Maximilian Zenk
Inclusive, Differentially Private Federated Learning for Clinical Data
May 28, 2025Abstract:Federated Learning (FL) offers a promising approach for training clinical AI models without centralizing sensitive patient data. However, its real-world adoption is hindered by challenges related to privacy, resource constraints, and compliance. Existing Differential Privacy (DP) approaches often apply uniform noise, which disproportionately degrades model performance, even among well-compliant institutions. In this work, we propose a novel compliance-aware FL framework that enhances DP by adaptively adjusting noise based on quantifiable client compliance scores. Additionally, we introduce a compliance scoring tool based on key healthcare and security standards to promote secure, inclusive, and equitable participation across diverse clinical settings. Extensive experiments on public datasets demonstrate that integrating under-resourced, less compliant clinics with highly regulated institutions yields accuracy improvements of up to 15% over traditional FL. This work advances FL by balancing privacy, compliance, and performance, making it a viable solution for real-world clinical workflows in global healthcare.
Advances in Automated Fetal Brain MRI Segmentation and Biometry: Insights from the FeTA 2024 Challenge
May 05, 2025


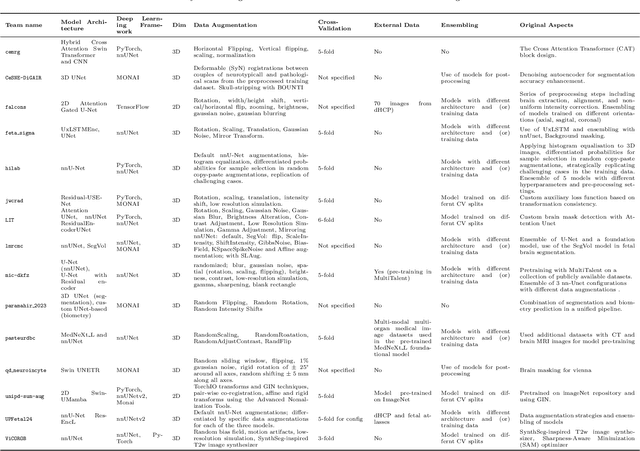
Abstract:Accurate fetal brain tissue segmentation and biometric analysis are essential for studying brain development in utero. The FeTA Challenge 2024 advanced automated fetal brain MRI analysis by introducing biometry prediction as a new task alongside tissue segmentation. For the first time, our diverse multi-centric test set included data from a new low-field (0.55T) MRI dataset. Evaluation metrics were also expanded to include the topology-specific Euler characteristic difference (ED). Sixteen teams submitted segmentation methods, most of which performed consistently across both high- and low-field scans. However, longitudinal trends indicate that segmentation accuracy may be reaching a plateau, with results now approaching inter-rater variability. The ED metric uncovered topological differences that were missed by conventional metrics, while the low-field dataset achieved the highest segmentation scores, highlighting the potential of affordable imaging systems when paired with high-quality reconstruction. Seven teams participated in the biometry task, but most methods failed to outperform a simple baseline that predicted measurements based solely on gestational age, underscoring the challenge of extracting reliable biometric estimates from image data alone. Domain shift analysis identified image quality as the most significant factor affecting model generalization, with super-resolution pipelines also playing a substantial role. Other factors, such as gestational age, pathology, and acquisition site, had smaller, though still measurable, effects. Overall, FeTA 2024 offers a comprehensive benchmark for multi-class segmentation and biometry estimation in fetal brain MRI, underscoring the need for data-centric approaches, improved topological evaluation, and greater dataset diversity to enable clinically robust and generalizable AI tools.
Benchmark of Segmentation Techniques for Pelvic Fracture in CT and X-ray: Summary of the PENGWIN 2024 Challenge
Apr 03, 2025Abstract:The segmentation of pelvic fracture fragments in CT and X-ray images is crucial for trauma diagnosis, surgical planning, and intraoperative guidance. However, accurately and efficiently delineating the bone fragments remains a significant challenge due to complex anatomy and imaging limitations. The PENGWIN challenge, organized as a MICCAI 2024 satellite event, aimed to advance automated fracture segmentation by benchmarking state-of-the-art algorithms on these complex tasks. A diverse dataset of 150 CT scans was collected from multiple clinical centers, and a large set of simulated X-ray images was generated using the DeepDRR method. Final submissions from 16 teams worldwide were evaluated under a rigorous multi-metric testing scheme. The top-performing CT algorithm achieved an average fragment-wise intersection over union (IoU) of 0.930, demonstrating satisfactory accuracy. However, in the X-ray task, the best algorithm attained an IoU of 0.774, highlighting the greater challenges posed by overlapping anatomical structures. Beyond the quantitative evaluation, the challenge revealed methodological diversity in algorithm design. Variations in instance representation, such as primary-secondary classification versus boundary-core separation, led to differing segmentation strategies. Despite promising results, the challenge also exposed inherent uncertainties in fragment definition, particularly in cases of incomplete fractures. These findings suggest that interactive segmentation approaches, integrating human decision-making with task-relevant information, may be essential for improving model reliability and clinical applicability.
Comparative Benchmarking of Failure Detection Methods in Medical Image Segmentation: Unveiling the Role of Confidence Aggregation
Jun 05, 2024



Abstract:Semantic segmentation is an essential component of medical image analysis research, with recent deep learning algorithms offering out-of-the-box applicability across diverse datasets. Despite these advancements, segmentation failures remain a significant concern for real-world clinical applications, necessitating reliable detection mechanisms. This paper introduces a comprehensive benchmarking framework aimed at evaluating failure detection methodologies within medical image segmentation. Through our analysis, we identify the strengths and limitations of current failure detection metrics, advocating for the risk-coverage analysis as a holistic evaluation approach. Utilizing a collective dataset comprising five public 3D medical image collections, we assess the efficacy of various failure detection strategies under realistic test-time distribution shifts. Our findings highlight the importance of pixel confidence aggregation and we observe superior performance of the pairwise Dice score (Roy et al., 2019) between ensemble predictions, positioning it as a simple and robust baseline for failure detection in medical image segmentation. To promote ongoing research, we make the benchmarking framework available to the community.
Real-World Federated Learning in Radiology: Hurdles to overcome and Benefits to gain
May 15, 2024



Abstract:Objective: Federated Learning (FL) enables collaborative model training while keeping data locally. Currently, most FL studies in radiology are conducted in simulated environments due to numerous hurdles impeding its translation into practice. The few existing real-world FL initiatives rarely communicate specific measures taken to overcome these hurdles, leaving behind a significant knowledge gap. Minding efforts to implement real-world FL, there is a notable lack of comprehensive assessment comparing FL to less complex alternatives. Materials & Methods: We extensively reviewed FL literature, categorizing insights along with our findings according to their nature and phase while establishing a FL initiative, summarized to a comprehensive guide. We developed our own FL infrastructure within the German Radiological Cooperative Network (RACOON) and demonstrated its functionality by training FL models on lung pathology segmentation tasks across six university hospitals. We extensively evaluated FL against less complex alternatives in three distinct evaluation scenarios. Results: The proposed guide outlines essential steps, identified hurdles, and proposed solutions for establishing successful FL initiatives conducting real-world experiments. Our experimental results show that FL outperforms less complex alternatives in all evaluation scenarios, justifying the effort required to translate FL into real-world applications. Discussion & Conclusion: Our proposed guide aims to aid future FL researchers in circumventing pitfalls and accelerating translation of FL into radiological applications. Our results underscore the value of efforts needed to translate FL into real-world applications by demonstrating advantageous performance over alternatives, and emphasize the importance of strategic organization, robust management of distributed data and infrastructure in real-world settings.
Mitigating False Predictions In Unreasonable Body Regions
Apr 24, 2024



Abstract:Despite considerable strides in developing deep learning models for 3D medical image segmentation, the challenge of effectively generalizing across diverse image distributions persists. While domain generalization is acknowledged as vital for robust application in clinical settings, the challenges stemming from training with a limited Field of View (FOV) remain unaddressed. This limitation leads to false predictions when applied to body regions beyond the FOV of the training data. In response to this problem, we propose a novel loss function that penalizes predictions in implausible body regions, applicable in both single-dataset and multi-dataset training schemes. It is realized with a Body Part Regression model that generates axial slice positional scores. Through comprehensive evaluation using a test set featuring varying FOVs, our approach demonstrates remarkable improvements in generalization capabilities. It effectively mitigates false positive tumor predictions up to 85% and significantly enhances overall segmentation performance.
Skeleton Recall Loss for Connectivity Conserving and Resource Efficient Segmentation of Thin Tubular Structures
Apr 03, 2024



Abstract:Accurately segmenting thin tubular structures, such as vessels, nerves, roads or concrete cracks, is a crucial task in computer vision. Standard deep learning-based segmentation loss functions, such as Dice or Cross-Entropy, focus on volumetric overlap, often at the expense of preserving structural connectivity or topology. This can lead to segmentation errors that adversely affect downstream tasks, including flow calculation, navigation, and structural inspection. Although current topology-focused losses mark an improvement, they introduce significant computational and memory overheads. This is particularly relevant for 3D data, rendering these losses infeasible for larger volumes as well as increasingly important multi-class segmentation problems. To mitigate this, we propose a novel Skeleton Recall Loss, which effectively addresses these challenges by circumventing intensive GPU-based calculations with inexpensive CPU operations. It demonstrates overall superior performance to current state-of-the-art approaches on five public datasets for topology-preserving segmentation, while substantially reducing computational overheads by more than 90%. In doing so, we introduce the first multi-class capable loss function for thin structure segmentation, excelling in both efficiency and efficacy for topology-preservation.
ValUES: A Framework for Systematic Validation of Uncertainty Estimation in Semantic Segmentation
Jan 16, 2024



Abstract:Uncertainty estimation is an essential and heavily-studied component for the reliable application of semantic segmentation methods. While various studies exist claiming methodological advances on the one hand, and successful application on the other hand, the field is currently hampered by a gap between theory and practice leaving fundamental questions unanswered: Can data-related and model-related uncertainty really be separated in practice? Which components of an uncertainty method are essential for real-world performance? Which uncertainty method works well for which application? In this work, we link this research gap to a lack of systematic and comprehensive evaluation of uncertainty methods. Specifically, we identify three key pitfalls in current literature and present an evaluation framework that bridges the research gap by providing 1) a controlled environment for studying data ambiguities as well as distribution shifts, 2) systematic ablations of relevant method components, and 3) test-beds for the five predominant uncertainty applications: OoD-detection, active learning, failure detection, calibration, and ambiguity modeling. Empirical results on simulated as well as real-world data demonstrate how the proposed framework is able to answer the predominant questions in the field revealing for instance that 1) separation of uncertainty types works on simulated data but does not necessarily translate to real-world data, 2) aggregation of scores is a crucial but currently neglected component of uncertainty methods, 3) While ensembles are performing most robustly across the different downstream tasks and settings, test-time augmentation often constitutes a light-weight alternative. Code is at: https://github.com/IML-DKFZ/values
Why is the winner the best?
Mar 30, 2023



Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
MultiTalent: A Multi-Dataset Approach to Medical Image Segmentation
Mar 25, 2023Abstract:The medical imaging community generates a wealth of datasets, many of which are openly accessible and annotated for specific diseases and tasks such as multi-organ or lesion segmentation. Current practices continue to limit model training and supervised pre-training to one or a few similar datasets, neglecting the synergistic potential of other available annotated data. We propose MultiTalent, a method that leverages multiple CT datasets with diverse and conflicting class definitions to train a single model for a comprehensive structure segmentation. Our results demonstrate improved segmentation performance compared to previous related approaches, systematically, also compared to single dataset training using state-of-the-art methods, especially for lesion segmentation and other challenging structures. We show that MultiTalent also represents a powerful foundation model that offers a superior pre-training for various segmentation tasks compared to commonly used supervised or unsupervised pre-training baselines. Our findings offer a new direction for the medical imaging community to effectively utilize the wealth of available data for improved segmentation performance. The code and model weights will be published here: [tba]
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge