Franz Thaler
Augmentation-based Domain Generalization and Joint Training from Multiple Source Domains for Whole Heart Segmentation
Aug 06, 2025Abstract:As the leading cause of death worldwide, cardiovascular diseases motivate the development of more sophisticated methods to analyze the heart and its substructures from medical images like Computed Tomography (CT) and Magnetic Resonance (MR). Semantic segmentations of important cardiac structures that represent the whole heart are useful to assess patient-specific cardiac morphology and pathology. Furthermore, accurate semantic segmentations can be used to generate cardiac digital twin models which allows e.g. electrophysiological simulation and personalized therapy planning. Even though deep learning-based methods for medical image segmentation achieved great advancements over the last decade, retaining good performance under domain shift -- i.e. when training and test data are sampled from different data distributions -- remains challenging. In order to perform well on domains known at training-time, we employ a (1) balanced joint training approach that utilizes CT and MR data in equal amounts from different source domains. Further, aiming to alleviate domain shift towards domains only encountered at test-time, we rely on (2) strong intensity and spatial augmentation techniques to greatly diversify the available training data. Our proposed whole heart segmentation method, a 5-fold ensemble with our contributions, achieves the best performance for MR data overall and a performance similar to the best performance for CT data when compared to a model trained solely on CT. With 93.33% DSC and 0.8388 mm ASSD for CT and 89.30% DSC and 1.2411 mm ASSD for MR data, our method demonstrates great potential to efficiently obtain accurate semantic segmentations from which patient-specific cardiac twin models can be generated.
LA-CaRe-CNN: Cascading Refinement CNN for Left Atrial Scar Segmentation
Aug 06, 2025Abstract:Atrial fibrillation (AF) represents the most prevalent type of cardiac arrhythmia for which treatment may require patients to undergo ablation therapy. In this surgery cardiac tissues are locally scarred on purpose to prevent electrical signals from causing arrhythmia. Patient-specific cardiac digital twin models show great potential for personalized ablation therapy, however, they demand accurate semantic segmentation of healthy and scarred tissue typically obtained from late gadolinium enhanced (LGE) magnetic resonance (MR) scans. In this work we propose the Left Atrial Cascading Refinement CNN (LA-CaRe-CNN), which aims to accurately segment the left atrium as well as left atrial scar tissue from LGE MR scans. LA-CaRe-CNN is a 2-stage CNN cascade that is trained end-to-end in 3D, where Stage 1 generates a prediction for the left atrium, which is then refined in Stage 2 in conjunction with the original image information to obtain a prediction for the left atrial scar tissue. To account for domain shift towards domains unknown during training, we employ strong intensity and spatial augmentation to increase the diversity of the training dataset. Our proposed method based on a 5-fold ensemble achieves great segmentation results, namely, 89.21% DSC and 1.6969 mm ASSD for the left atrium, as well as 64.59% DSC and 91.80% G-DSC for the more challenging left atrial scar tissue. Thus, segmentations obtained through LA-CaRe-CNN show great potential for the generation of patient-specific cardiac digital twin models and downstream tasks like personalized targeted ablation therapy to treat AF.
Gaussian Process Emulators for Few-Shot Segmentation in Cardiac MRI
Nov 12, 2024Abstract:Segmentation of cardiac magnetic resonance images (MRI) is crucial for the analysis and assessment of cardiac function, helping to diagnose and treat various cardiovascular diseases. Most recent techniques rely on deep learning and usually require an extensive amount of labeled data. To overcome this problem, few-shot learning has the capability of reducing data dependency on labeled data. In this work, we introduce a new method that merges few-shot learning with a U-Net architecture and Gaussian Process Emulators (GPEs), enhancing data integration from a support set for improved performance. GPEs are trained to learn the relation between the support images and the corresponding masks in latent space, facilitating the segmentation of unseen query images given only a small labeled support set at inference. We test our model with the M&Ms-2 public dataset to assess its ability to segment the heart in cardiac magnetic resonance imaging from different orientations, and compare it with state-of-the-art unsupervised and few-shot methods. Our architecture shows higher DICE coefficients compared to these methods, especially in the more challenging setups where the size of the support set is considerably small.
CaRe-CNN: Cascading Refinement CNN for Myocardial Infarct Segmentation with Microvascular Obstructions
Dec 19, 2023



Abstract:Late gadolinium enhanced (LGE) magnetic resonance (MR) imaging is widely established to assess the viability of myocardial tissue of patients after acute myocardial infarction (MI). We propose the Cascading Refinement CNN (CaRe-CNN), which is a fully 3D, end-to-end trained, 3-stage CNN cascade that exploits the hierarchical structure of such labeled cardiac data. Throughout the three stages of the cascade, the label definition changes and CaRe-CNN learns to gradually refine its intermediate predictions accordingly. Furthermore, to obtain more consistent qualitative predictions, we propose a series of post-processing steps that take anatomical constraints into account. Our CaRe-CNN was submitted to the FIMH 2023 MYOSAIQ challenge, where it ranked second out of 18 participating teams. CaRe-CNN showed great improvements most notably when segmenting the difficult but clinically most relevant myocardial infarct tissue (MIT) as well as microvascular obstructions (MVO). When computing the average scores over all labels, our method obtained the best score in eight out of ten metrics. Thus, accurate cardiac segmentation after acute MI via our CaRe-CNN allows generating patient-specific models of the heart serving as an important step towards personalized medicine.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Efficient Multi-Organ Segmentation Using SpatialConfiguration-Net with Low GPU Memory Requirements
Nov 26, 2021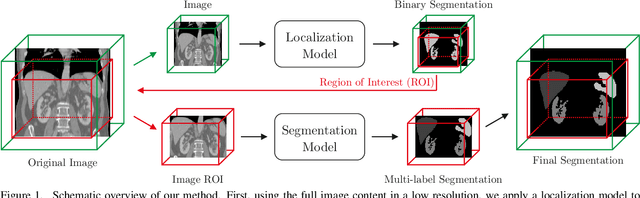

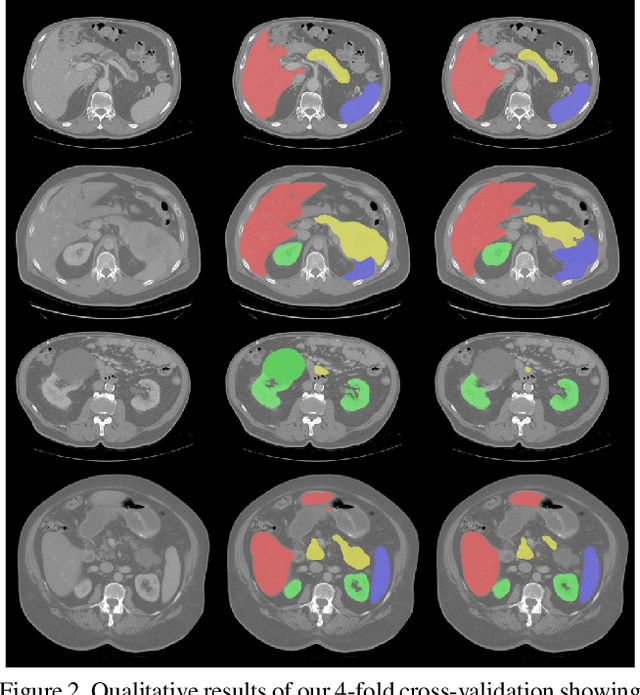
Abstract:Even though many semantic segmentation methods exist that are able to perform well on many medical datasets, often, they are not designed for direct use in clinical practice. The two main concerns are generalization to unseen data with a different visual appearance, e.g., images acquired using a different scanner, and efficiency in terms of computation time and required Graphics Processing Unit (GPU) memory. In this work, we employ a multi-organ segmentation model based on the SpatialConfiguration-Net (SCN), which integrates prior knowledge of the spatial configuration among the labelled organs to resolve spurious responses in the network outputs. Furthermore, we modified the architecture of the segmentation model to reduce its memory footprint as much as possible without drastically impacting the quality of the predictions. Lastly, we implemented a minimal inference script for which we optimized both, execution time and required GPU memory.
Modeling Annotation Uncertainty with Gaussian Heatmaps in Landmark Localization
Sep 21, 2021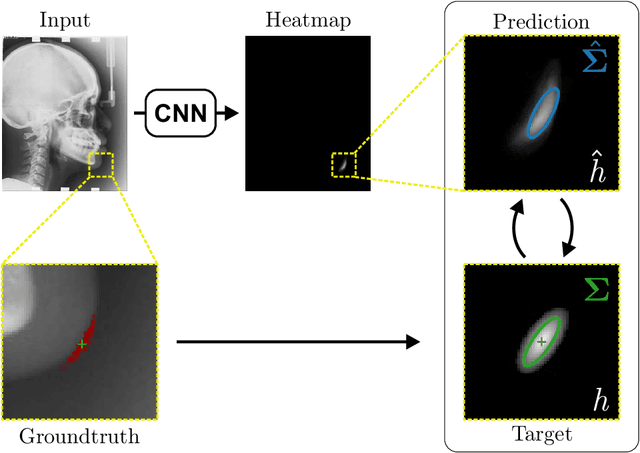
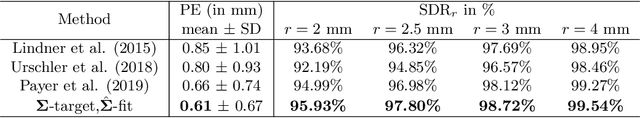
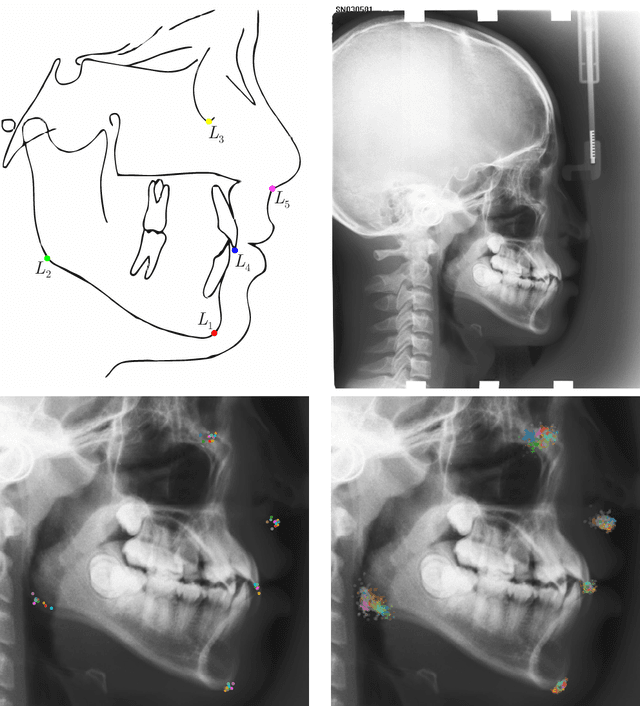
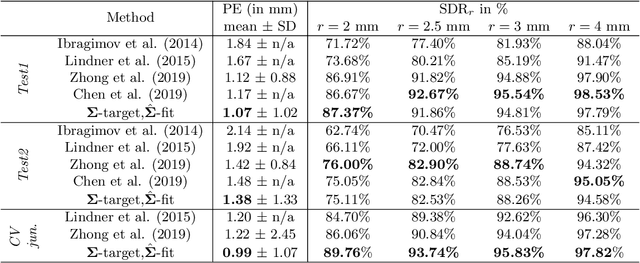
Abstract:In landmark localization, due to ambiguities in defining their exact position, landmark annotations may suffer from large observer variabilities, which result in uncertain annotations. To model the annotation ambiguities of the training dataset, we propose to learn anisotropic Gaussian parameters modeling the shape of the target heatmap during optimization. Furthermore, our method models the prediction uncertainty of individual samples by fitting anisotropic Gaussian functions to the predicted heatmaps during inference. Besides state-of-the-art results, our experiments on datasets of hand radiographs and lateral cephalograms also show that Gaussian functions are correlated with both localization accuracy and observer variability. As a final experiment, we show the importance of integrating the uncertainty into decision making by measuring the influence of the predicted location uncertainty on the classification of anatomical abnormalities in lateral cephalograms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge