Uncertainty-Aware Multiple-Instance Learning for Reliable Classification: Application to Optical Coherence Tomography
Feb 06, 2023Coen de Vente, Bram van Ginneken, Carel B. Hoyng, Caroline C. W. Klaver, Clara I. Sánchez
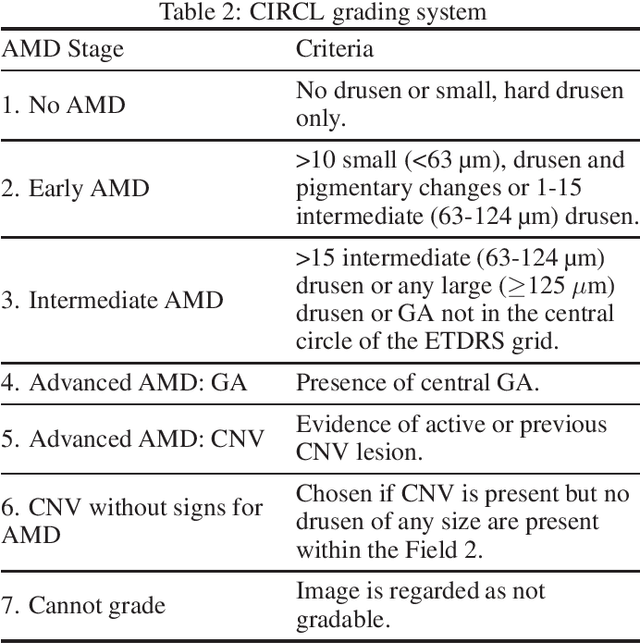
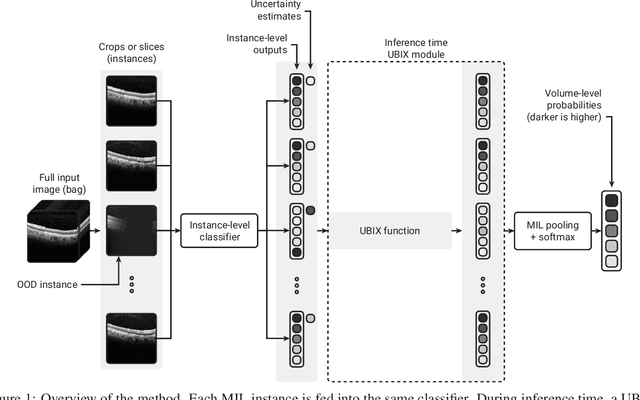
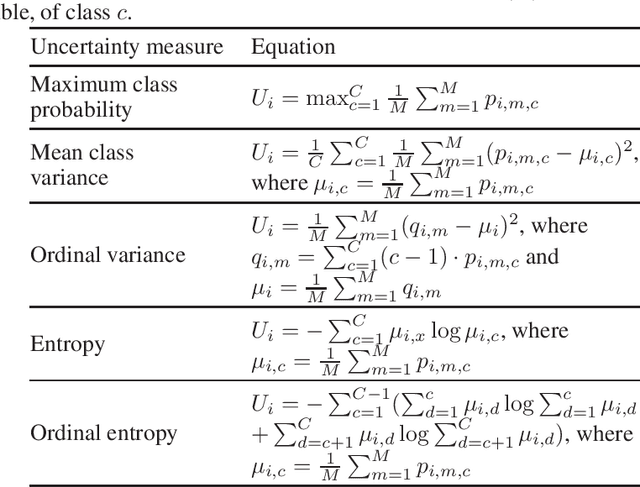
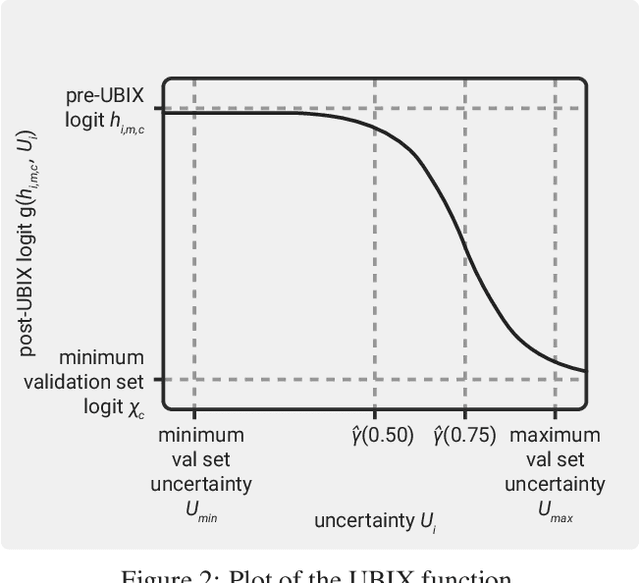

Deep learning classification models for medical image analysis often perform well on data from scanners that were used during training. However, when these models are applied to data from different vendors, their performance tends to drop substantially. Artifacts that only occur within scans from specific scanners are major causes of this poor generalizability. We aimed to improve the reliability of deep learning classification models by proposing Uncertainty-Based Instance eXclusion (UBIX). This technique, based on multiple-instance learning, reduces the effect of corrupted instances on the bag-classification by seamlessly integrating out-of-distribution (OOD) instance detection during inference. Although UBIX is generally applicable to different medical images and diverse classification tasks, we focused on staging of age-related macular degeneration in optical coherence tomography. After being trained using images from one vendor, UBIX showed a reliable behavior, with a slight decrease in performance (a decrease of the quadratic weighted kappa ($\kappa_w$) from 0.861 to 0.708), when applied to images from different vendors containing artifacts; while a state-of-the-art 3D neural network suffered from a significant detriment of performance ($\kappa_w$ from 0.852 to 0.084) on the same test set. We showed that instances with unseen artifacts can be identified with OOD detection and their contribution to the bag-level predictions can be reduced, improving reliability without the need for retraining on new data. This potentially increases the applicability of artificial intelligence models to data from other scanners than the ones for which they were developed.
Understanding metric-related pitfalls in image analysis validation
Feb 03, 2023Annika Reinke, Minu D. Tizabi, Michael Baumgartner, Matthias Eisenmann, Doreen Heckmann-Nötzel, A. Emre Kavu, Tim Rädsch, Carole H. Sudre, Laura Acion, Michela Antonelli, Tal Arbel, Spyridon Bakas, Arriel Benis, Matthew Blaschko, Florian Büttner, M. Jorge Cardoso, Veronika Cheplygina, Jianxu Chen, Evangelia Christodoulou, Beth A. Cimini, Gary S. Collins, Keyvan Farahani, Luciana Ferrer, Adrian Galdran, Bram van Ginneken, Ben Glocker, Patrick Godau, Robert Haase, Daniel A. Hashimoto, Michael M. Hoffman, Merel Huisman, Fabian Isensee, Pierre Jannin, Charles E. Kahn, Dagmar Kainmueller, Bernhard Kainz, Alexandros Karargyris, Alan Karthikesalingam, Hannes Kenngott, Jens Kleesiek, Florian Kofler, Thijs Kooi, Annette Kopp-Schneider, Michal Kozubek, Anna Kreshuk, Tahsin Kurc, Bennett A. Landman, Geert Litjens, Amin Madani, Klaus Maier-Hein, Anne L. Martel, Peter Mattson, Erik Meijering, Bjoern Menze, Karel G. M. Moons, Henning Müller, Brennan Nichyporuk, Felix Nickel, Jens Petersen, Susanne M. Rafelski, Nasir Rajpoot, Mauricio Reyes, Michael A. Riegler, Nicola Rieke, Julio Saez-Rodriguez, Clara I. Sánchez, Shravya Shetty, Maarten van Smeden, Ronald M. Summers, Abdel A. Taha, Aleksei Tiulpin, Sotirios A. Tsaftaris, Ben Van Calster, Gaël Varoquaux, Manuel Wiesenfarth, Ziv R. Yaniv, Paul F. Jäger, Lena Maier-Hein
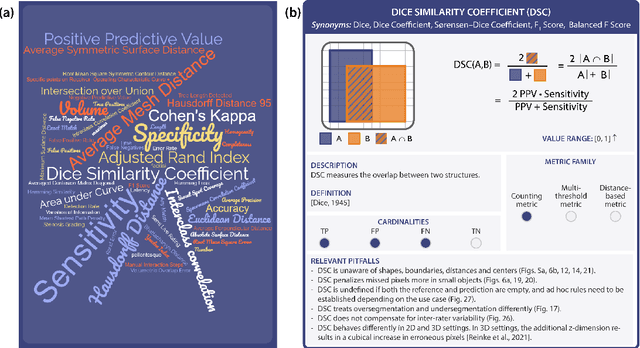
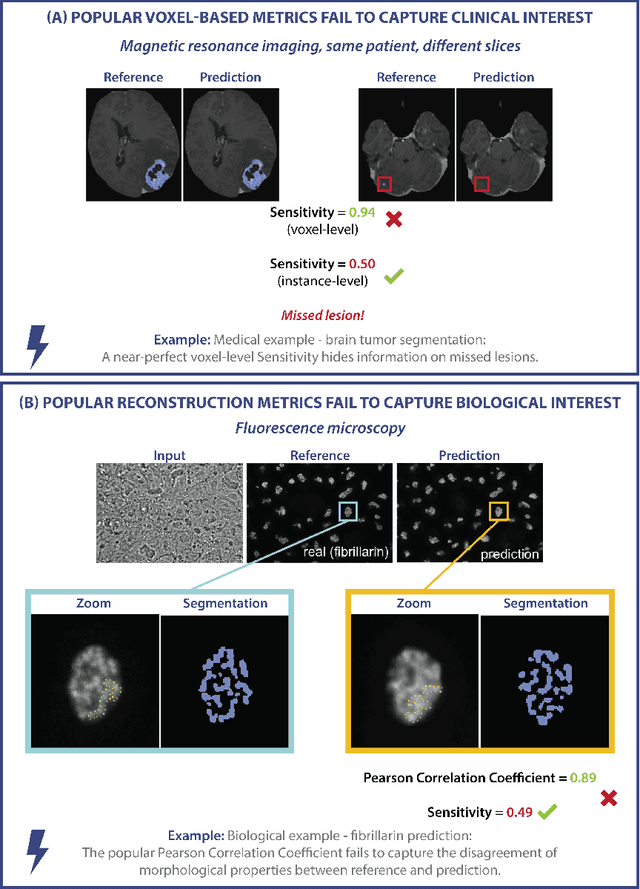
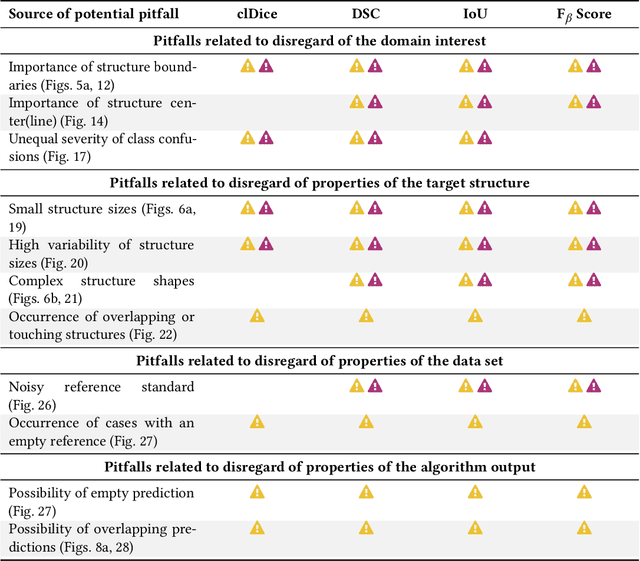

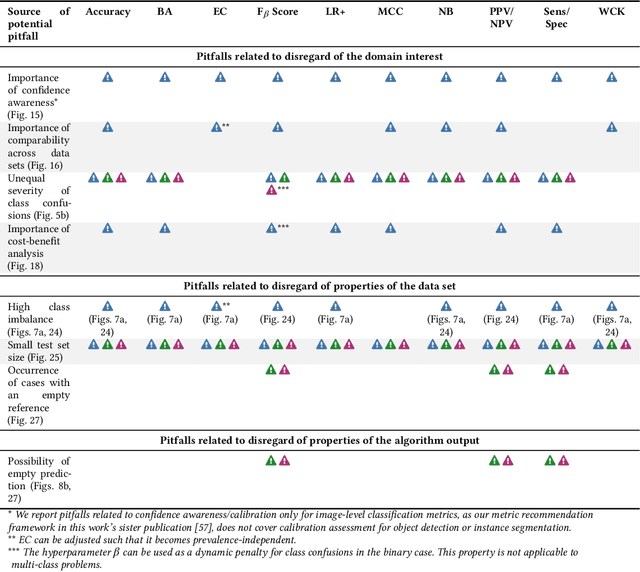
Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, the present work provides the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
AIROGS: Artificial Intelligence for RObust Glaucoma Screening Challenge
Feb 03, 2023Coen de Vente, Koenraad A. Vermeer, Nicolas Jaccard, He Wang, Hongyi Sun, Firas Khader, Daniel Truhn, Temirgali Aimyshev, Yerkebulan Zhanibekuly, Tien-Dung Le, Adrian Galdran, Miguel Ángel González Ballester, Gustavo Carneiro, Devika R G, Hrishikesh P S, Densen Puthussery, Hong Liu, Zekang Yang, Satoshi Kondo, Satoshi Kasai, Edward Wang, Ashritha Durvasula, Jónathan Heras, Miguel Ángel Zapata, Teresa Araújo, Guilherme Aresta, Hrvoje Bogunović, Mustafa Arikan, Yeong Chan Lee, Hyun Bin Cho, Yoon Ho Choi, Abdul Qayyum, Imran Razzak, Bram van Ginneken, Hans G. Lemij, Clara I. Sánchez
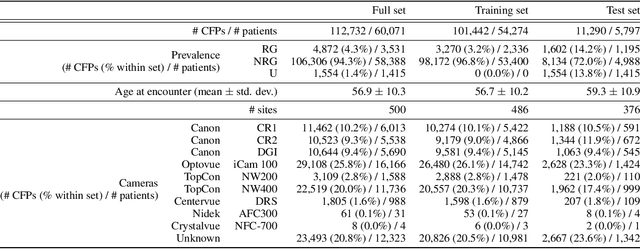
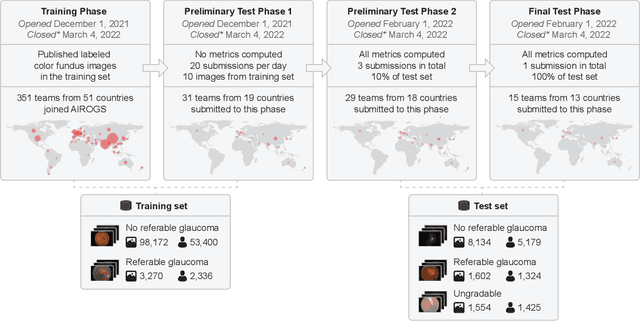
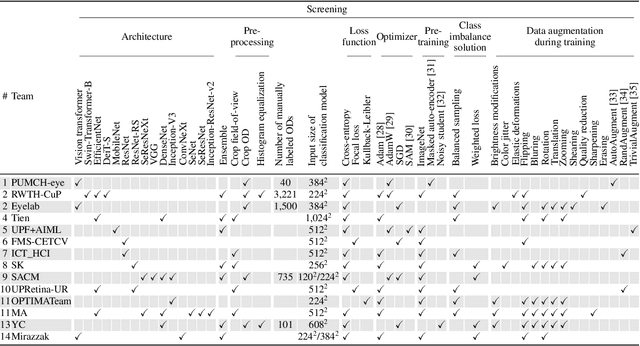
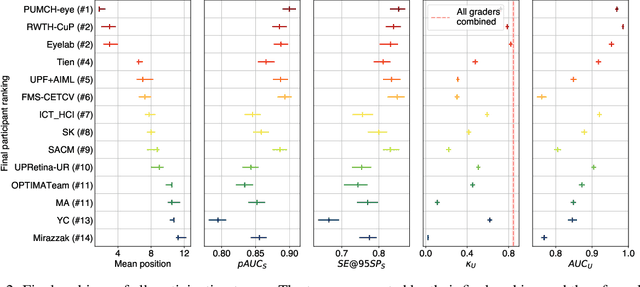
The early detection of glaucoma is essential in preventing visual impairment. Artificial intelligence (AI) can be used to analyze color fundus photographs (CFPs) in a cost-effective manner, making glaucoma screening more accessible. While AI models for glaucoma screening from CFPs have shown promising results in laboratory settings, their performance decreases significantly in real-world scenarios due to the presence of out-of-distribution and low-quality images. To address this issue, we propose the Artificial Intelligence for Robust Glaucoma Screening (AIROGS) challenge. This challenge includes a large dataset of around 113,000 images from about 60,000 patients and 500 different screening centers, and encourages the development of algorithms that are robust to ungradable and unexpected input data. We evaluated solutions from 14 teams in this paper, and found that the best teams performed similarly to a set of 20 expert ophthalmologists and optometrists. The highest-scoring team achieved an area under the receiver operating characteristic curve of 0.99 (95% CI: 0.98-0.99) for detecting ungradable images on-the-fly. Additionally, many of the algorithms showed robust performance when tested on three other publicly available datasets. These results demonstrate the feasibility of robust AI-enabled glaucoma screening.
Domain adaptation strategies for cancer-independent detection of lymph node metastases
Jul 13, 2022Péter Bándi, Maschenka Balkenhol, Marcory van Dijk, Bram van Ginneken, Jeroen van der Laak, Geert Litjens
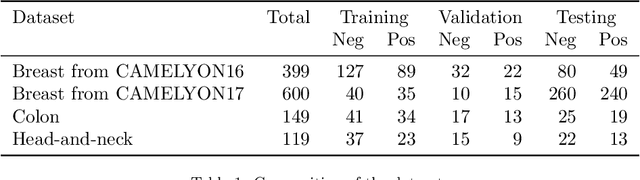
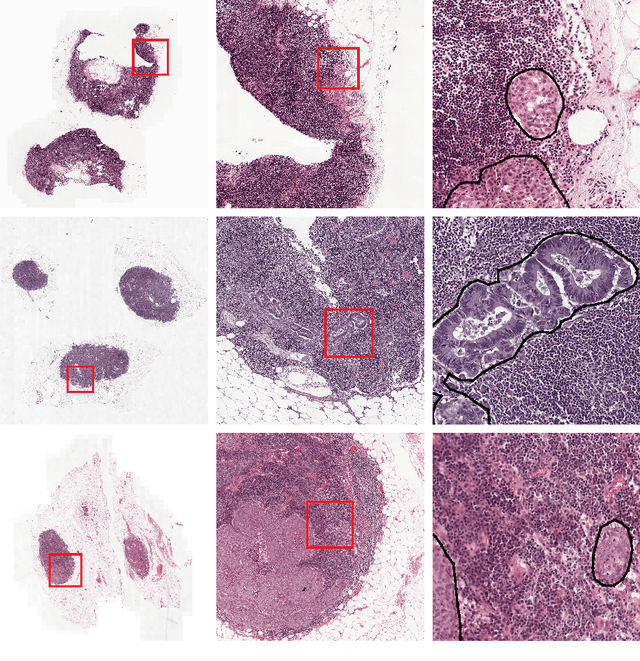
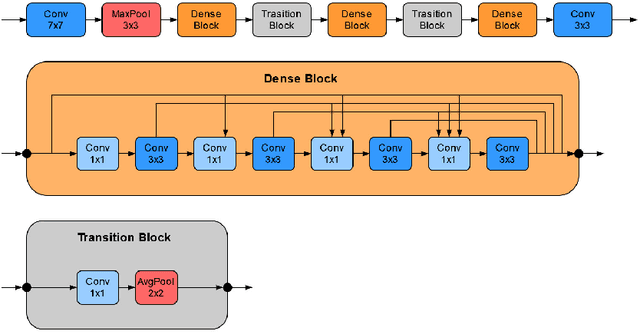
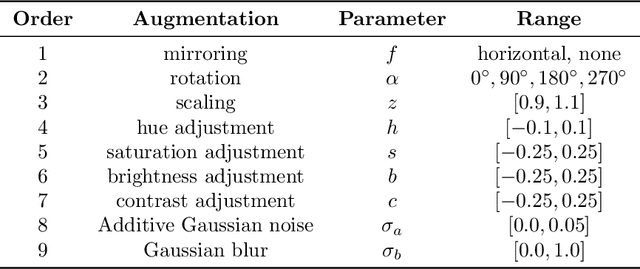
Recently, large, high-quality public datasets have led to the development of convolutional neural networks that can detect lymph node metastases of breast cancer at the level of expert pathologists. Many cancers, regardless of the site of origin, can metastasize to lymph nodes. However, collecting and annotating high-volume, high-quality datasets for every cancer type is challenging. In this paper we investigate how to leverage existing high-quality datasets most efficiently in multi-task settings for closely related tasks. Specifically, we will explore different training and domain adaptation strategies, including prevention of catastrophic forgetting, for colon and head-and-neck cancer metastasis detection in lymph nodes. Our results show state-of-the-art performance on both cancer metastasis detection tasks. Furthermore, we show the effectiveness of repeated adaptation of networks from one cancer type to another to obtain multi-task metastasis detection networks. Last, we show that leveraging existing high-quality datasets can significantly boost performance on new target tasks and that catastrophic forgetting can be effectively mitigated using regularization.
Metrics reloaded: Pitfalls and recommendations for image analysis validation
Jun 03, 2022Lena Maier-Hein, Annika Reinke, Evangelia Christodoulou, Ben Glocker, Patrick Godau, Fabian Isensee, Jens Kleesiek, Michal Kozubek, Mauricio Reyes, Michael A. Riegler, Manuel Wiesenfarth, Michael Baumgartner, Matthias Eisenmann, Doreen Heckmann-Nötzel, A. Emre Kavur, Tim Rädsch, Minu D. Tizabi, Laura Acion, Michela Antonelli, Tal Arbel, Spyridon Bakas, Peter Bankhead, Arriel Benis, M. Jorge Cardoso, Veronika Cheplygina, Beth Cimini, Gary S. Collins, Keyvan Farahani, Bram van Ginneken, Daniel A. Hashimoto, Michael M. Hoffman, Merel Huisman, Pierre Jannin, Charles E. Kahn, Alexandros Karargyris, Alan Karthikesalingam, Hannes Kenngott, Annette Kopp-Schneider, Anna Kreshuk, Tahsin Kurc, Bennett A. Landman, Geert Litjens, Amin Madani, Klaus Maier-Hein, Anne L. Martel, Peter Mattson, Erik Meijering, Bjoern Menze, David Moher, Karel G. M. Moons, Henning Müller, Felix Nickel, Brennan Nichyporuk, Jens Petersen, Nasir Rajpoot, Nicola Rieke, Julio Saez-Rodriguez, Clarisa Sánchez Gutiérrez, Shravya Shetty, Maarten van Smeden, Carole H. Sudre, Ronald M. Summers, Abdel A. Taha, Sotirios A. Tsaftaris, Ben Van Calster, Gaël Varoquaux, Paul F. Jäger
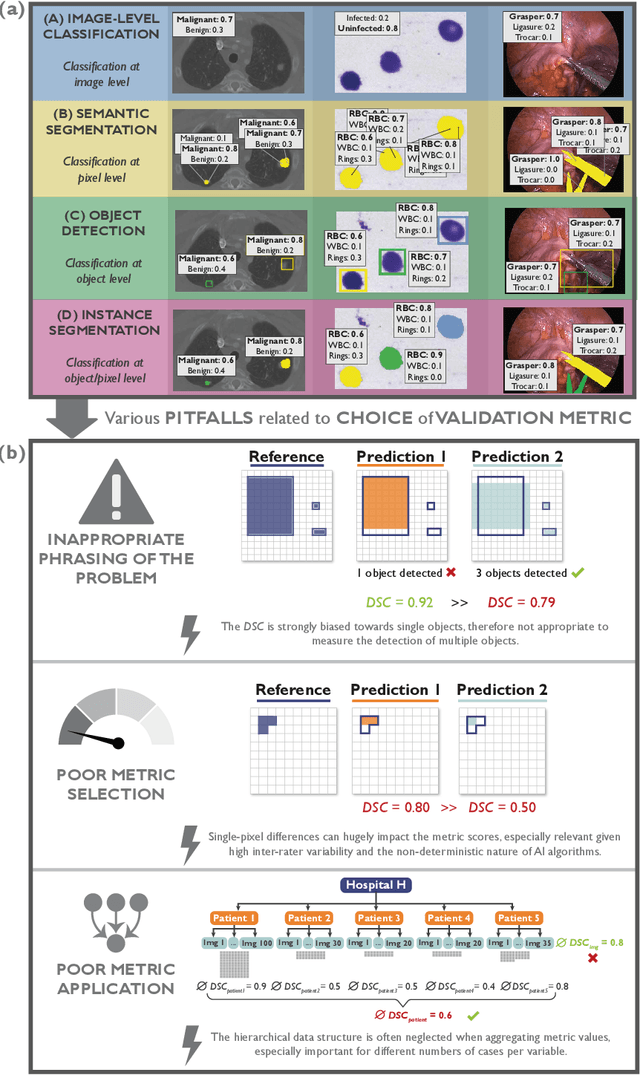
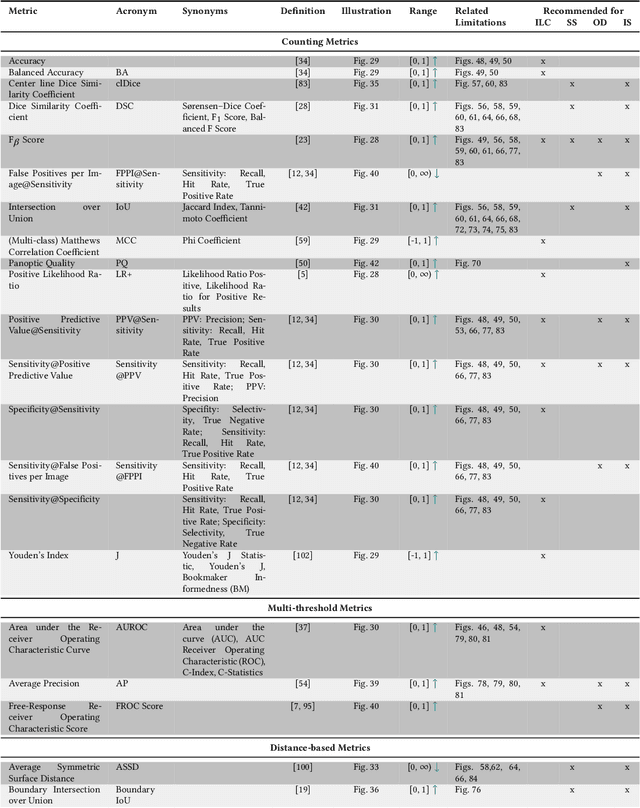
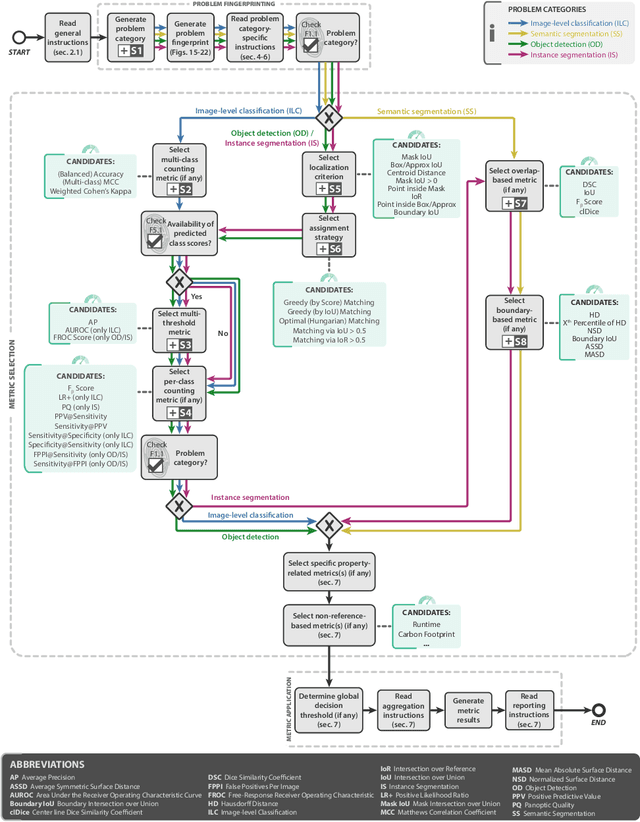
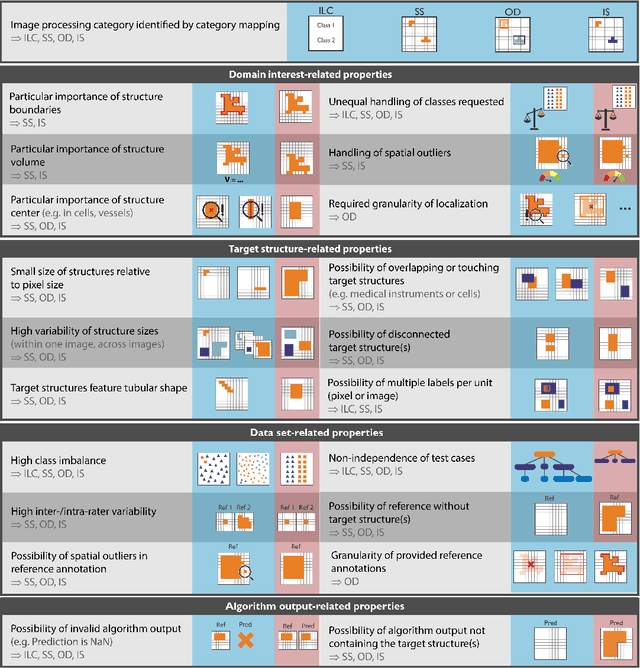
The field of automatic biomedical image analysis crucially depends on robust and meaningful performance metrics for algorithm validation. Current metric usage, however, is often ill-informed and does not reflect the underlying domain interest. Here, we present a comprehensive framework that guides researchers towards choosing performance metrics in a problem-aware manner. Specifically, we focus on biomedical image analysis problems that can be interpreted as a classification task at image, object or pixel level. The framework first compiles domain interest-, target structure-, data set- and algorithm output-related properties of a given problem into a problem fingerprint, while also mapping it to the appropriate problem category, namely image-level classification, semantic segmentation, instance segmentation, or object detection. It then guides users through the process of selecting and applying a set of appropriate validation metrics while making them aware of potential pitfalls related to individual choices. In this paper, we describe the current status of the Metrics Reloaded recommendation framework, with the goal of obtaining constructive feedback from the image analysis community. The current version has been developed within an international consortium of more than 60 image analysis experts and will be made openly available as a user-friendly toolkit after community-driven optimization.
Structure and position-aware graph neural network for airway labeling
Jan 12, 2022Weiyi Xie, Colin Jacobs, Jean-Paul Charbonnier, Bram van Ginneken
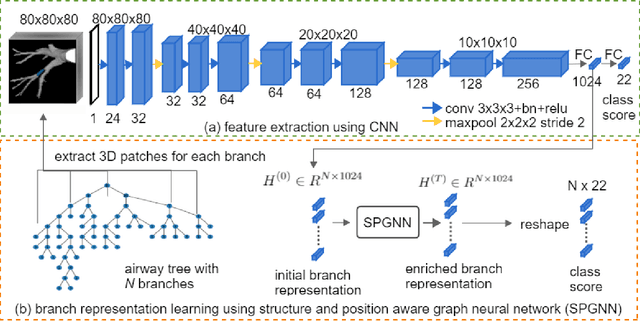
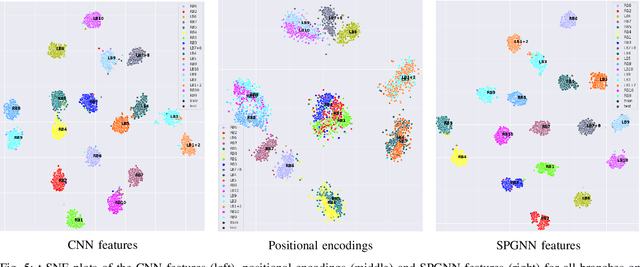
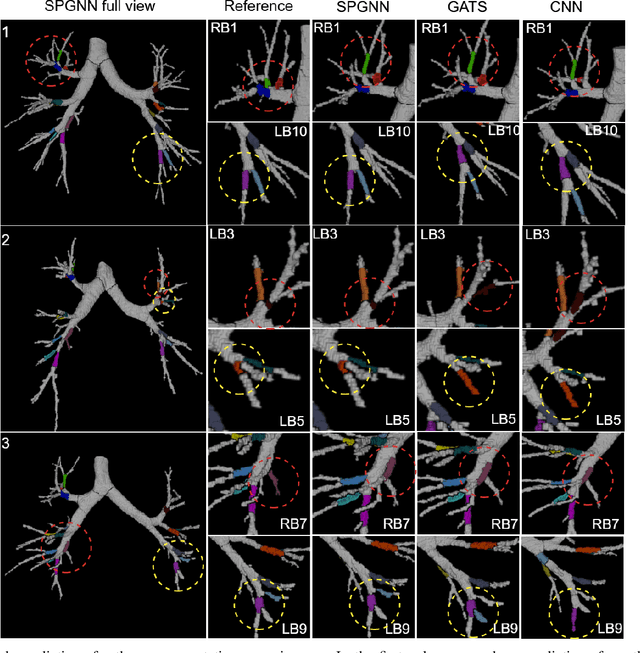
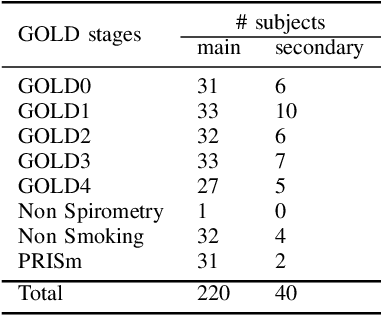
We present a novel graph-based approach for labeling the anatomical branches of a given airway tree segmentation. The proposed method formulates airway labeling as a branch classification problem in the airway tree graph, where branch features are extracted using convolutional neural networks (CNN) and enriched using graph neural networks. Our graph neural network is structure-aware by having each node aggregate information from its local neighbors and position-aware by encoding node positions in the graph. We evaluated the proposed method on 220 airway trees from subjects with various severity stages of Chronic Obstructive Pulmonary Disease (COPD). The results demonstrate that our approach is computationally efficient and significantly improves branch classification performance than the baseline method. The overall average accuracy of our method reaches 91.18\% for labeling all 18 segmental airway branches, compared to 83.83\% obtained by the standard CNN method. We published our source code at https://github.com/DIAGNijmegen/spgnn. The proposed algorithm is also publicly available at https://grand-challenge.org/algorithms/airway-anatomical-labeling/.
Learn2Reg: comprehensive multi-task medical image registration challenge, dataset and evaluation in the era of deep learning
Dec 23, 2021Alessa Hering, Lasse Hansen, Tony C. W. Mok, Albert C. S. Chung, Hanna Siebert, Stephanie Häger, Annkristin Lange, Sven Kuckertz, Stefan Heldmann, Wei Shao, Sulaiman Vesal, Mirabela Rusu, Geoffrey Sonn, Théo Estienne, Maria Vakalopoulou, Luyi Han, Yunzhi Huang, Mikael Brudfors, Yaël Balbastre, SamuelJ outard, Marc Modat, Gal Lifshitz, Dan Raviv, Jinxin Lv, Qiang Li, Vincent Jaouen, Dimitris Visvikis, Constance Fourcade, Mathieu Rubeaux, Wentao Pan, Zhe Xu, Bailiang Jian, Francesca De Benetti, Marek Wodzinski, Niklas Gunnarsson, Jens Sjölund, Huaqi Qiu, Zeju Li, Christoph Großbröhmer, Andrew Hoopes, Ingerid Reinertsen, Yiming Xiao, Bennett Landman, Yuankai Huo, Keelin Murphy, Nikolas Lessmann, Bram van Ginneken, Adrian V. Dalca, Mattias P. Heinrich
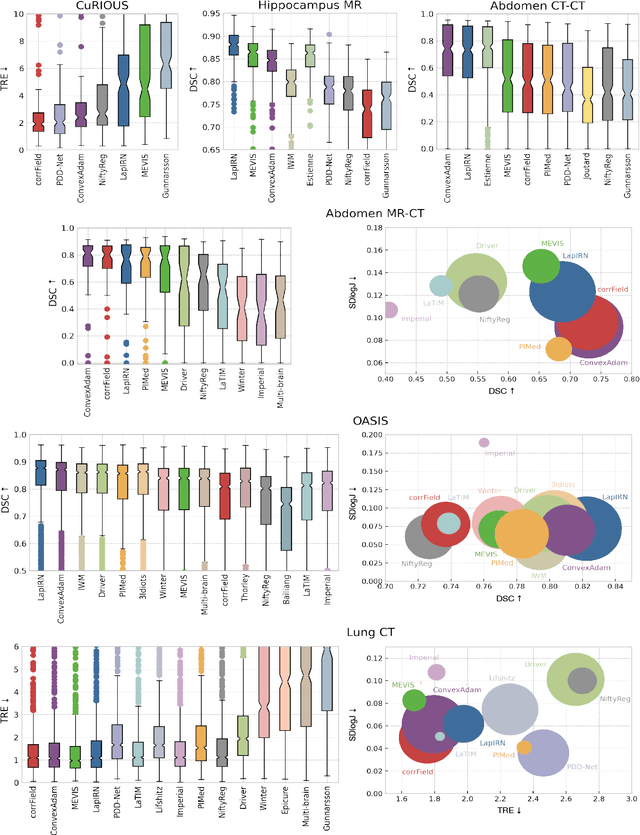
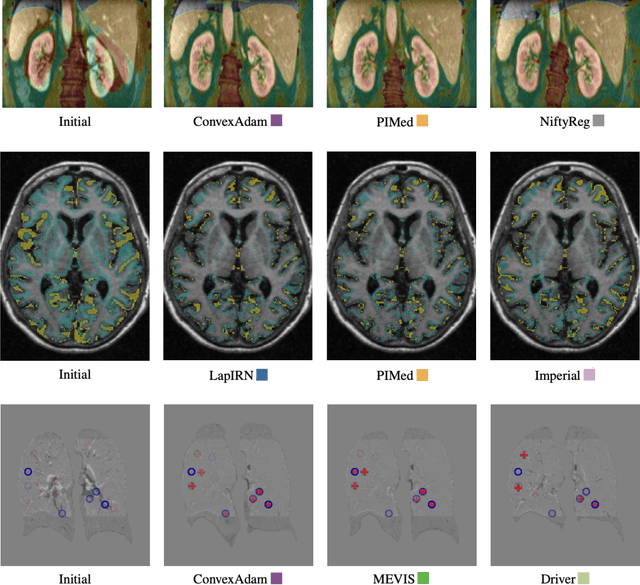
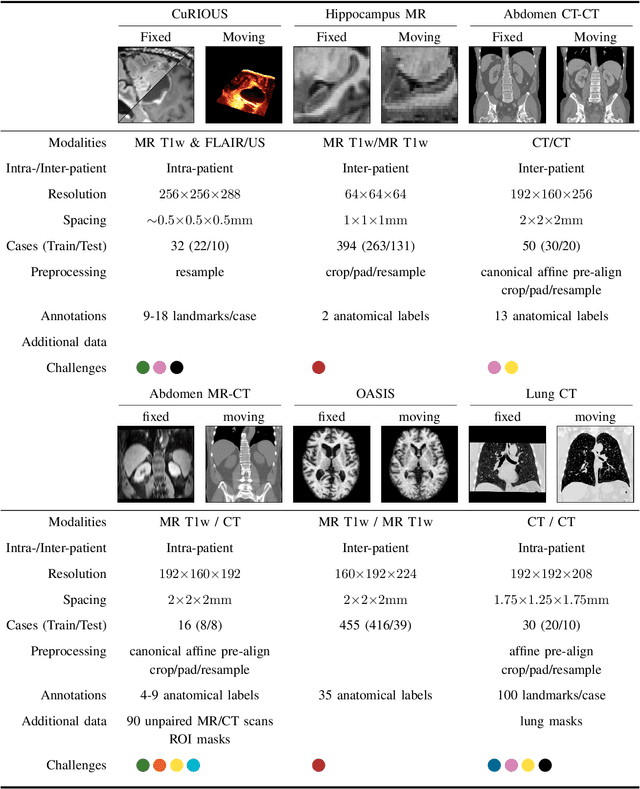
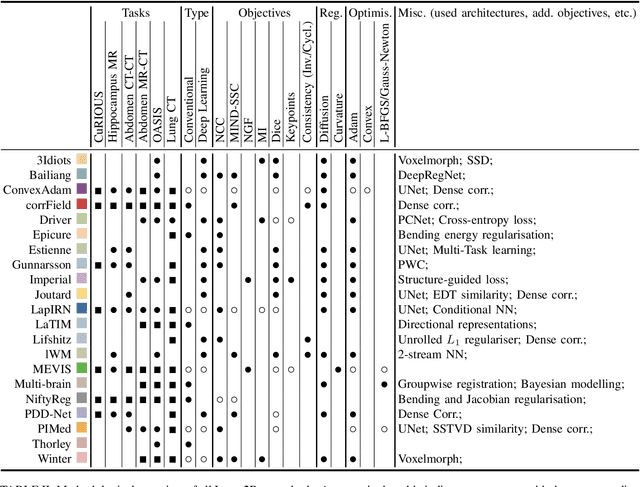
Image registration is a fundamental medical image analysis task, and a wide variety of approaches have been proposed. However, only a few studies have comprehensively compared medical image registration approaches on a wide range of clinically relevant tasks, in part because of the lack of availability of such diverse data. This limits the development of registration methods, the adoption of research advances into practice, and a fair benchmark across competing approaches. The Learn2Reg challenge addresses these limitations by providing a multi-task medical image registration benchmark for comprehensive characterisation of deformable registration algorithms. A continuous evaluation will be possible at https://learn2reg.grand-challenge.org. Learn2Reg covers a wide range of anatomies (brain, abdomen, and thorax), modalities (ultrasound, CT, MR), availability of annotations, as well as intra- and inter-patient registration evaluation. We established an easily accessible framework for training and validation of 3D registration methods, which enabled the compilation of results of over 65 individual method submissions from more than 20 unique teams. We used a complementary set of metrics, including robustness, accuracy, plausibility, and runtime, enabling unique insight into the current state-of-the-art of medical image registration. This paper describes datasets, tasks, evaluation methods and results of the challenge, and the results of further analysis of transferability to new datasets, the importance of label supervision, and resulting bias.
Robust Segmentation Models using an Uncertainty Slice Sampling Based Annotation Workflow
Sep 30, 2021Grzegorz Chlebus, Andrea Schenk, Horst K. Hahn, Bram van Ginneken, Hans Meine
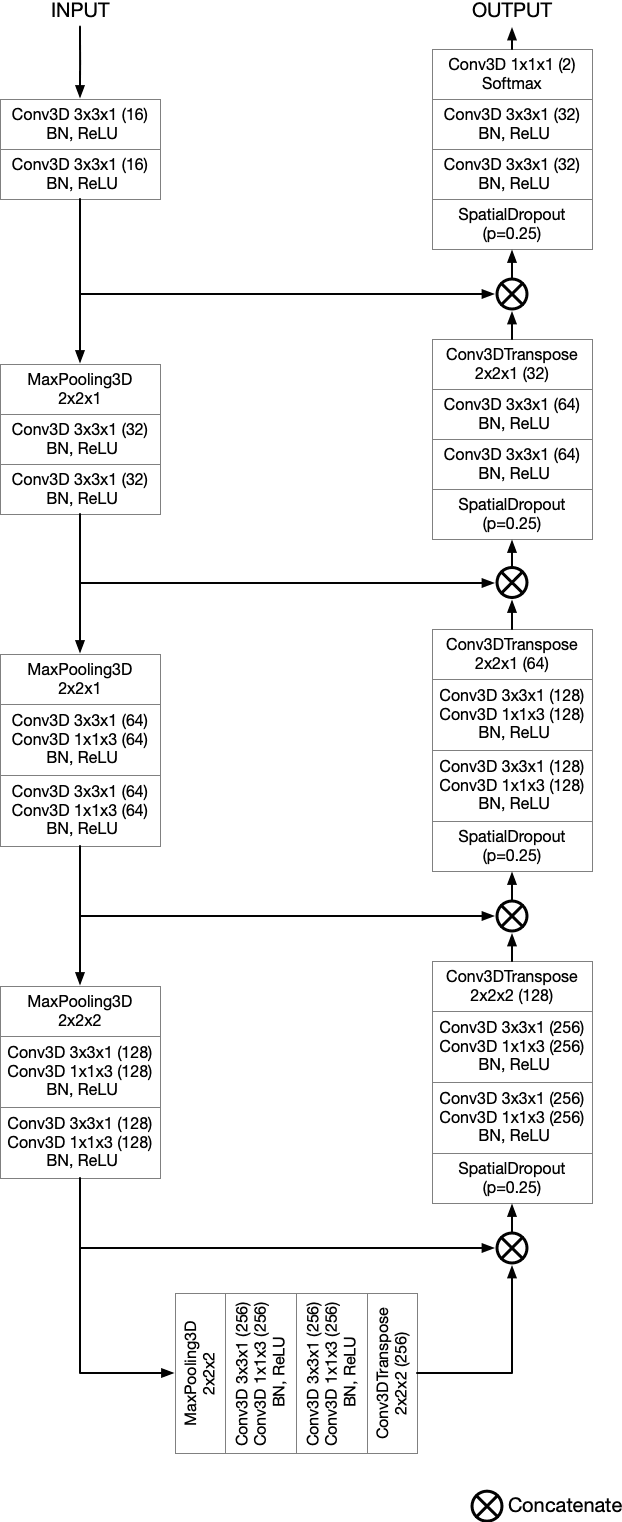
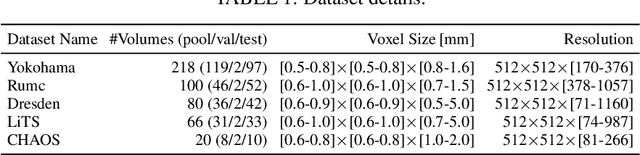
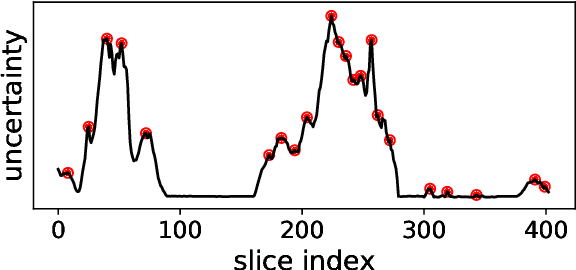
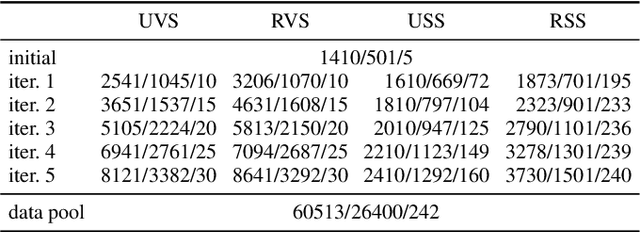
Semantic segmentation neural networks require pixel-level annotations in large quantities to achieve a good performance. In the medical domain, such annotations are expensive, because they are time-consuming and require expert knowledge. Active learning optimizes the annotation effort by devising strategies to select cases for labeling that are most informative to the model. In this work, we propose an uncertainty slice sampling (USS) strategy for semantic segmentation of 3D medical volumes that selects 2D image slices for annotation and compare it with various other strategies. We demonstrate the efficiency of USS on a CT liver segmentation task using multi-site data. After five iterations, the training data resulting from USS consisted of 2410 slices (4% of all slices in the data pool) compared to 8121 (13%), 8641 (14%), and 3730 (6%) for uncertainty volume (UVS), random volume (RVS), and random slice (RSS) sampling, respectively. Despite being trained on the smallest amount of data, the model based on the USS strategy evaluated on 234 test volumes significantly outperformed models trained according to other strategies and achieved a mean Dice index of 0.964, a relative volume error of 4.2%, a mean surface distance of 1.35 mm, and a Hausdorff distance of 23.4 mm. This was only slightly inferior to 0.967, 3.8%, 1.18 mm, and 22.9 mm achieved by a model trained on all available data, but the robustness analysis using the 5th percentile of Dice and the 95th percentile of the remaining metrics demonstrated that USS resulted not only in the most robust model compared to other sampling schemes, but also outperformed the model trained on all data according to Dice (0.946 vs. 0.945) and mean surface distance (1.92 mm vs. 2.03 mm).
The Medical Segmentation Decathlon
Jun 10, 2021Michela Antonelli, Annika Reinke, Spyridon Bakas, Keyvan Farahani, AnnetteKopp-Schneider, Bennett A. Landman, Geert Litjens, Bjoern Menze, Olaf Ronneberger, Ronald M. Summers, Bram van Ginneken, Michel Bilello, Patrick Bilic, Patrick F. Christ, Richard K. G. Do, Marc J. Gollub, Stephan H. Heckers, Henkjan Huisman, William R. Jarnagin, Maureen K. McHugo, Sandy Napel, Jennifer S. Goli Pernicka, Kawal Rhode, Catalina Tobon-Gomez, Eugene Vorontsov, Henkjan Huisman, James A. Meakin, Sebastien Ourselin, Manuel Wiesenfarth, Pablo Arbelaez, Byeonguk Bae, Sihong Chen, Laura Daza, Jianjiang Feng, Baochun He, Fabian Isensee, Yuanfeng Ji, Fucang Jia, Namkug Kim, Ildoo Kim, Dorit Merhof, Akshay Pai, Beomhee Park, Mathias Perslev, Ramin Rezaiifar, Oliver Rippel, Ignacio Sarasua, Wei Shen, Jaemin Son, Christian Wachinger, Liansheng Wang, Yan Wang, Yingda Xia, Daguang Xu, Zhanwei Xu, Yefeng Zheng, Amber L. Simpson, Lena Maier-Hein, M. Jorge Cardoso
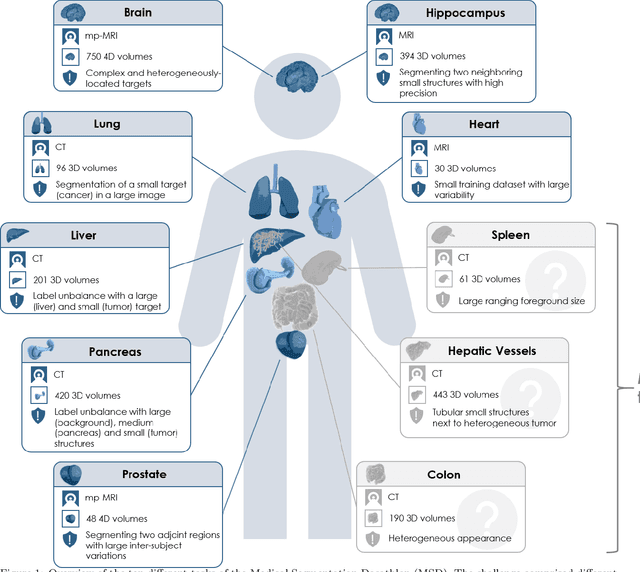
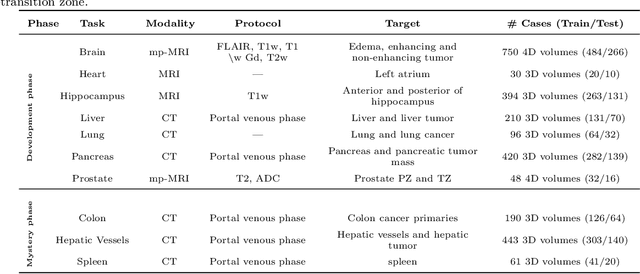
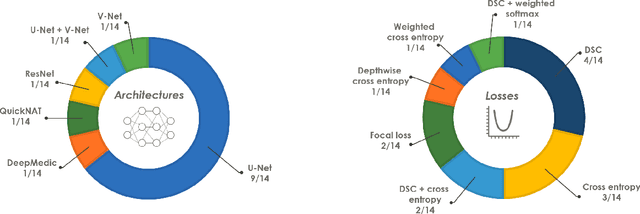
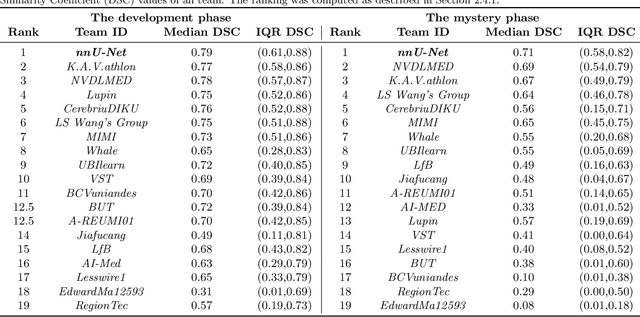
International challenges have become the de facto standard for comparative assessment of image analysis algorithms given a specific task. Segmentation is so far the most widely investigated medical image processing task, but the various segmentation challenges have typically been organized in isolation, such that algorithm development was driven by the need to tackle a single specific clinical problem. We hypothesized that a method capable of performing well on multiple tasks will generalize well to a previously unseen task and potentially outperform a custom-designed solution. To investigate the hypothesis, we organized the Medical Segmentation Decathlon (MSD) - a biomedical image analysis challenge, in which algorithms compete in a multitude of both tasks and modalities. The underlying data set was designed to explore the axis of difficulties typically encountered when dealing with medical images, such as small data sets, unbalanced labels, multi-site data and small objects. The MSD challenge confirmed that algorithms with a consistent good performance on a set of tasks preserved their good average performance on a different set of previously unseen tasks. Moreover, by monitoring the MSD winner for two years, we found that this algorithm continued generalizing well to a wide range of other clinical problems, further confirming our hypothesis. Three main conclusions can be drawn from this study: (1) state-of-the-art image segmentation algorithms are mature, accurate, and generalize well when retrained on unseen tasks; (2) consistent algorithmic performance across multiple tasks is a strong surrogate of algorithmic generalizability; (3) the training of accurate AI segmentation models is now commoditized to non AI experts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge