Maschenka Balkenhol
DALPHIN: Benchmarking Digital Pathology AI Copilots Against Pathologists on an Open Multicentric Dataset
May 05, 2026Abstract:Foundation models with visual question answering capabilities for digital pathology are emerging. Such unprecedented technology requires independent benchmarking to assess its potential in assisting pathologists in routine diagnostics. We created DALPHIN, the first multicentric open benchmark for pathology AI copilots, comprising 1236 images from 300 cases, spanning 130 rare to common diagnoses, 6 countries, and 14 subspecialties. The DALPHIN design and dataset are introduced alongside a human performance benchmark of 31 pathologists from 10 countries with varying expertise. We report results for two general-purpose (GPT-5, Gemini 2.5 Pro) and one pathology-specific copilot (PathChat+) for sequential and independent answer generation. We observed no statistically significant difference from expert-level performance in four of six tasks for PathChat, 2/6 tasks for Gemini, and 1/6 tasks for GPT. DALPHIN is publicly released with sequestered, indirectly accessible ground truth to foster robust and enduring benchmarking. Data, methods, and the evaluation platform are accessible through dalphin.grand-challenge.org.
A Multicentric Dataset for Training and Benchmarking Breast Cancer Segmentation in H&E Slides
Oct 02, 2025


Abstract:Automated semantic segmentation of whole-slide images (WSIs) stained with hematoxylin and eosin (H&E) is essential for large-scale artificial intelligence-based biomarker analysis in breast cancer. However, existing public datasets for breast cancer segmentation lack the morphological diversity needed to support model generalizability and robust biomarker validation across heterogeneous patient cohorts. We introduce BrEast cancEr hisTopathoLogy sEgmentation (BEETLE), a dataset for multiclass semantic segmentation of H&E-stained breast cancer WSIs. It consists of 587 biopsies and resections from three collaborating clinical centers and two public datasets, digitized using seven scanners, and covers all molecular subtypes and histological grades. Using diverse annotation strategies, we collected annotations across four classes - invasive epithelium, non-invasive epithelium, necrosis, and other - with particular focus on morphologies underrepresented in existing datasets, such as ductal carcinoma in situ and dispersed lobular tumor cells. The dataset's diversity and relevance to the rapidly growing field of automated biomarker quantification in breast cancer ensure its high potential for reuse. Finally, we provide a well-curated, multicentric external evaluation set to enable standardized benchmarking of breast cancer segmentation models.
Domain adaptation strategies for cancer-independent detection of lymph node metastases
Jul 13, 2022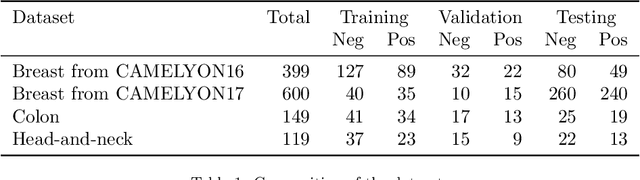
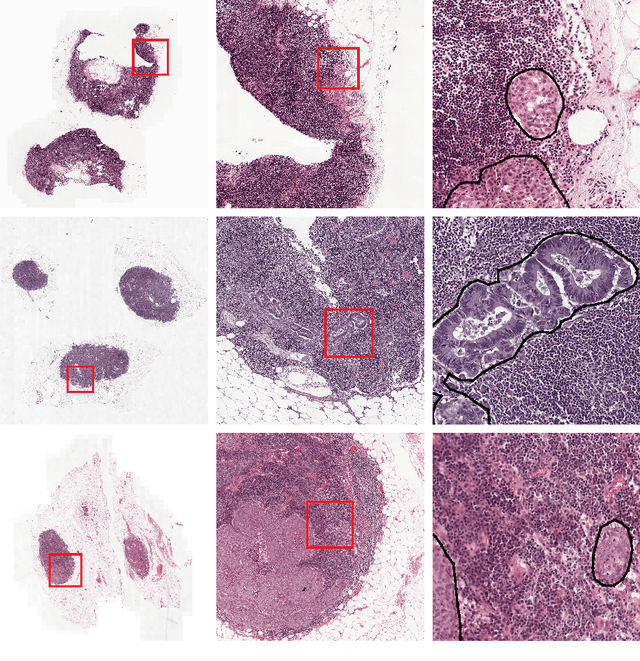
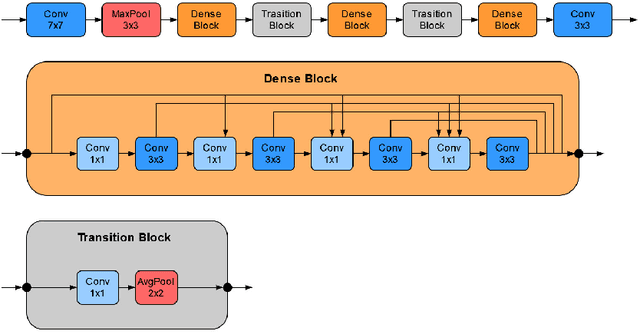
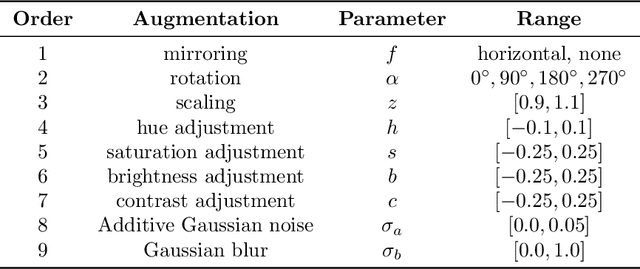
Abstract:Recently, large, high-quality public datasets have led to the development of convolutional neural networks that can detect lymph node metastases of breast cancer at the level of expert pathologists. Many cancers, regardless of the site of origin, can metastasize to lymph nodes. However, collecting and annotating high-volume, high-quality datasets for every cancer type is challenging. In this paper we investigate how to leverage existing high-quality datasets most efficiently in multi-task settings for closely related tasks. Specifically, we will explore different training and domain adaptation strategies, including prevention of catastrophic forgetting, for colon and head-and-neck cancer metastasis detection in lymph nodes. Our results show state-of-the-art performance on both cancer metastasis detection tasks. Furthermore, we show the effectiveness of repeated adaptation of networks from one cancer type to another to obtain multi-task metastasis detection networks. Last, we show that leveraging existing high-quality datasets can significantly boost performance on new target tasks and that catastrophic forgetting can be effectively mitigated using regularization.
Automated Scoring of Nuclear Pleomorphism Spectrum with Pathologist-level Performance in Breast Cancer
Dec 24, 2020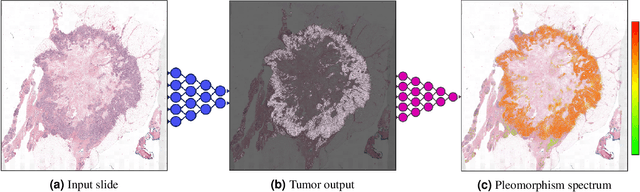
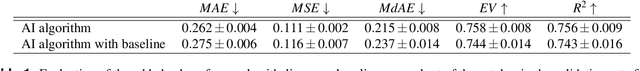
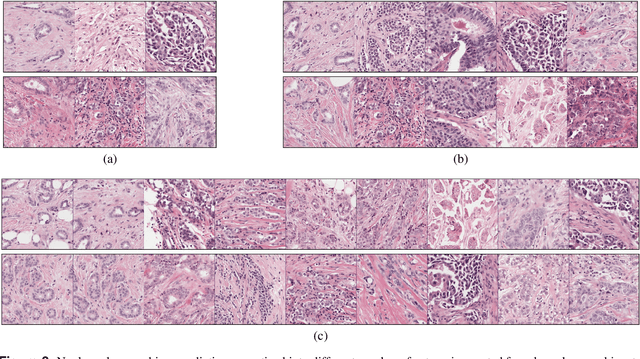
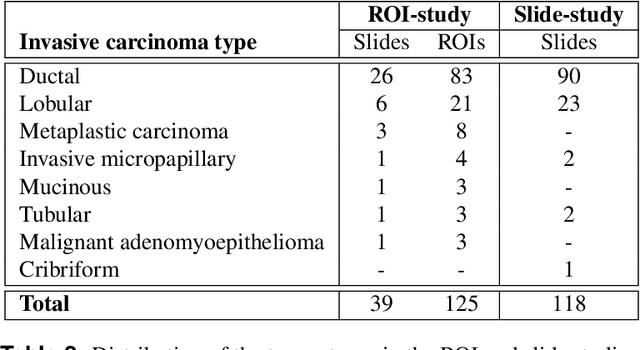
Abstract:Nuclear pleomorphism, defined herein as the extent of abnormalities in the overall appearance of tumor nuclei, is one of the components of the three-tiered breast cancer grading. Given that nuclear pleomorphism reflects a continuous spectrum of variation, we trained a deep neural network on a large variety of tumor regions from the collective knowledge of several pathologists, without constraining the network to the traditional three-category classification. We also motivate an additional approach in which we discuss the additional benefit of normal epithelium as baseline, following the routine clinical practice where pathologists are trained to score nuclear pleomorphism in tumor, having the normal breast epithelium for comparison. In multiple experiments, our fully-automated approach could achieve top pathologist-level performance in select regions of interest as well as at whole slide images, compared to ten and four pathologists, respectively.
HookNet: multi-resolution convolutional neural networks for semantic segmentation in histopathology whole-slide images
Jun 22, 2020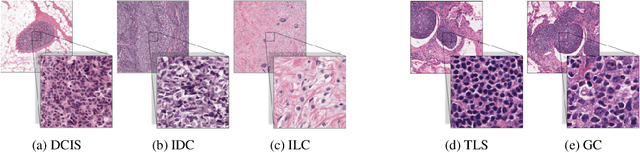
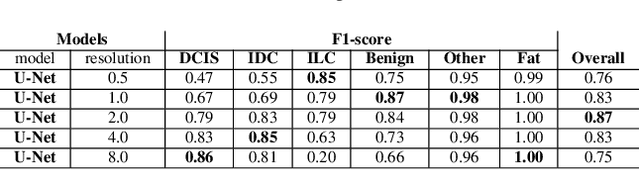
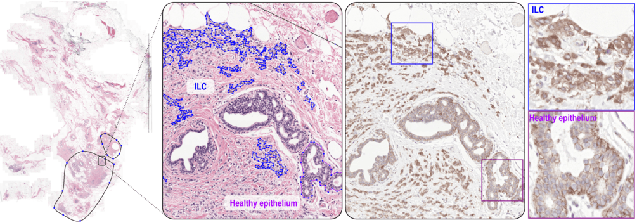
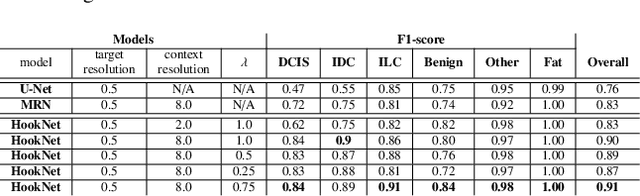
Abstract:We propose HookNet, a semantic segmentation model for histopathology whole-slide images, which combines context and details via multiple branches of encoder-decoder convolutional neural networks. Concentricpatches at multiple resolutions with different fields of view are used to feed different branches of HookNet, and intermediate representations are combined via a hooking mechanism. We describe a framework to design and train HookNet for achieving high-resolution semantic segmentation and introduce constraints to guarantee pixel-wise alignment in feature maps during hooking. We show the advantages of using HookNet in two histopathology image segmentation tasks where tissue type prediction accuracy strongly depends on contextual information, namely (1) multi-class tissue segmentation in breast cancer and, (2) segmentation of tertiary lymphoid structures and germinal centers in lung cancer. Weshow the superiority of HookNet when compared with single-resolution U-Net models working at different resolutions as well as with a recently published multi-resolution model for histopathology image segmentation
Artificial Intelligence Assistance Significantly Improves Gleason Grading of Prostate Biopsies by Pathologists
Feb 11, 2020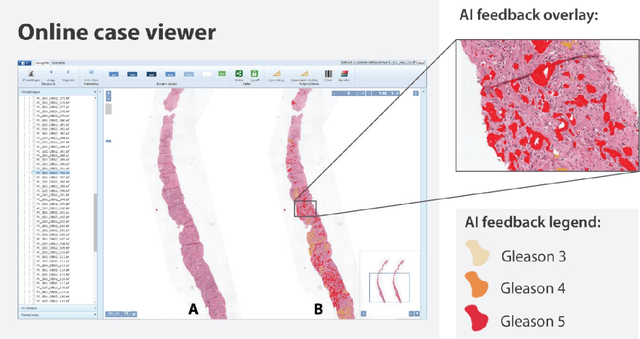
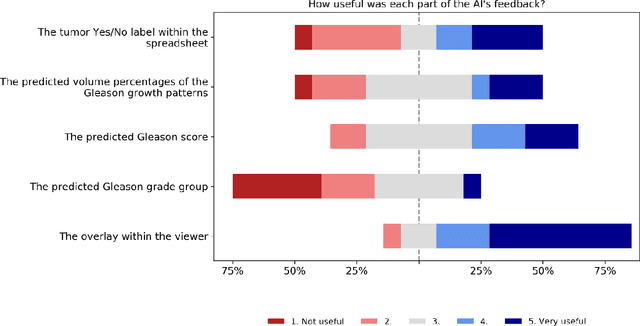
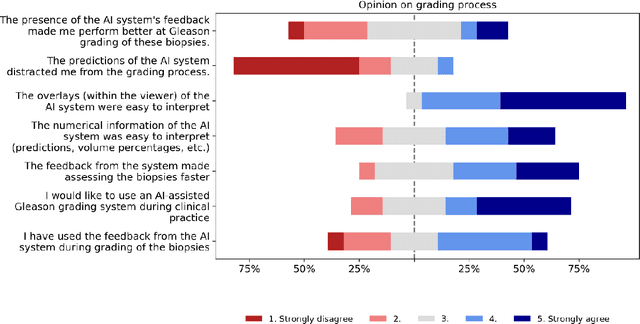
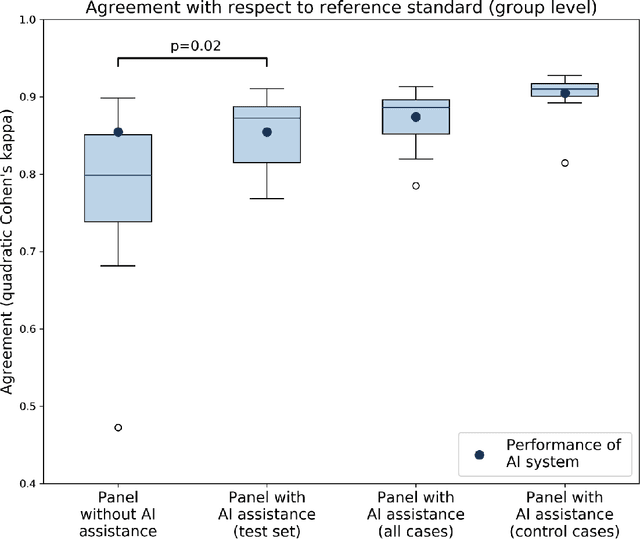
Abstract:While the Gleason score is the most important prognostic marker for prostate cancer patients, it suffers from significant observer variability. Artificial Intelligence (AI) systems, based on deep learning, have proven to achieve pathologist-level performance at Gleason grading. However, the performance of such systems can degrade in the presence of artifacts, foreign tissue, or other anomalies. Pathologists integrating their expertise with feedback from an AI system could result in a synergy that outperforms both the individual pathologist and the system. Despite the hype around AI assistance, existing literature on this topic within the pathology domain is limited. We investigated the value of AI assistance for grading prostate biopsies. A panel of fourteen observers graded 160 biopsies with and without AI assistance. Using AI, the agreement of the panel with an expert reference standard significantly increased (quadratically weighted Cohen's kappa, 0.799 vs 0.872; p=0.018). Our results show the added value of AI systems for Gleason grading, but more importantly, show the benefits of pathologist-AI synergy.
Whole-Slide Mitosis Detection in H&E Breast Histology Using PHH3 as a Reference to Train Distilled Stain-Invariant Convolutional Networks
Aug 17, 2018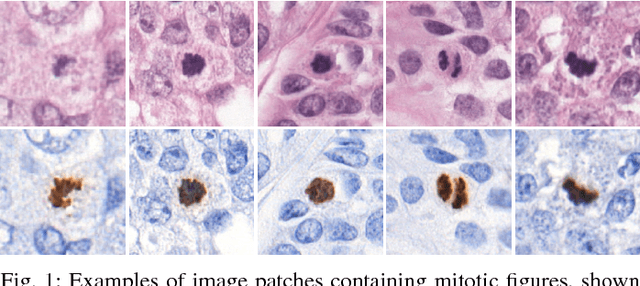
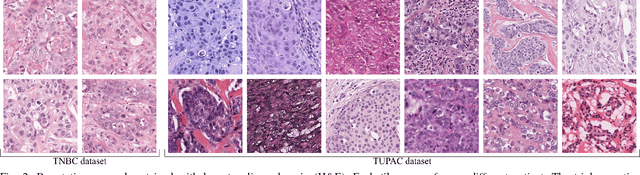
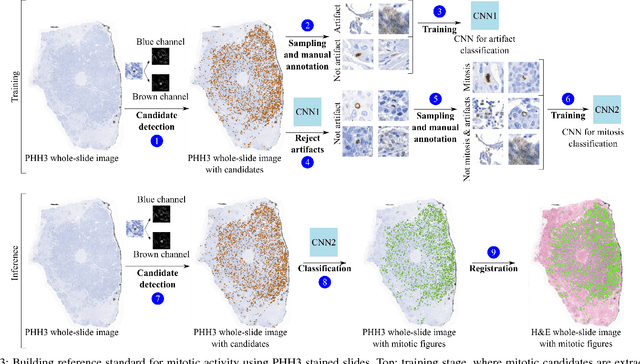
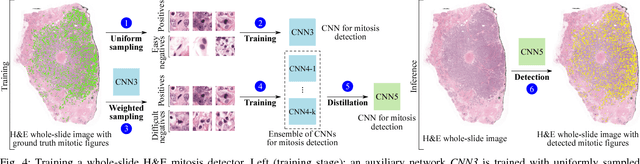
Abstract:Manual counting of mitotic tumor cells in tissue sections constitutes one of the strongest prognostic markers for breast cancer. This procedure, however, is time-consuming and error-prone. We developed a method to automatically detect mitotic figures in breast cancer tissue sections based on convolutional neural networks (CNNs). Application of CNNs to hematoxylin and eosin (H&E) stained histological tissue sections is hampered by: (1) noisy and expensive reference standards established by pathologists, (2) lack of generalization due to staining variation across laboratories, and (3) high computational requirements needed to process gigapixel whole-slide images (WSIs). In this paper, we present a method to train and evaluate CNNs to specifically solve these issues in the context of mitosis detection in breast cancer WSIs. First, by combining image analysis of mitotic activity in phosphohistone-H3 (PHH3) restained slides and registration, we built a reference standard for mitosis detection in entire H&E WSIs requiring minimal manual annotation effort. Second, we designed a data augmentation strategy that creates diverse and realistic H&E stain variations by modifying the hematoxylin and eosin color channels directly. Using it during training combined with network ensembling resulted in a stain invariant mitosis detector. Third, we applied knowledge distillation to reduce the computational requirements of the mitosis detection ensemble with a negligible loss of performance. The system was trained in a single-center cohort and evaluated in an independent multicenter cohort from The Cancer Genome Atlas on the three tasks of the Tumor Proliferation Assessment Challenge (TUPAC). We obtained a performance within the top-3 best methods for most of the tasks of the challenge.
Context-aware stacked convolutional neural networks for classification of breast carcinomas in whole-slide histopathology images
May 10, 2017Abstract:Automated classification of histopathological whole-slide images (WSI) of breast tissue requires analysis at very high resolutions with a large contextual area. In this paper, we present context-aware stacked convolutional neural networks (CNN) for classification of breast WSIs into normal/benign, ductal carcinoma in situ (DCIS), and invasive ductal carcinoma (IDC). We first train a CNN using high pixel resolution patches to capture cellular level information. The feature responses generated by this model are then fed as input to a second CNN, stacked on top of the first. Training of this stacked architecture with large input patches enables learning of fine-grained (cellular) details and global interdependence of tissue structures. Our system is trained and evaluated on a dataset containing 221 WSIs of H&E stained breast tissue specimens. The system achieves an AUC of 0.962 for the binary classification of non-malignant and malignant slides and obtains a three class accuracy of 81.3% for classification of WSIs into normal/benign, DCIS, and IDC, demonstrating its potentials for routine diagnostics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge