Bennett Landman
Senior Member, IEEE
Scale-up Unlearnable Examples Learning with High-Performance Computing
Jan 10, 2025



Abstract:Recent advancements in AI models are structured to retain user interactions, which could inadvertently include sensitive healthcare data. In the healthcare field, particularly when radiologists use AI-driven diagnostic tools hosted on online platforms, there is a risk that medical imaging data may be repurposed for future AI training without explicit consent, spotlighting critical privacy and intellectual property concerns around healthcare data usage. Addressing these privacy challenges, a novel approach known as Unlearnable Examples (UEs) has been introduced, aiming to make data unlearnable to deep learning models. A prominent method within this area, called Unlearnable Clustering (UC), has shown improved UE performance with larger batch sizes but was previously limited by computational resources. To push the boundaries of UE performance with theoretically unlimited resources, we scaled up UC learning across various datasets using Distributed Data Parallel (DDP) training on the Summit supercomputer. Our goal was to examine UE efficacy at high-performance computing (HPC) levels to prevent unauthorized learning and enhance data security, particularly exploring the impact of batch size on UE's unlearnability. Utilizing the robust computational capabilities of the Summit, extensive experiments were conducted on diverse datasets such as Pets, MedMNist, Flowers, and Flowers102. Our findings reveal that both overly large and overly small batch sizes can lead to performance instability and affect accuracy. However, the relationship between batch size and unlearnability varied across datasets, highlighting the necessity for tailored batch size strategies to achieve optimal data protection. Our results underscore the critical role of selecting appropriate batch sizes based on the specific characteristics of each dataset to prevent learning and ensure data security in deep learning applications.
Field-of-View Extension for Diffusion MRI via Deep Generative Models
May 06, 2024



Abstract:Purpose: In diffusion MRI (dMRI), the volumetric and bundle analyses of whole-brain tissue microstructure and connectivity can be severely impeded by an incomplete field-of-view (FOV). This work aims to develop a method for imputing the missing slices directly from existing dMRI scans with an incomplete FOV. We hypothesize that the imputed image with complete FOV can improve the whole-brain tractography for corrupted data with incomplete FOV. Therefore, our approach provides a desirable alternative to discarding the valuable dMRI data, enabling subsequent tractography analyses that would otherwise be challenging or unattainable with corrupted data. Approach: We propose a framework based on a deep generative model that estimates the absent brain regions in dMRI scans with incomplete FOV. The model is capable of learning both the diffusion characteristics in diffusion-weighted images (DWI) and the anatomical features evident in the corresponding structural images for efficiently imputing missing slices of DWI outside of incomplete FOV. Results: For evaluating the imputed slices, on the WRAP dataset the proposed framework achieved PSNRb0=22.397, SSIMb0=0.905, PSNRb1300=22.479, SSIMb1300=0.893; on the NACC dataset it achieved PSNRb0=21.304, SSIMb0=0.892, PSNRb1300=21.599, SSIMb1300= 0.877. The proposed framework improved the tractography accuracy, as demonstrated by an increased average Dice score for 72 tracts (p < 0.001) on both the WRAP and NACC datasets. Conclusions: Results suggest that the proposed framework achieved sufficient imputation performance in dMRI data with incomplete FOV for improving whole-brain tractography, thereby repairing the corrupted data. Our approach achieved more accurate whole-brain tractography results with extended and complete FOV and reduced the uncertainty when analyzing bundles associated with Alzheimer's Disease.
Disruptive Autoencoders: Leveraging Low-level features for 3D Medical Image Pre-training
Jul 31, 2023Abstract:Harnessing the power of pre-training on large-scale datasets like ImageNet forms a fundamental building block for the progress of representation learning-driven solutions in computer vision. Medical images are inherently different from natural images as they are acquired in the form of many modalities (CT, MR, PET, Ultrasound etc.) and contain granulated information like tissue, lesion, organs etc. These characteristics of medical images require special attention towards learning features representative of local context. In this work, we focus on designing an effective pre-training framework for 3D radiology images. First, we propose a new masking strategy called local masking where the masking is performed across channel embeddings instead of tokens to improve the learning of local feature representations. We combine this with classical low-level perturbations like adding noise and downsampling to further enable low-level representation learning. To this end, we introduce Disruptive Autoencoders, a pre-training framework that attempts to reconstruct the original image from disruptions created by a combination of local masking and low-level perturbations. Additionally, we also devise a cross-modal contrastive loss (CMCL) to accommodate the pre-training of multiple modalities in a single framework. We curate a large-scale dataset to enable pre-training of 3D medical radiology images (MRI and CT). The proposed pre-training framework is tested across multiple downstream tasks and achieves state-of-the-art performance. Notably, our proposed method tops the public test leaderboard of BTCV multi-organ segmentation challenge.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
A Comparative Study of Confidence Calibration in Deep Learning: From Computer Vision to Medical Imaging
Jun 17, 2022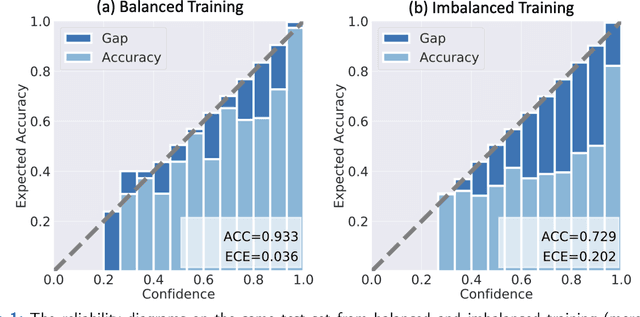
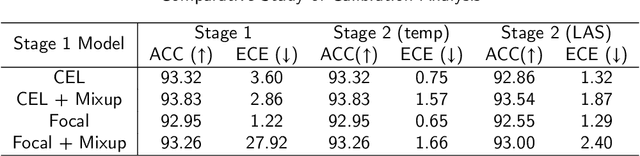
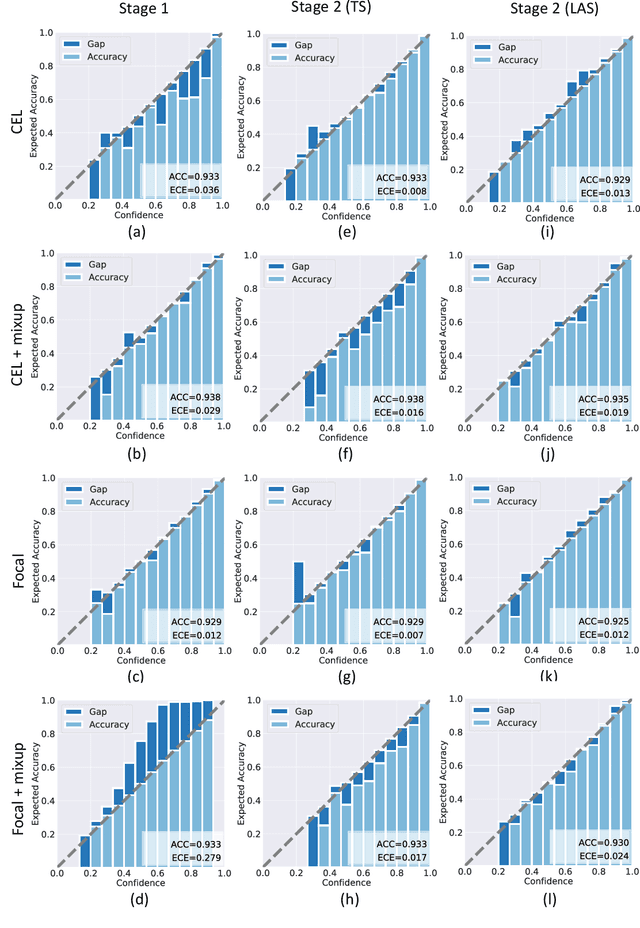
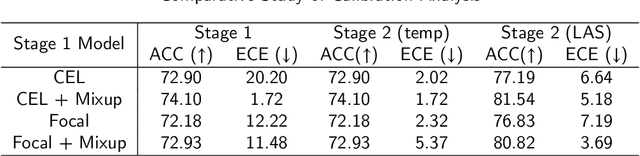
Abstract:Although deep learning prediction models have been successful in the discrimination of different classes, they can often suffer from poor calibration across challenging domains including healthcare. Moreover, the long-tail distribution poses great challenges in deep learning classification problems including clinical disease prediction. There are approaches proposed recently to calibrate deep prediction in computer vision, but there are no studies found to demonstrate how the representative models work in different challenging contexts. In this paper, we bridge the confidence calibration from computer vision to medical imaging with a comparative study of four high-impact calibration models. Our studies are conducted in different contexts (natural image classification and lung cancer risk estimation) including in balanced vs. imbalanced training sets and in computer vision vs. medical imaging. Our results support key findings: (1) We achieve new conclusions which are not studied under different learning contexts, e.g., combining two calibration models that both mitigate the overconfident prediction can lead to under-confident prediction, and simpler calibration models from the computer vision domain tend to be more generalizable to medical imaging. (2) We highlight the gap between general computer vision tasks and medical imaging prediction, e.g., calibration methods ideal for general computer vision tasks may in fact damage the calibration of medical imaging prediction. (3) We also reinforce previous conclusions in natural image classification settings. We believe that this study has merits to guide readers to choose calibration models and understand gaps between general computer vision and medical imaging domains.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
Learn2Reg: comprehensive multi-task medical image registration challenge, dataset and evaluation in the era of deep learning
Dec 23, 2021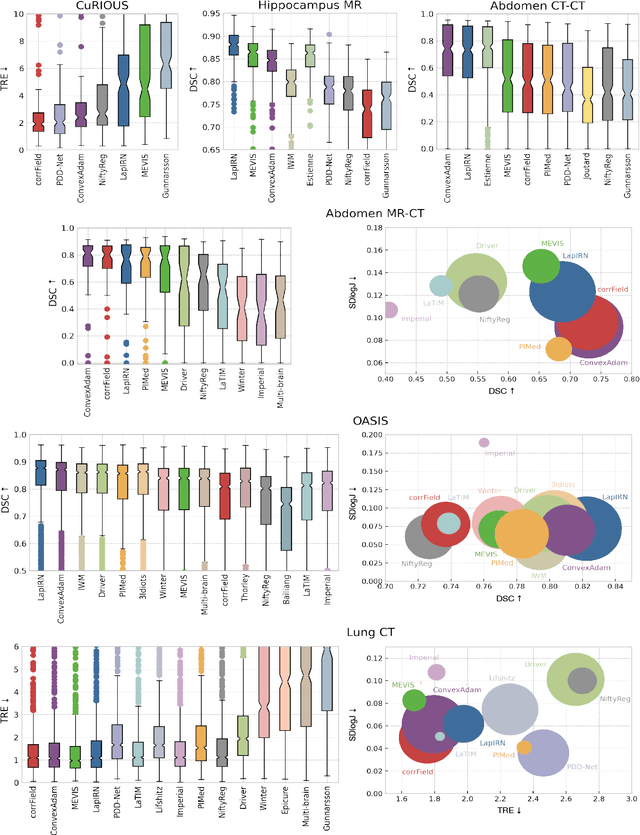
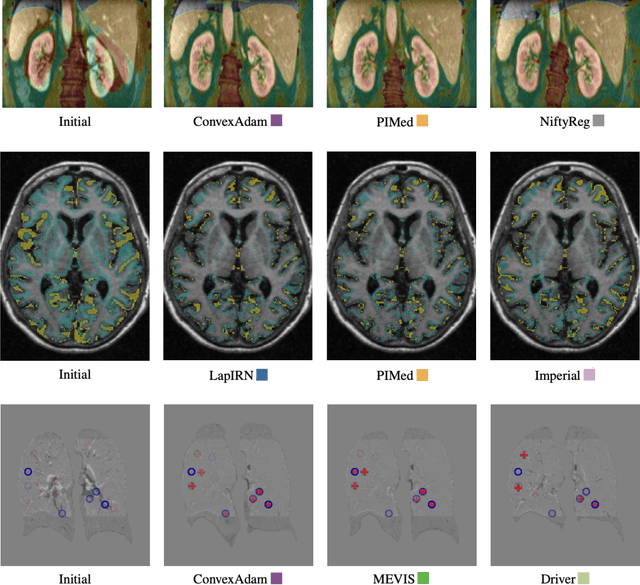
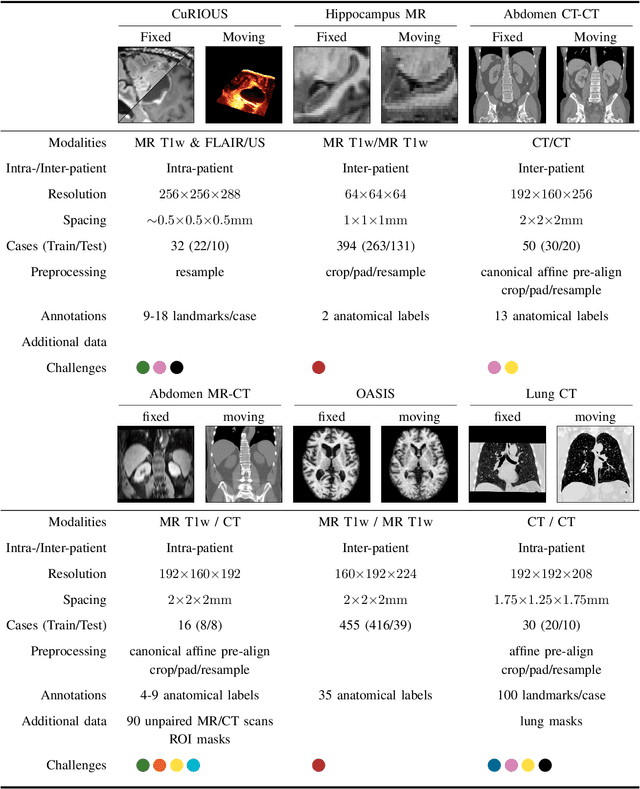
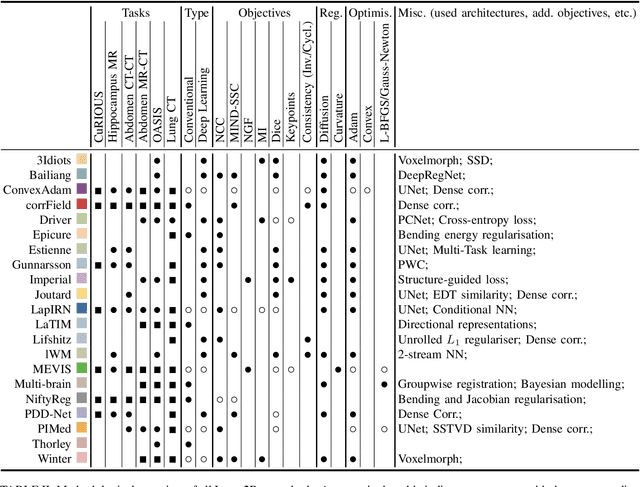
Abstract:Image registration is a fundamental medical image analysis task, and a wide variety of approaches have been proposed. However, only a few studies have comprehensively compared medical image registration approaches on a wide range of clinically relevant tasks, in part because of the lack of availability of such diverse data. This limits the development of registration methods, the adoption of research advances into practice, and a fair benchmark across competing approaches. The Learn2Reg challenge addresses these limitations by providing a multi-task medical image registration benchmark for comprehensive characterisation of deformable registration algorithms. A continuous evaluation will be possible at https://learn2reg.grand-challenge.org. Learn2Reg covers a wide range of anatomies (brain, abdomen, and thorax), modalities (ultrasound, CT, MR), availability of annotations, as well as intra- and inter-patient registration evaluation. We established an easily accessible framework for training and validation of 3D registration methods, which enabled the compilation of results of over 65 individual method submissions from more than 20 unique teams. We used a complementary set of metrics, including robustness, accuracy, plausibility, and runtime, enabling unique insight into the current state-of-the-art of medical image registration. This paper describes datasets, tasks, evaluation methods and results of the challenge, and the results of further analysis of transferability to new datasets, the importance of label supervision, and resulting bias.
Self-Supervised Pre-Training of Swin Transformers for 3D Medical Image Analysis
Nov 29, 2021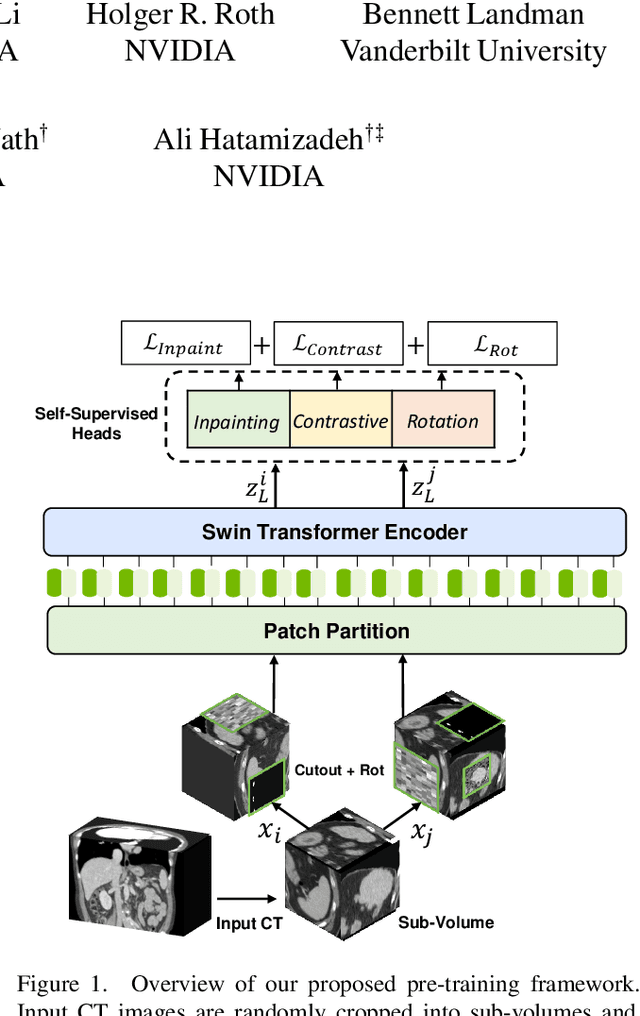
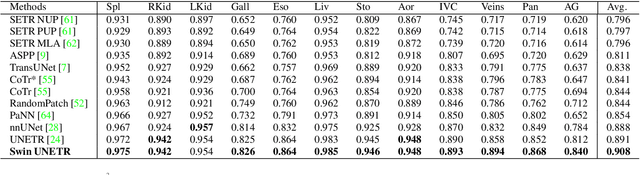
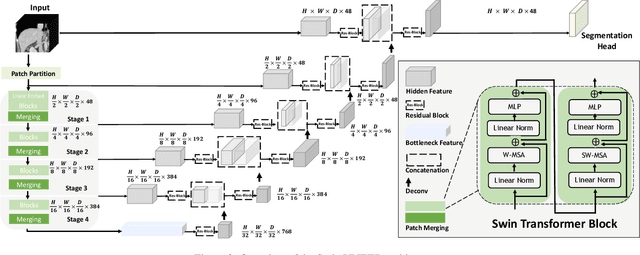
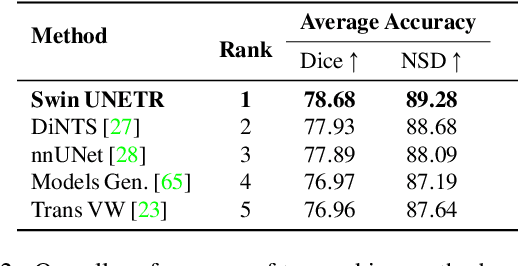
Abstract:Vision Transformers (ViT)s have shown great performance in self-supervised learning of global and local representations that can be transferred to downstream applications. Inspired by these results, we introduce a novel self-supervised learning framework with tailored proxy tasks for medical image analysis. Specifically, we propose: (i) a new 3D transformer-based model, dubbed Swin UNEt TRansformers (Swin UNETR), with a hierarchical encoder for self-supervised pre-training; (ii) tailored proxy tasks for learning the underlying pattern of human anatomy. We demonstrate successful pre-training of the proposed model on 5,050 publicly available computed tomography (CT) images from various body organs. The effectiveness of our approach is validated by fine-tuning the pre-trained models on the Beyond the Cranial Vault (BTCV) Segmentation Challenge with 13 abdominal organs and segmentation tasks from the Medical Segmentation Decathlon (MSD) dataset. Our model is currently the state-of-the-art (i.e. ranked 1st) on the public test leaderboards of both MSD and BTCV datasets. Code: https://monai.io/research/swin-unetr
Multi-Site Infant Brain Segmentation Algorithms: The iSeg-2019 Challenge
Jul 11, 2020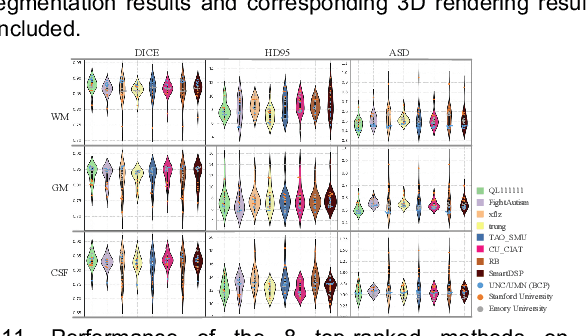
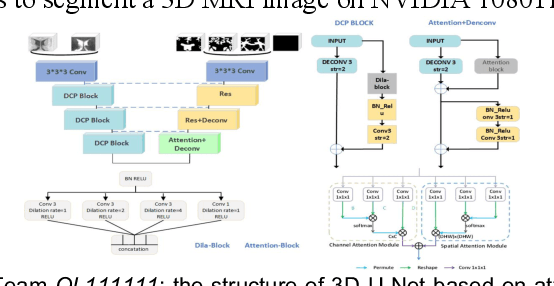
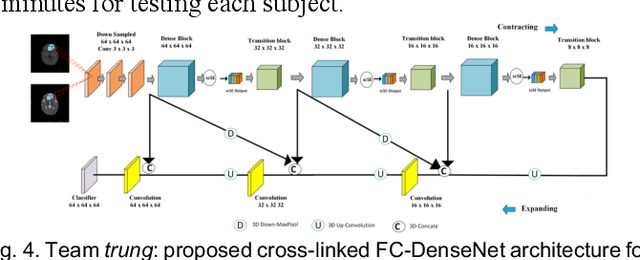
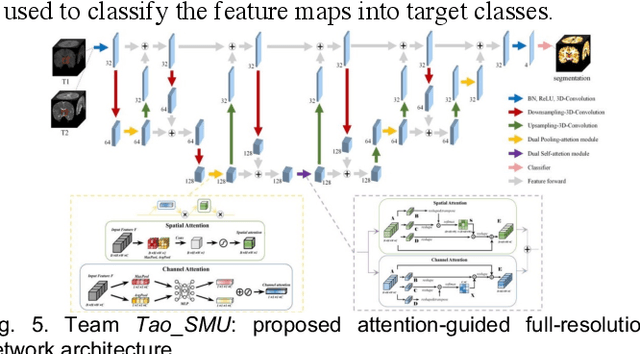
Abstract:To better understand early brain growth patterns in health and disorder, it is critical to accurately segment infant brain magnetic resonance (MR) images into white matter (WM), gray matter (GM), and cerebrospinal fluid (CSF). Deep learning-based methods have achieved state-of-the-art performance; however, one of major limitations is that the learning-based methods may suffer from the multi-site issue, that is, the models trained on a dataset from one site may not be applicable to the datasets acquired from other sites with different imaging protocols/scanners. To promote methodological development in the community, iSeg-2019 challenge (http://iseg2019.web.unc.edu) provides a set of 6-month infant subjects from multiple sites with different protocols/scanners for the participating methods. Training/validation subjects are from UNC (MAP) and testing subjects are from UNC/UMN (BCP), Stanford University, and Emory University. By the time of writing, there are 30 automatic segmentation methods participating in iSeg-2019. We review the 8 top-ranked teams by detailing their pipelines/implementations, presenting experimental results and evaluating performance in terms of the whole brain, regions of interest, and gyral landmark curves. We also discuss their limitations and possible future directions for the multi-site issue. We hope that the multi-site dataset in iSeg-2019 and this review article will attract more researchers on the multi-site issue.
The Future of Digital Health with Federated Learning
Mar 18, 2020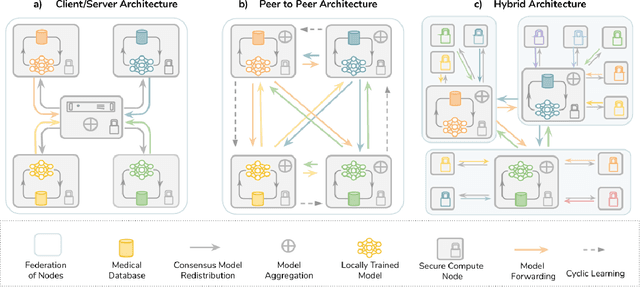
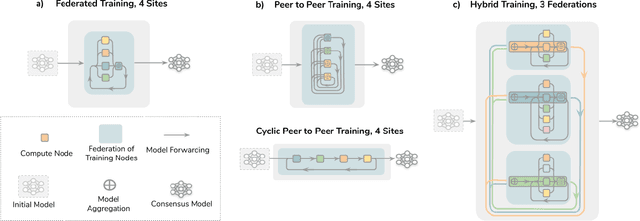
Abstract:Data-driven Machine Learning has emerged as a promising approach for building accurate and robust statistical models from medical data, which is collected in huge volumes by modern healthcare systems. Existing medical data is not fully exploited by ML primarily because it sits in data silos and privacy concerns restrict access to this data. However, without access to sufficient data, ML will be prevented from reaching its full potential and, ultimately, from making the transition from research to clinical practice. This paper considers key factors contributing to this issue, explores how Federated Learning (FL) may provide a solution for the future of digital health and highlights the challenges and considerations that need to be addressed.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge