Jinwook Choi
LoraMap: Harnessing the Power of LoRA Connections
Aug 29, 2024



Abstract:Large Language Models (LLMs) can benefit from mitigating hallucinations through fact-checking and overcoming substantial computational overhead with parameter-efficient techniques such as Low-Rank Adaptation (LoRA). While some studies have explored the parallel integration of multiple LoRAs, these approaches need attention to the connections between them. This paper investigates methods to establish connections among multiple LoRAs. We create three reasoning datasets tailored to fact-checking and fine-tune individual LoRAs, allowing them to view and reason from diverse perspectives. Then, we explore strategies for allocating these reasoning LoRAs and introduce LoraMap, an approach to map connections between them. The results on the fact-checking task demonstrate that the performance of LoraMap is superior to LoraHub, an existing LoRA composition method. LoraMap also outperforms with significantly fewer parameters than LoraConcat, which concatenates LoRAs and further fine-tunes them.
Automated Information Extraction from Thyroid Operation Narrative: A Comparative Study of GPT-4 and Fine-tuned KoELECTRA
Jun 12, 2024



Abstract:In the rapidly evolving field of healthcare, the integration of artificial intelligence (AI) has become a pivotal component in the automation of clinical workflows, ushering in a new era of efficiency and accuracy. This study focuses on the transformative capabilities of the fine-tuned KoELECTRA model in comparison to the GPT-4 model, aiming to facilitate automated information extraction from thyroid operation narratives. The current research landscape is dominated by traditional methods heavily reliant on regular expressions, which often face challenges in processing free-style text formats containing critical details of operation records, including frozen biopsy reports. Addressing this, the study leverages advanced natural language processing (NLP) techniques to foster a paradigm shift towards more sophisticated data processing systems. Through this comparative study, we aspire to unveil a more streamlined, precise, and efficient approach to document processing in the healthcare domain, potentially revolutionizing the way medical data is handled and analyzed.
* 9 pages, 2 figures, 3 tables
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Clinical Text Generation through Leveraging Medical Concept and Relations
Oct 02, 2019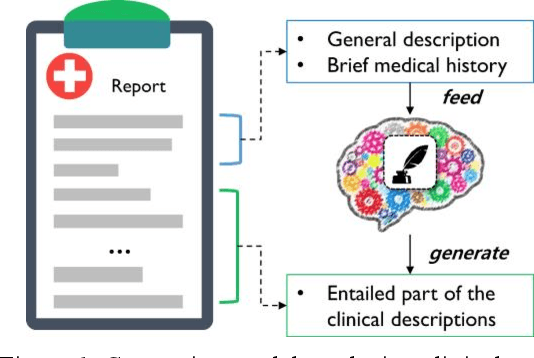
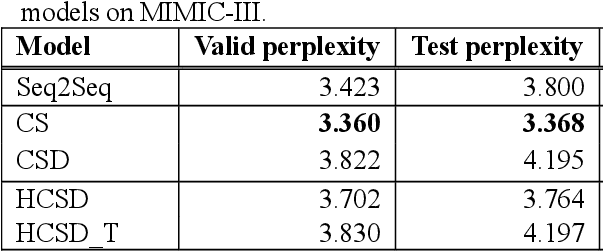
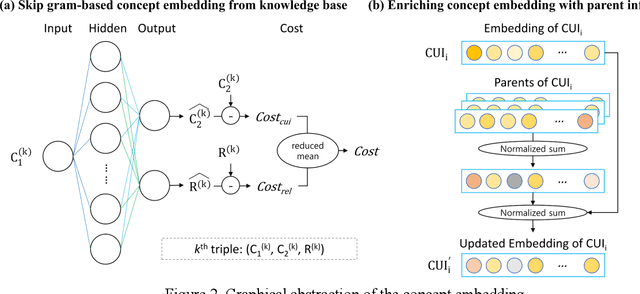
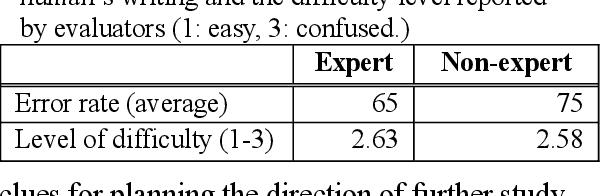
Abstract:With a neural sequence generation model, this study aims to develop a method of writing the patient clinical texts given a brief medical history. As a proof-of-a-concept, we have demonstrated that it can be workable to use medical concept embedding in clinical text generation. Our model was based on the Sequence-to-Sequence architecture and trained with a large set of de-identified clinical text data. The quantitative result shows that our concept embedding method decreased the perplexity of the baseline architecture. Also, we discuss the analyzed results from a human evaluation performed by medical doctors.
Gradient-based Camera Exposure Control for Outdoor Mobile Platforms
Jun 13, 2018
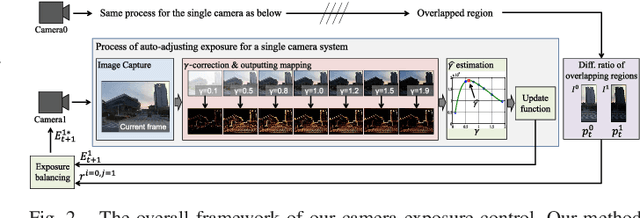
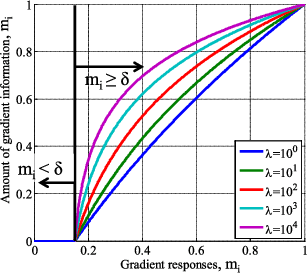
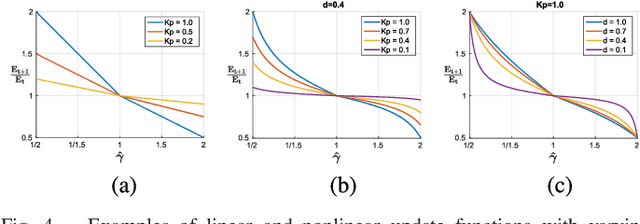
Abstract:We introduce a novel method to automatically adjust camera exposure for image processing and computer vision applications on mobile robot platforms. Because most image processing algorithms rely heavily on low-level image features that are based mainly on local gradient information, we consider that gradient quantity can determine the proper exposure level, allowing a camera to capture the important image features in a manner robust to illumination conditions. We then extend this concept to a multi-camera system and present a new control algorithm to achieve both brightness consistency between adjacent cameras and a proper exposure level for each camera. We implement our prototype system with off-the-shelf machine-vision cameras and demonstrate the effectiveness of the proposed algorithms on practical applications, including pedestrian detection, visual odometry, surround-view imaging, panoramic imaging and stereo matching.
Connecting Distant Entities with Induction through Conditional Random Fields for Named Entity Recognition: Precursor-Induced CRF
May 26, 2018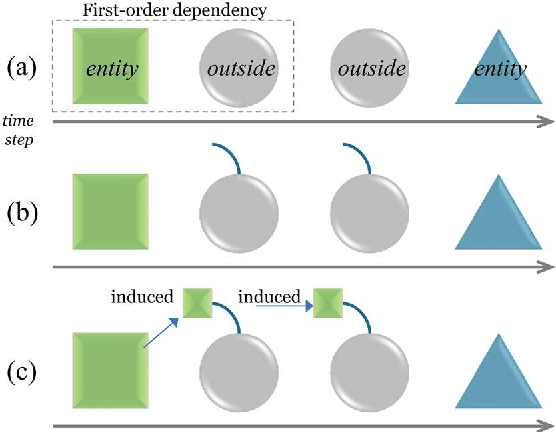
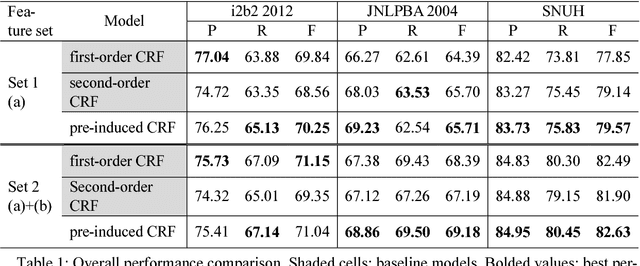
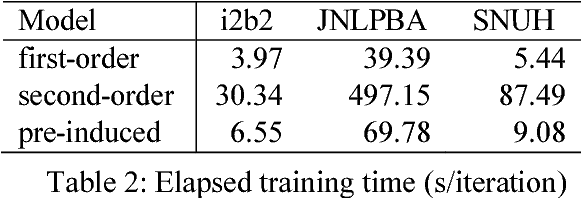
Abstract:This paper presents a method of designing specific high-order dependency factor on the linear chain conditional random fields (CRFs) for named entity recognition (NER). Named entities tend to be separated from each other by multiple outside tokens in a text, and thus the first-order CRF, as well as the second-order CRF, may innately lose transition information between distant named entities. The proposed design uses outside label in NER as a transmission medium of precedent entity information on the CRF. Then, empirical results apparently demonstrate that it is possible to exploit long-distance label dependency in the original first-order linear chain CRF structure upon NER while reducing computational loss rather than in the second-order CRF.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge