Seyed Hossein Mirjahanmardi
Domain Adaptation using Silver Standard Labels for Ki-67 Scoring in Digital Pathology: A Step Closer to Widescale Deployment
Jul 08, 2023



Abstract:Deep learning systems have been proposed to improve the objectivity and efficiency of Ki- 67 PI scoring. The challenge is that while very accurate, deep learning techniques suffer from reduced performance when applied to out-of-domain data. This is a critical challenge for clinical translation, as models are typically trained using data available to the vendor, which is not from the target domain. To address this challenge, this study proposes a domain adaptation pipeline that employs an unsupervised framework to generate silver standard (pseudo) labels in the target domain, which is used to augment the gold standard (GS) source domain data. Five training regimes were tested on two validated Ki-67 scoring architectures (UV-Net and piNET), (1) SS Only: trained on target silver standard (SS) labels, (2) GS Only: trained on source GS labels, (3) Mixed: trained on target SS and source GS labels, (4) GS+SS: trained on source GS labels and fine-tuned on target SS labels, and our proposed method (5) SS+GS: trained on source SS labels and fine-tuned on source GS labels. The SS+GS method yielded significantly (p < 0.05) higher PI accuracy (95.9%) and more consistent results compared to the GS Only model on target data. Analysis of t-SNE plots showed features learned by the SS+GS models are more aligned for source and target data, resulting in improved generalization. The proposed pipeline provides an efficient method for learning the target distribution without manual annotations, which are time-consuming and costly to generate for medical images. This framework can be applied to any target site as a per-laboratory calibration method, for widescale deployment.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Preserving Dense Features for Ki67 Nuclei Detection
Nov 10, 2021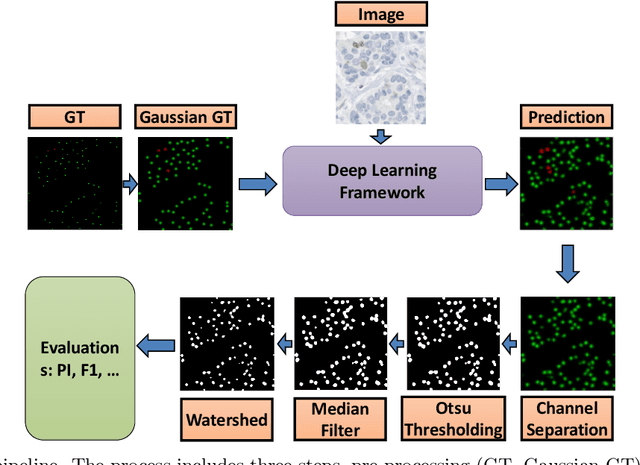
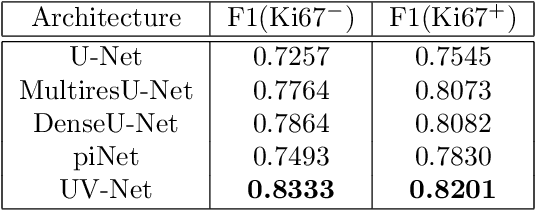
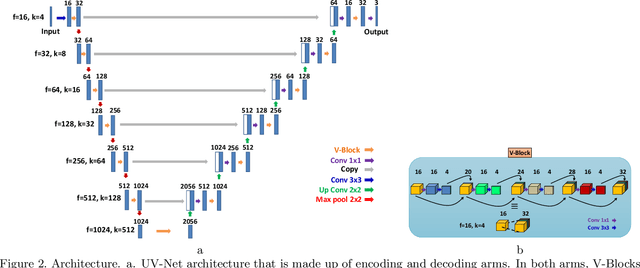
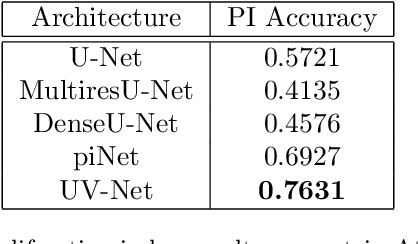
Abstract:Nuclei detection is a key task in Ki67 proliferation index estimation in breast cancer images. Deep learning algorithms have shown strong potential in nuclei detection tasks. However, they face challenges when applied to pathology images with dense medium and overlapping nuclei since fine details are often diluted or completely lost by early maxpooling layers. This paper introduces an optimized UV-Net architecture, specifically developed to recover nuclear details with high-resolution through feature preservation for Ki67 proliferation index computation. UV-Net achieves an average F1-score of 0.83 on held-out test patch data, while other architectures obtain 0.74-0.79. On tissue microarrays (unseen) test data obtained from multiple centers, UV-Net's accuracy exceeds other architectures by a wide margin, including 9-42\% on Ontario Veterinary College, 7-35\% on Protein Atlas and 0.3-3\% on University Health Network.
Low-Dispersive Permittivity Measurement Based on Transmitted Power Only
Sep 14, 2021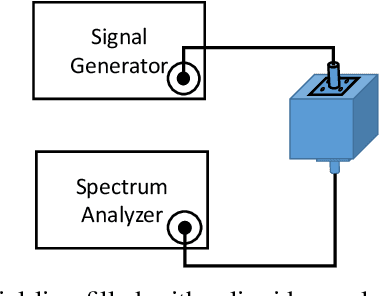
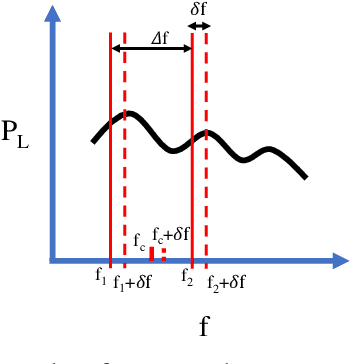
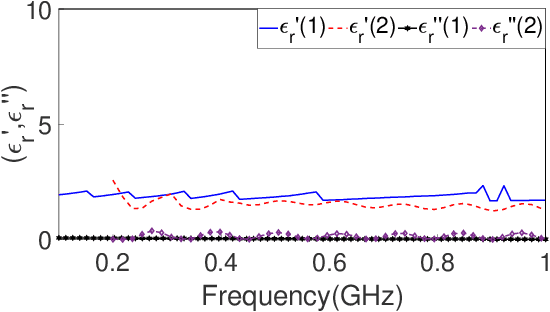
Abstract:This paper presents a complex permittivity measurement method for low-dispersive materials as a function of frequency. The introduced method relies only on transmitted power signals which are collected using a spectrum analyzer/power meter, removing the need for phase measurements and a vector network analyzer. This method provides a very good accuracy along with easy and inexpensive permittivity measurements.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge