, Brennan Nichyporuk, Bruno Oliveira, David Owen, Jimut Bahan Pal, Junwen Pan, Wentao Pan, Winnie Pang, Bogyu Park, Vivek Pawar, Kamlesh Pawar, Michael Peven, Lena Philipp, Tomasz Pieciak, Szymon Plotka, Marcel Plutat, Fattaneh Pourakpour, Domen Preložnik, Kumaradevan Punithakumar, Abdul Qayyum, Sandro Queirós, Arman Rahmim, Salar Razavi, Jintao Ren, Mina Rezaei, Jonathan Adam Rico, ZunHyan Rieu, Markus Rink, Johannes Roth, Yusely Ruiz-Gonzalez, Numan Saeed, Anindo Saha, Mostafa Salem, Ricardo Sanchez-Matilla, Kurt Schilling, Wei Shao, Zhiqiang Shen, Ruize Shi, Pengcheng Shi, Daniel Sobotka, Théodore Soulier, Bella Specktor Fadida, Danail Stoyanov, Timothy Sum Hon Mun, Xiaowu Sun, Rong Tao, Franz Thaler, Antoine Théberge, Felix Thielke, Helena Torres, Kareem A. Wahid, Jiacheng Wang, YiFei Wang, Wei Wang, Xiong Wang, Jianhui Wen, Ning Wen, Marek Wodzinski, Ye Wu, Fangfang Xia, Tianqi Xiang, Chen Xiaofei, Lizhan Xu, Tingting Xue, Yuxuan Yang, Lin Yang, Kai Yao, Huifeng Yao, Amirsaeed Yazdani, Michael Yip, Hwanseung Yoo, Fereshteh Yousefirizi, Shunkai Yu, Lei Yu, Jonathan Zamora, Ramy Ashraf Zeineldin, Dewen Zeng, Jianpeng Zhang, Bokai Zhang, Jiapeng Zhang, Fan Zhang, Huahong Zhang, Zhongchen Zhao, Zixuan Zhao, Jiachen Zhao, Can Zhao, Qingshuo Zheng, Yuheng Zhi, Ziqi Zhou, Baosheng Zou, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
MLC at HECKTOR 2022: The Effect and Importance of Training Data when Analyzing Cases of Head and Neck Tumors using Machine Learning
Nov 30, 2022Vajira Thambawita, Andrea M. Storås, Steven A. Hicks, Pål Halvorsen, Michael A. Riegler
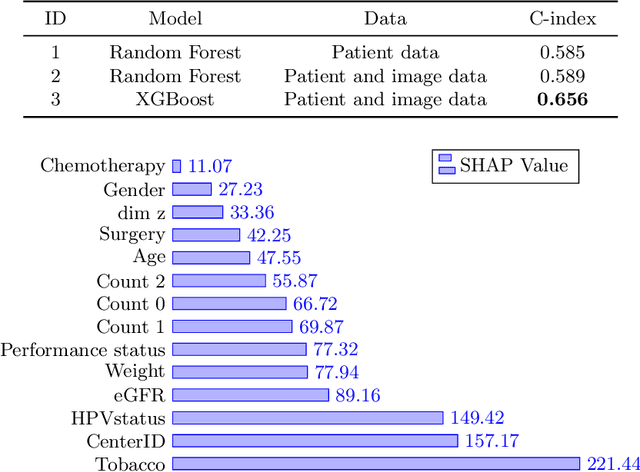
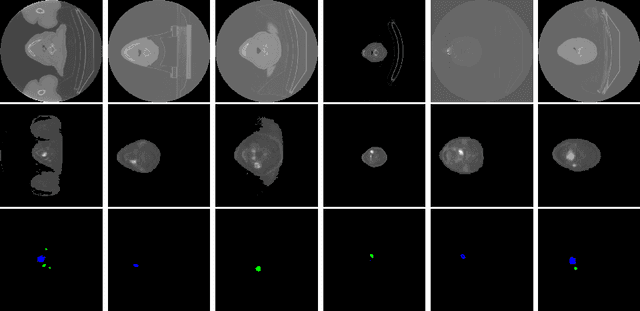
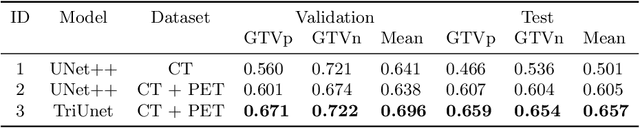
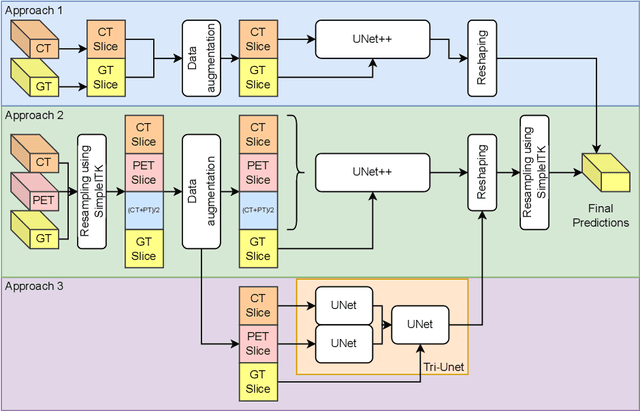

Head and neck cancers are the fifth most common cancer worldwide, and recently, analysis of Positron Emission Tomography (PET) and Computed Tomography (CT) images has been proposed to identify patients with a prognosis. Even though the results look promising, more research is needed to further validate and improve the results. This paper presents the work done by team MLC for the 2022 version of the HECKTOR grand challenge held at MICCAI 2022. For Task 1, the automatic segmentation task, our approach was, in contrast to earlier solutions using 3D segmentation, to keep it as simple as possible using a 2D model, analyzing every slice as a standalone image. In addition, we were interested in understanding how different modalities influence the results. We proposed two approaches; one using only the CT scans to make predictions and another using a combination of the CT and PET scans. For Task 2, the prediction of recurrence-free survival, we first proposed two approaches, one where we only use patient data and one where we combined the patient data with segmentations from the image model. For the prediction of the first two approaches, we used Random Forest. In our third approach, we combined patient data and image data using XGBoost. Low kidney function might worsen cancer prognosis. In this approach, we therefore estimated the kidney function of the patients and included it as a feature. Overall, we conclude that our simple methods were not able to compete with the highest-ranking submissions, but we still obtained reasonably good scores. We also got interesting insights into how the combination of different modalities can influence the segmentation and predictions.
Segmentation Consistency Training: Out-of-Distribution Generalization for Medical Image Segmentation
May 30, 2022Birk Torpmann-Hagen, Vajira Thambawita, Kyrre Glette, Pål Halvorsen, Michael A. Riegler
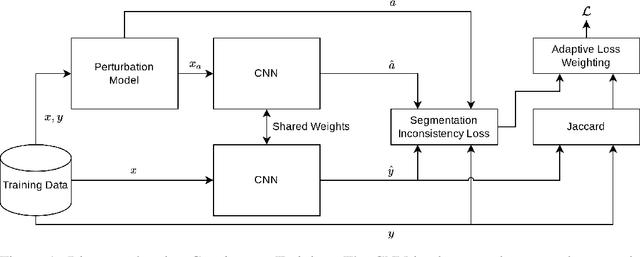
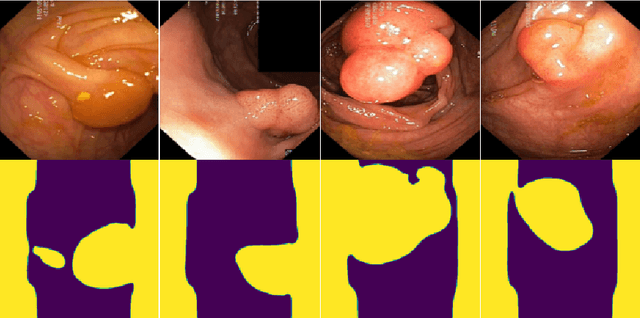

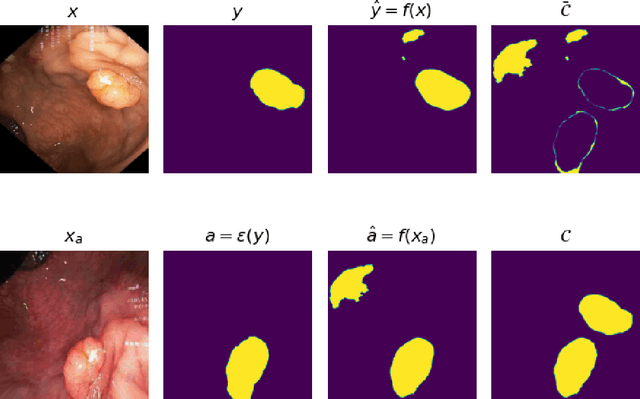
Generalizability is seen as one of the major challenges in deep learning, in particular in the domain of medical imaging, where a change of hospital or in imaging routines can lead to a complete failure of a model. To tackle this, we introduce Consistency Training, a training procedure and alternative to data augmentation based on maximizing models' prediction consistency across augmented and unaugmented data in order to facilitate better out-of-distribution generalization. To this end, we develop a novel region-based segmentation loss function called Segmentation Inconsistency Loss (SIL), which considers the differences between pairs of augmented and unaugmented predictions and labels. We demonstrate that Consistency Training outperforms conventional data augmentation on several out-of-distribution datasets on polyp segmentation, a popular medical task.
PolypConnect: Image inpainting for generating realistic gastrointestinal tract images with polyps
May 30, 2022Jan Andre Fagereng, Vajira Thambawita, Andrea M. Storås, Sravanthi Parasa, Thomas de Lange, Pål Halvorsen, Michael A. Riegler
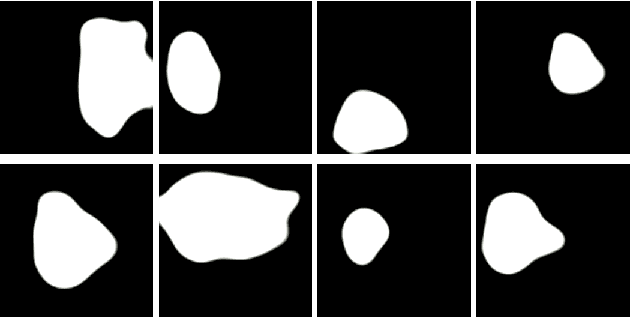
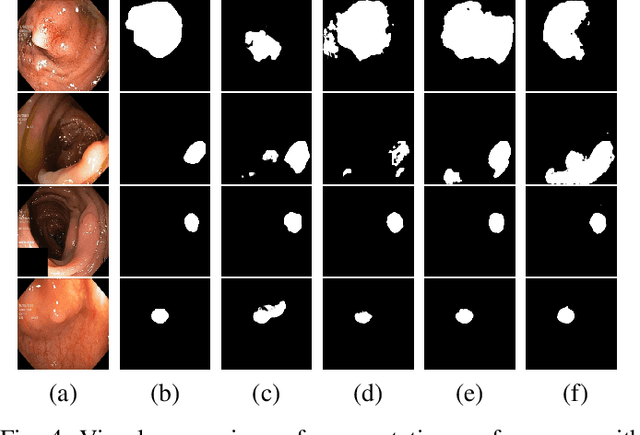
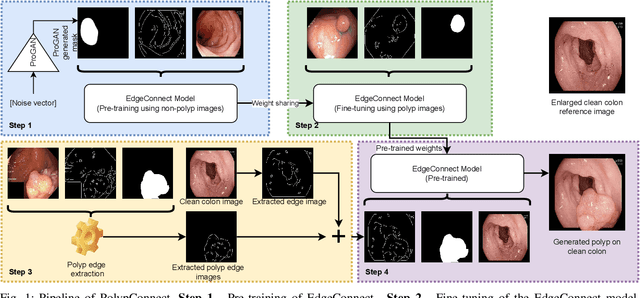



Early identification of a polyp in the lower gastrointestinal (GI) tract can lead to prevention of life-threatening colorectal cancer. Developing computer-aided diagnosis (CAD) systems to detect polyps can improve detection accuracy and efficiency and save the time of the domain experts called endoscopists. Lack of annotated data is a common challenge when building CAD systems. Generating synthetic medical data is an active research area to overcome the problem of having relatively few true positive cases in the medical domain. To be able to efficiently train machine learning (ML) models, which are the core of CAD systems, a considerable amount of data should be used. In this respect, we propose the PolypConnect pipeline, which can convert non-polyp images into polyp images to increase the size of training datasets for training. We present the whole pipeline with quantitative and qualitative evaluations involving endoscopists. The polyp segmentation model trained using synthetic data, and real data shows a 5.1% improvement of mean intersection over union (mIOU), compared to the model trained only using real data. The codes of all the experiments are available on GitHub to reproduce the results.
Grid HTM: Hierarchical Temporal Memory for Anomaly Detection in Videos
May 30, 2022Vladimir Monakhov, Vajira Thambawita, Pål Halvorsen, Michael A. Riegler
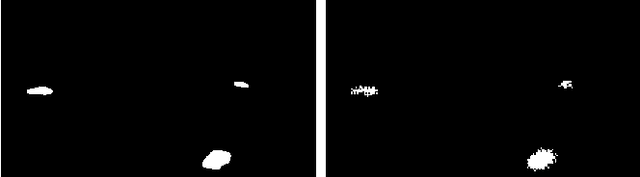


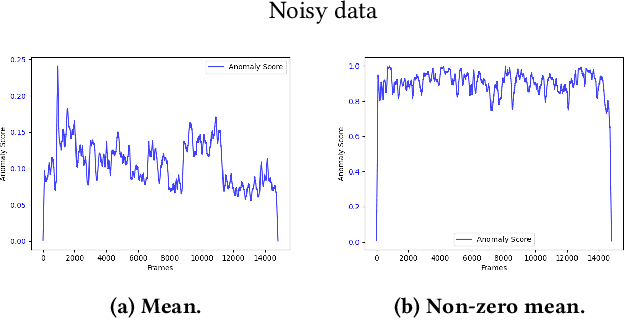
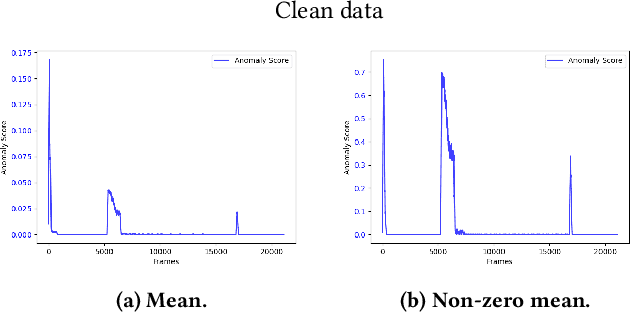
The interest for video anomaly detection systems has gained traction for the past few years. The current approaches use deep learning to perform anomaly detection in videos, but this approach has multiple problems. For starters, deep learning in general has issues with noise, concept drift, explainability, and training data volumes. Additionally, anomaly detection in itself is a complex task and faces challenges such as unknowness, heterogeneity, and class imbalance. Anomaly detection using deep learning is therefore mainly constrained to generative models such as generative adversarial networks and autoencoders due to their unsupervised nature, but even they suffer from general deep learning issues and are hard to train properly. In this paper, we explore the capabilities of the Hierarchical Temporal Memory (HTM) algorithm to perform anomaly detection in videos, as it has favorable properties such as noise tolerance and online learning which combats concept drift. We introduce a novel version of HTM, namely, Grid HTM, which is an HTM-based architecture specifically for anomaly detection in complex videos such as surveillance footage.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge