James A. Meakin
and on behalf of the UNICORN consortium
Designing UNICORN: a Unified Benchmark for Imaging in Computational Pathology, Radiology, and Natural Language
Mar 03, 2026Abstract:Medical foundation models show promise to learn broadly generalizable features from large, diverse datasets. This could be the base for reliable cross-modality generalization and rapid adaptation to new, task-specific goals, with only a few task-specific examples. Yet, evidence for this is limited by the lack of public, standardized, and reproducible evaluation frameworks, as existing public benchmarks are often fragmented across task-, organ-, or modality-specific settings, limiting assessment of cross-task generalization. We introduce UNICORN, a public benchmark designed to systematically evaluate medical foundation models under a unified protocol. To isolate representation quality, we built the benchmark on a novel two-step framework that decouples model inference from task-specific evaluation based on standardized few-shot adaptation. As a central design choice, we constructed indirectly accessible sequestered test sets derived from clinically relevant cohorts, along with standardized evaluation code and a submission interface on an open benchmarking platform. Performance is aggregated into a single UNICORN Score, a new metric that we introduce to support direct comparison of foundation models across diverse medical domains, modalities, and task types. The UNICORN test dataset includes data from more than 2,400 patients, including over 3,700 vision cases and over 2,400 clinical reports collected from 17 institutions across eight countries. The benchmark spans eight anatomical regions and four imaging modalities. Both task-specific and aggregated leaderboards enable accessible, standardized, and reproducible evaluation. By standardizing multi-task, multi-modality assessment, UNICORN establishes a foundation for reproducible benchmarking of medical foundation models. Data, baseline methods, and the evaluation platform are publicly available via unicorn.grand-challenge.org.
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024



Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
The STOIC2021 COVID-19 AI challenge: applying reusable training methodologies to private data
Jun 25, 2023Abstract:Challenges drive the state-of-the-art of automated medical image analysis. The quantity of public training data that they provide can limit the performance of their solutions. Public access to the training methodology for these solutions remains absent. This study implements the Type Three (T3) challenge format, which allows for training solutions on private data and guarantees reusable training methodologies. With T3, challenge organizers train a codebase provided by the participants on sequestered training data. T3 was implemented in the STOIC2021 challenge, with the goal of predicting from a computed tomography (CT) scan whether subjects had a severe COVID-19 infection, defined as intubation or death within one month. STOIC2021 consisted of a Qualification phase, where participants developed challenge solutions using 2000 publicly available CT scans, and a Final phase, where participants submitted their training methodologies with which solutions were trained on CT scans of 9724 subjects. The organizers successfully trained six of the eight Final phase submissions. The submitted codebases for training and running inference were released publicly. The winning solution obtained an area under the receiver operating characteristic curve for discerning between severe and non-severe COVID-19 of 0.815. The Final phase solutions of all finalists improved upon their Qualification phase solutions.HSUXJM-TNZF9CHSUXJM-TNZF9C
The Medical Segmentation Decathlon
Jun 10, 2021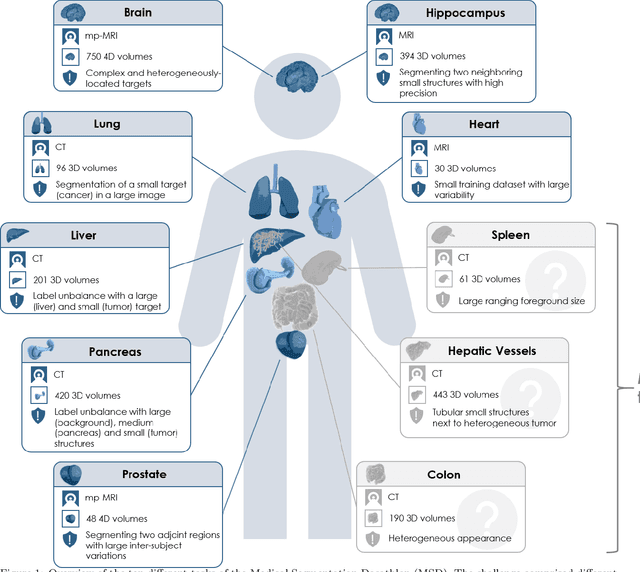
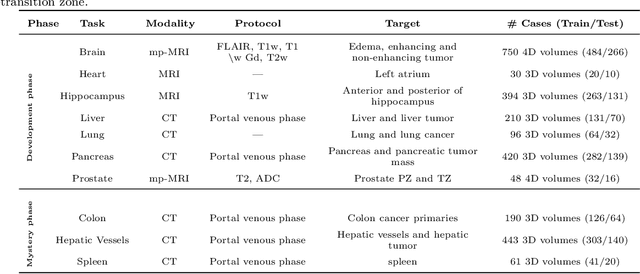
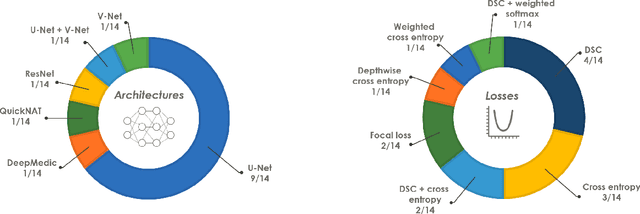
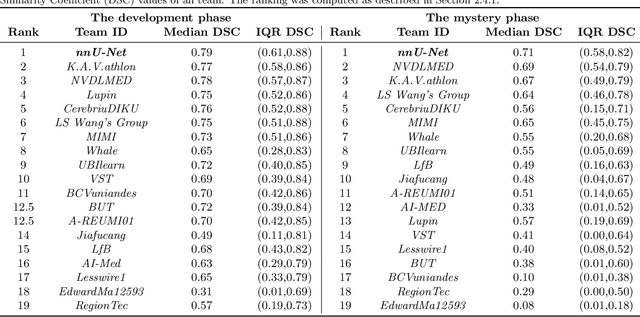
Abstract:International challenges have become the de facto standard for comparative assessment of image analysis algorithms given a specific task. Segmentation is so far the most widely investigated medical image processing task, but the various segmentation challenges have typically been organized in isolation, such that algorithm development was driven by the need to tackle a single specific clinical problem. We hypothesized that a method capable of performing well on multiple tasks will generalize well to a previously unseen task and potentially outperform a custom-designed solution. To investigate the hypothesis, we organized the Medical Segmentation Decathlon (MSD) - a biomedical image analysis challenge, in which algorithms compete in a multitude of both tasks and modalities. The underlying data set was designed to explore the axis of difficulties typically encountered when dealing with medical images, such as small data sets, unbalanced labels, multi-site data and small objects. The MSD challenge confirmed that algorithms with a consistent good performance on a set of tasks preserved their good average performance on a different set of previously unseen tasks. Moreover, by monitoring the MSD winner for two years, we found that this algorithm continued generalizing well to a wide range of other clinical problems, further confirming our hypothesis. Three main conclusions can be drawn from this study: (1) state-of-the-art image segmentation algorithms are mature, accurate, and generalize well when retrained on unseen tasks; (2) consistent algorithmic performance across multiple tasks is a strong surrogate of algorithmic generalizability; (3) the training of accurate AI segmentation models is now commoditized to non AI experts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge