Dorit Merhof
Gated Differential Linear Attention: A Linear-Time Decoder for High-Fidelity Medical Segmentation
Mar 05, 2026Abstract:Medical image segmentation requires models that preserve fine anatomical boundaries while remaining efficient for clinical deployment. While transformers capture long-range dependencies, they suffer from quadratic attention cost and large data requirements, whereas CNNs are compute-friendly yet struggle with global reasoning. Linear attention offers $\mathcal{O}(N)$ scaling, but often exhibits training instability and attention dilution, yielding diffuse maps. We introduce PVT-GDLA, a decoder-centric Transformer that restores sharp, long-range dependencies at linear time. Its core, Gated Differential Linear Attention (GDLA), computes two kernelized attention paths on complementary query/key subspaces and subtracts them with a learnable, channel-wise scale to cancel common-mode noise and amplify relevant context. A lightweight, head-specific gate injects nonlinearity and input-adaptive sparsity, mitigating attention sink, and a parallel local token-mixing branch with depthwise convolution strengthens neighboring-token interactions, improving boundary fidelity, all while retaining $\mathcal{O}(N)$ complexity and low parameter overhead. Coupled with a pretrained Pyramid Vision Transformer (PVT) encoder, PVT-GDLA achieves state-of-the-art accuracy across CT, MRI, ultrasound, and dermoscopy benchmarks under equal training budgets, with comparable parameters but lower FLOPs than CNN-, Transformer-, hybrid-, and linear-attention baselines. PVT-GDLA provides a practical path to fast, scalable, high-fidelity medical segmentation in clinical environments and other resource-constrained settings.
Footprint-Guided Exemplar-Free Continual Histopathology Report Generation
Feb 27, 2026Abstract:Rapid progress in vision-language modeling has enabled pathology report generation from gigapixel whole-slide images, but most approaches assume static training with simultaneous access to all data. In clinical deployment, however, new organs, institutions, and reporting conventions emerge over time, and sequential fine-tuning can cause catastrophic forgetting. We introduce an exemplar-free continual learning framework for WSI-to-report generation that avoids storing raw slides or patch exemplars. The core idea is a compact domain footprint built in a frozen patch-embedding space: a small codebook of representative morphology tokens together with slide-level co-occurrence summaries and lightweight patch-count priors. These footprints support generative replay by synthesizing pseudo-WSI representations that reflect domain-specific morphological mixtures, while a teacher snapshot provides pseudo-reports to supervise the updated model without retaining past data. To address shifting reporting conventions, we distill domain-specific linguistic characteristics into a compact style descriptor and use it to steer generation. At inference, the model identifies the most compatible descriptor directly from the slide signal, enabling domain-agnostic setup without requiring explicit domain identifiers. Evaluated across multiple public continual learning benchmarks, our approach outperforms exemplar-free and limited-buffer rehearsal baselines, highlighting footprint-based generative replay as a practical solution for deployment in evolving clinical settings.
Towards Modality-Agnostic Continual Domain-Incremental Brain Lesion Segmentation
Jan 20, 2026Abstract:Brain lesion segmentation from multi-modal MRI often assumes fixed modality sets or predefined pathologies, making existing models difficult to adapt across cohorts and imaging protocols. Continual learning (CL) offers a natural solution but current approaches either impose a maximum modality configuration or suffer from severe forgetting in buffer-free settings. We introduce CLMU-Net, a replay-based CL framework for 3D brain lesion segmentation that supports arbitrary and variable modality combinations without requiring prior knowledge of the maximum set. A conceptually simple yet effective channel-inflation strategy maps any modality subset into a unified multi-channel representation, enabling a single model to operate across diverse datasets. To enrich inherently local 3D patch features, we incorporate lightweight domain-conditioned textual embeddings that provide global modality-disease context for each training case. Forgetting is further reduced through principled replay using a compact buffer composed of both prototypical and challenging samples. Experiments on five heterogeneous MRI brain datasets demonstrate that CLMU-Net consistently outperforms popular CL baselines. Notably, our method yields an average Dice score improvement of $\geq$ 18\% while remaining robust under heterogeneous-modality conditions. These findings underscore the value of flexible modality handling, targeted replay, and global contextual cues for continual medical image segmentation. Our implementation is available at https://github.com/xmindflow/CLMU-Net.
Tissue Classification and Whole-Slide Images Analysis via Modeling of the Tumor Microenvironment and Biological Pathways
Jan 13, 2026Abstract:Automatic integration of whole slide images (WSIs) and gene expression profiles has demonstrated substantial potential in precision clinical diagnosis and cancer progression studies. However, most existing studies focus on individual gene sequences and slide level classification tasks, with limited attention to spatial transcriptomics and patch level applications. To address this limitation, we propose a multimodal network, BioMorphNet, which automatically integrates tissue morphological features and spatial gene expression to support tissue classification and differential gene analysis. For considering morphological features, BioMorphNet constructs a graph to model the relationships between target patches and their neighbors, and adjusts the response strength based on morphological and molecular level similarity, to better characterize the tumor microenvironment. In terms of multimodal interactions, BioMorphNet derives clinical pathway features from spatial transcriptomic data based on a predefined pathway database, serving as a bridge between tissue morphology and gene expression. In addition, a novel learnable pathway module is designed to automatically simulate the biological pathway formation process, providing a complementary representation to existing clinical pathways. Compared with the latest morphology gene multimodal methods, BioMorphNet's average classification metrics improve by 2.67%, 5.48%, and 6.29% for prostate cancer, colorectal cancer, and breast cancer datasets, respectively. BioMorphNet not only classifies tissue categories within WSIs accurately to support tumor localization, but also analyzes differential gene expression between tissue categories based on prediction confidence, contributing to the discovery of potential tumor biomarkers.
Deep Learning-Based Desikan-Killiany Parcellation of the Brain Using Diffusion MRI
Aug 11, 2025



Abstract:Accurate brain parcellation in diffusion MRI (dMRI) space is essential for advanced neuroimaging analyses. However, most existing approaches rely on anatomical MRI for segmentation and inter-modality registration, a process that can introduce errors and limit the versatility of the technique. In this study, we present a novel deep learning-based framework for direct parcellation based on the Desikan-Killiany (DK) atlas using only diffusion MRI data. Our method utilizes a hierarchical, two-stage segmentation network: the first stage performs coarse parcellation into broad brain regions, and the second stage refines the segmentation to delineate more detailed subregions within each coarse category. We conduct an extensive ablation study to evaluate various diffusion-derived parameter maps, identifying an optimal combination of fractional anisotropy, trace, sphericity, and maximum eigenvalue that enhances parellation accuracy. When evaluated on the Human Connectome Project and Consortium for Neuropsychiatric Phenomics datasets, our approach achieves superior Dice Similarity Coefficients compared to existing state-of-the-art models. Additionally, our method demonstrates robust generalization across different image resolutions and acquisition protocols, producing more homogeneous parcellations as measured by the relative standard deviation within regions. This work represents a significant advancement in dMRI-based brain segmentation, providing a precise, reliable, and registration-free solution that is critical for improved structural connectivity and microstructural analyses in both research and clinical applications. The implementation of our method is publicly available on github.com/xmindflow/DKParcellationdMRI.
Spatial Transcriptomics Expression Prediction from Histopathology Based on Cross-Modal Mask Reconstruction and Contrastive Learning
Jun 10, 2025Abstract:Spatial transcriptomics is a technology that captures gene expression levels at different spatial locations, widely used in tumor microenvironment analysis and molecular profiling of histopathology, providing valuable insights into resolving gene expression and clinical diagnosis of cancer. Due to the high cost of data acquisition, large-scale spatial transcriptomics data remain challenging to obtain. In this study, we develop a contrastive learning-based deep learning method to predict spatially resolved gene expression from whole-slide images. Evaluation across six different disease datasets demonstrates that, compared to existing studies, our method improves Pearson Correlation Coefficient (PCC) in the prediction of highly expressed genes, highly variable genes, and marker genes by 6.27%, 6.11%, and 11.26% respectively. Further analysis indicates that our method preserves gene-gene correlations and applies to datasets with limited samples. Additionally, our method exhibits potential in cancer tissue localization based on biomarker expression.
CENet: Context Enhancement Network for Medical Image Segmentation
May 23, 2025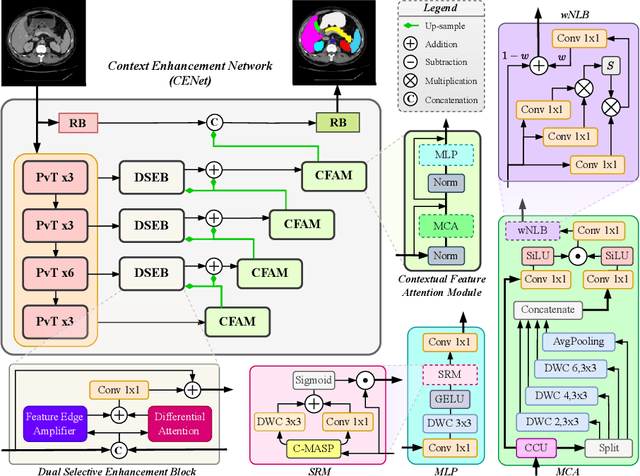
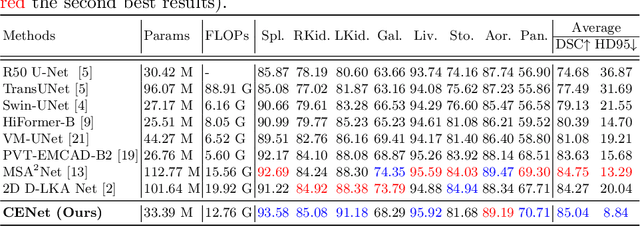
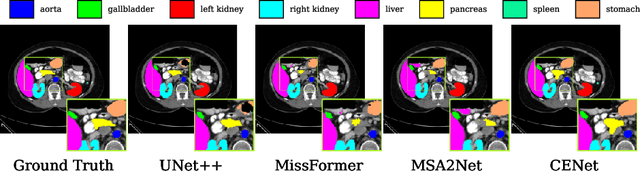
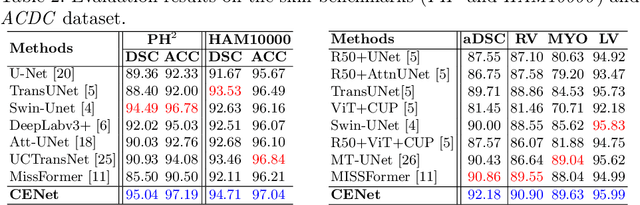
Abstract:Medical image segmentation, particularly in multi-domain scenarios, requires precise preservation of anatomical structures across diverse representations. While deep learning has advanced this field, existing models often struggle with accurate boundary representation, variability in organ morphology, and information loss during downsampling, limiting their accuracy and robustness. To address these challenges, we propose the Context Enhancement Network (CENet), a novel segmentation framework featuring two key innovations. First, the Dual Selective Enhancement Block (DSEB) integrated into skip connections enhances boundary details and improves the detection of smaller organs in a context-aware manner. Second, the Context Feature Attention Module (CFAM) in the decoder employs a multi-scale design to maintain spatial integrity, reduce feature redundancy, and mitigate overly enhanced representations. Extensive evaluations on both radiology and dermoscopic datasets demonstrate that CENet outperforms state-of-the-art (SOTA) methods in multi-organ segmentation and boundary detail preservation, offering a robust and accurate solution for complex medical image analysis tasks. The code is publicly available at https://github.com/xmindflow/cenet.
Attention-based Generative Latent Replay: A Continual Learning Approach for WSI Analysis
May 13, 2025



Abstract:Whole slide image (WSI) classification has emerged as a powerful tool in computational pathology, but remains constrained by domain shifts, e.g., due to different organs, diseases, or institution-specific variations. To address this challenge, we propose an Attention-based Generative Latent Replay Continual Learning framework (AGLR-CL), in a multiple instance learning (MIL) setup for domain incremental WSI classification. Our method employs Gaussian Mixture Models (GMMs) to synthesize WSI representations and patch count distributions, preserving knowledge of past domains without explicitly storing original data. A novel attention-based filtering step focuses on the most salient patch embeddings, ensuring high-quality synthetic samples. This privacy-aware strategy obviates the need for replay buffers and outperforms other buffer-free counterparts while matching the performance of buffer-based solutions. We validate AGLR-CL on clinically relevant biomarker detection and molecular status prediction across multiple public datasets with diverse centers, organs, and patient cohorts. Experimental results confirm its ability to retain prior knowledge and adapt to new domains, offering an effective, privacy-preserving avenue for domain incremental continual learning in WSI classification.
Towards Robust and Generalizable Gerchberg Saxton based Physics Inspired Neural Networks for Computer Generated Holography: A Sensitivity Analysis Framework
Apr 30, 2025Abstract:Computer-generated holography (CGH) enables applications in holographic augmented reality (AR), 3D displays, systems neuroscience, and optical trapping. The fundamental challenge in CGH is solving the inverse problem of phase retrieval from intensity measurements. Physics-inspired neural networks (PINNs), especially Gerchberg-Saxton-based PINNs (GS-PINNs), have advanced phase retrieval capabilities. However, their performance strongly depends on forward models (FMs) and their hyperparameters (FMHs), limiting generalization, complicating benchmarking, and hindering hardware optimization. We present a systematic sensitivity analysis framework based on Saltelli's extension of Sobol's method to quantify FMH impacts on GS-PINN performance. Our analysis demonstrates that SLM pixel-resolution is the primary factor affecting neural network sensitivity, followed by pixel-pitch, propagation distance, and wavelength. Free space propagation forward models demonstrate superior neural network performance compared to Fourier holography, providing enhanced parameterization and generalization. We introduce a composite evaluation metric combining performance consistency, generalization capability, and hyperparameter perturbation resilience, establishing a unified benchmarking standard across CGH configurations. Our research connects physics-inspired deep learning theory with practical CGH implementations through concrete guidelines for forward model selection, neural network architecture, and performance evaluation. Our contributions advance the development of robust, interpretable, and generalizable neural networks for diverse holographic applications, supporting evidence-based decisions in CGH research and implementation.
Modality-Independent Brain Lesion Segmentation with Privacy-aware Continual Learning
Mar 26, 2025



Abstract:Traditional brain lesion segmentation models for multi-modal MRI are typically tailored to specific pathologies, relying on datasets with predefined modalities. Adapting to new MRI modalities or pathologies often requires training separate models, which contrasts with how medical professionals incrementally expand their expertise by learning from diverse datasets over time. Inspired by this human learning process, we propose a unified segmentation model capable of sequentially learning from multiple datasets with varying modalities and pathologies. Our approach leverages a privacy-aware continual learning framework that integrates a mixture-of-experts mechanism and dual knowledge distillation to mitigate catastrophic forgetting while not compromising performance on newly encountered datasets. Extensive experiments across five diverse brain MRI datasets and four dataset sequences demonstrate the effectiveness of our framework in maintaining a single adaptable model, capable of handling varying hospital protocols, imaging modalities, and disease types. Compared to widely used privacy-aware continual learning methods such as LwF, SI, EWC, and MiB, our method achieves an average Dice score improvement of approximately 11%. Our framework represents a significant step toward more versatile and practical brain lesion segmentation models, with implementation available at \href{https://github.com/xmindflow/BrainCL}{GitHub}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge