Quoc-Huy Trinh
Firebolt-VL: Efficient Vision-Language Understanding with Cross-Modality Modulation
Apr 07, 2026Abstract:Recent advances in multimodal large language models (MLLMs) have enabled impressive progress in vision-language understanding, yet their high computational cost limits deployment in resource-constrained scenarios such as personal assistants, document understanding, and smart cameras. Most existing methods rely on Transformer-based cross-attention, whose quadratic complexity hinders efficiency. Moreover, small vision-language models often struggle to precisely capture fine-grained, task-relevant visual regions, leading to degraded performance on fine-grained reasoning tasks that limit their effectiveness in the real world. To address these issues, we introduce Firebolt-VL, an efficient vision-language model that replaces the Transformer-based decoder with a Liquid Foundation Model (LFM) decoder. To further enhance visual grounding, we propose a Token-Grid Correlation Module, which computes lightweight correlations between text tokens and image patches and modulates via the state-space model with FiLM conditioning. This enables the model to selectively emphasize visual regions relevant to the textual prompt while maintaining linear-time inference. Experimental results across multiple benchmarks demonstrate that Firebolt-VL achieves accurate, fine-grained understanding with significantly improved efficiency. Our model and code are available at: https://fireboltvl.github.io
Beyond Medical Diagnostics: How Medical Multimodal Large Language Models Think in Space
Mar 14, 2026Abstract:Visual spatial intelligence is critical for medical image interpretation, yet remains largely unexplored in Multimodal Large Language Models (MLLMs) for 3D imaging. This gap persists due to a systemic lack of datasets featuring structured 3D spatial annotations beyond basic labels. In this study, we introduce an agentic pipeline that autonomously synthesizes spatial visual question-answering (VQA) data by orchestrating computational tools such as volume and distance calculators with multi-agent collaboration and expert radiologist validation. We present SpatialMed, the first comprehensive benchmark for evaluating 3D spatial intelligence in medical MLLMs, comprising nearly 10K question-answer pairs across multiple organs and tumor types. Our evaluations on 14 state-of-the-art MLLMs and extensive analyses reveal that current models lack robust spatial reasoning capabilities for medical imaging.
Viper-F1: Fast and Fine-Grained Multimodal Understanding with Cross-Modal State-Space Modulation
Nov 18, 2025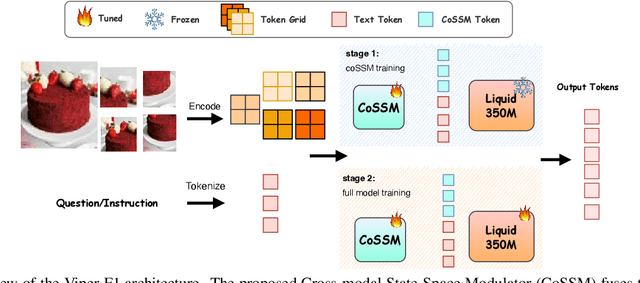
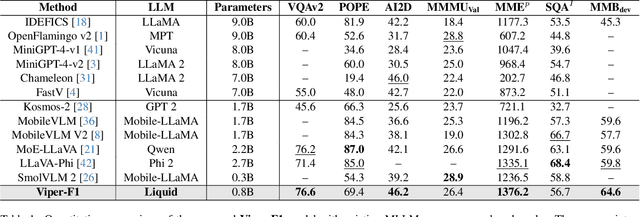
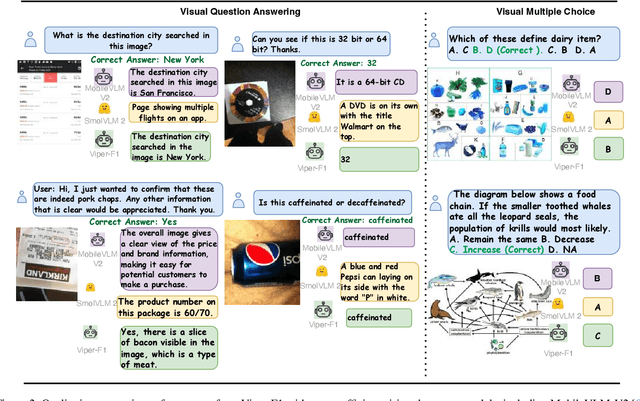
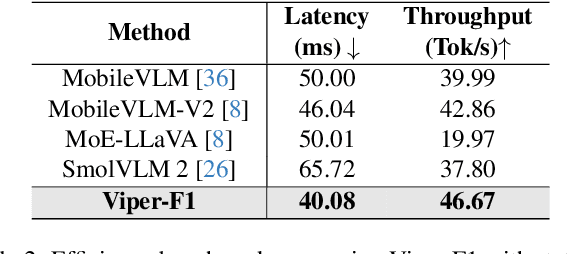
Abstract:Recent advances in multimodal large language models (MLLMs) have enabled impressive progress in vision-language understanding, yet their high computational cost limits deployment in resource-constrained scenarios such as robotic manipulation, personal assistants, and smart cameras. Most existing methods rely on Transformer-based cross-attention, whose quadratic complexity hinders efficiency. Moreover, small vision-language models often struggle to precisely capture fine-grained, task-relevant visual regions, leading to degraded performance on fine-grained reasoning tasks that limit their effectiveness in the real world. To address these issues, we introduce Viper-F1, a Hybrid State-Space Vision-Language Model that replaces attention with efficient Liquid State-Space Dynamics. To further enhance visual grounding, we propose a Token-Grid Correlation Module, which computes lightweight correlations between text tokens and image patches and modulates the state-space dynamics via FiLM conditioning. This enables the model to selectively emphasize visual regions relevant to the textual prompt while maintaining linear-time inference. Experimental results across multiple benchmarks demonstrate that Viper-F1 achieves accurate, fine-grained understanding with significantly improved efficiency.
SRMA-Mamba: Spatial Reverse Mamba Attention Network for Pathological Liver Segmentation in MRI Volumes
Aug 17, 2025



Abstract:Liver Cirrhosis plays a critical role in the prognosis of chronic liver disease. Early detection and timely intervention are critical in significantly reducing mortality rates. However, the intricate anatomical architecture and diverse pathological changes of liver tissue complicate the accurate detection and characterization of lesions in clinical settings. Existing methods underutilize the spatial anatomical details in volumetric MRI data, thereby hindering their clinical effectiveness and explainability. To address this challenge, we introduce a novel Mamba-based network, SRMA-Mamba, designed to model the spatial relationships within the complex anatomical structures of MRI volumes. By integrating the Spatial Anatomy-Based Mamba module (SABMamba), SRMA-Mamba performs selective Mamba scans within liver cirrhotic tissues and combines anatomical information from the sagittal, coronal, and axial planes to construct a global spatial context representation, enabling efficient volumetric segmentation of pathological liver structures. Furthermore, we introduce the Spatial Reverse Attention module (SRMA), designed to progressively refine cirrhotic details in the segmentation map, utilizing both the coarse segmentation map and hierarchical encoding features. Extensive experiments demonstrate that SRMA-Mamba surpasses state-of-the-art methods, delivering exceptional performance in 3D pathological liver segmentation. Our code is available for public: {\color{blue}{https://github.com/JunZengz/SRMA-Mamba}}.
PRS-Med: Position Reasoning Segmentation with Vision-Language Model in Medical Imaging
May 17, 2025



Abstract:Recent advancements in prompt-based medical image segmentation have enabled clinicians to identify tumors using simple input like bounding boxes or text prompts. However, existing methods face challenges when doctors need to interact through natural language or when position reasoning is required - understanding spatial relationships between anatomical structures and pathologies. We present PRS-Med, a framework that integrates vision-language models with segmentation capabilities to generate both accurate segmentation masks and corresponding spatial reasoning outputs. Additionally, we introduce the MMRS dataset (Multimodal Medical in Positional Reasoning Segmentation), which provides diverse, spatially-grounded question-answer pairs to address the lack of position reasoning data in medical imaging. PRS-Med demonstrates superior performance across six imaging modalities (CT, MRI, X-ray, ultrasound, endoscopy, RGB), significantly outperforming state-of-the-art methods in both segmentation accuracy and position reasoning. Our approach enables intuitive doctor-system interaction through natural language, facilitating more efficient diagnoses. Our dataset pipeline, model, and codebase will be released to foster further research in spatially-aware multimodal reasoning for medical applications.
Sing-On-Your-Beat: Simple Text-Controllable Accompaniment Generations
Nov 03, 2024



Abstract:Singing is one of the most cherished forms of human entertainment. However, creating a beautiful song requires an accompaniment that complements the vocals and aligns well with the song instruments and genre. With advancements in deep learning, previous research has focused on generating suitable accompaniments but often lacks precise alignment with the desired instrumentation and genre. To address this, we propose a straightforward method that enables control over the accompaniment through text prompts, allowing the generation of music that complements the vocals and aligns with the song instrumental and genre requirements. Through extensive experiments, we successfully generate 10-second accompaniments using vocal input and text control.
NeIn: Telling What You Don't Want
Sep 09, 2024



Abstract:Negation is a fundamental linguistic concept used by humans to convey information that they do not desire. Despite this, there has been minimal research specifically focused on negation within vision-language tasks. This lack of research means that vision-language models (VLMs) may struggle to understand negation, implying that they struggle to provide accurate results. One barrier to achieving human-level intelligence is the lack of a standard collection by which research into negation can be evaluated. This paper presents the first large-scale dataset, Negative Instruction (NeIn), for studying negation within the vision-language domain. Our dataset comprises 530,694 quadruples, i.e., source image, original caption, negative sentence, and target image in total, including 495,694 queries for training and 35,000 queries for benchmarking across multiple vision-language tasks. Specifically, we automatically generate NeIn based on a large, existing vision-language dataset, MS-COCO, via two steps: generation and filtering. During the generation phase, we leverage two VLMs, BLIP and MagicBrush, to generate the target image and a negative clause that expresses the content of the source image. In the subsequent filtering phase, we apply BLIP to remove erroneous samples. Additionally, we introduce an evaluation protocol for negation understanding of image editing models. Extensive experiments using our dataset across multiple VLMs for instruction-based image editing tasks demonstrate that even recent state-of-the-art VLMs struggle to understand negative queries. The project page is: https://tanbuinhat.github.io/NeIn/
RotCAtt-TransUNet++: Novel Deep Neural Network for Sophisticated Cardiac Segmentation
Sep 09, 2024



Abstract:Cardiovascular disease is a major global health concern, contributing significantly to global mortality. Accurately segmenting cardiac medical imaging data is crucial for reducing fatality rates associated with these conditions. However, current state-of-the-art (SOTA) neural networks, including CNN-based and Transformer-based approaches, face challenges in capturing both inter-slice connections and intra-slice details, especially in datasets featuring intricate, long-range details along the z-axis like coronary arteries. Existing methods also struggle with differentiating non-cardiac components from the myocardium, resulting in segmentation inaccuracies and the "spraying" phenomenon. To address these issues, we introduce RotCAtt-TransUNet++, a novel architecture designed for robust segmentation of intricate cardiac structures. Our approach enhances global context modeling through multiscale feature aggregation and nested skip connections in the encoder. Transformer layers facilitate capturing intra-slice interactions, while a rotatory attention mechanism handles inter-slice connectivity. A channel-wise cross-attention gate integrates multiscale information and decoder features, effectively bridging semantic gaps. Experimental results across multiple datasets demonstrate superior performance over current methods, achieving near-perfect annotation of coronary arteries and myocardium. Ablation studies confirm that our rotatory attention mechanism significantly improves segmentation accuracy by transforming embedded vectorized patches in semantic dimensional space.
* 6 pages, 5 figures
SAM-EG: Segment Anything Model with Egde Guidance framework for efficient Polyp Segmentation
Jun 21, 2024



Abstract:Polyp segmentation, a critical concern in medical imaging, has prompted numerous proposed methods aimed at enhancing the quality of segmented masks. While current state-of-the-art techniques produce impressive results, the size and computational cost of these models pose challenges for practical industry applications. Recently, the Segment Anything Model (SAM) has been proposed as a robust foundation model, showing promise for adaptation to medical image segmentation. Inspired by this concept, we propose SAM-EG, a framework that guides small segmentation models for polyp segmentation to address the computation cost challenge. Additionally, in this study, we introduce the Edge Guiding module, which integrates edge information into image features to assist the segmentation model in addressing boundary issues from current segmentation model in this task. Through extensive experiments, our small models showcase their efficacy by achieving competitive results with state-of-the-art methods, offering a promising approach to developing compact models with high accuracy for polyp segmentation and in the broader field of medical imaging.
KDAS3: Knowledge distillation via Attention Supervision, and Symmetrical structure guiding for Polyp Segmentation
Dec 13, 2023Abstract:Polyp segmentation, a contentious issue in medical imaging, has seen numerous proposed methods aimed at improving the quality of segmented masks. Currently, state-of-the-art techniques yield impressive results. However, the sheer size of these models poses challenges for practical industry applications. To address this, we present a Knowledge Distillation framework, incorporating attention supervision and the symmetrical guiding method. This framework is designed to facilitate knowledge transfer from a teacher model to a more compact student model with fewer parameters. Our experimental evaluation of the framework assesses its effectiveness in enabling the student model to acquire knowledge from the teacher efficiently. Additionally, our method serves to prevent the student model from incorporating redundant features that could lead to inaccurate predictions. Consequently, our method, boasting approximately 5 million parameters, achieves competitive results comparable to the state-of-the-art approaches. The implementation can be found at: https://github.com/huyquoctrinh/KDAS3
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge