Ziheng Zhang
WARDEN: Endangered Indigenous Language Transcription and Translation with 6 Hours of Training Data
May 13, 2026Abstract:This paper introduces WARDEN, an early language model system capable of transcribing and translating Wardaman, an endangered Australian indigenous language into English. The significant challenge we face is the lack of large-scale training data: in fact, we only have 6 hours of annotated audio. Therefore, while it is common practice to train a single model for transcription and translation using large datasets (like English to French), this practice is no longer viable in the Wardaman to English context. To tackle the low-resource challenge, we design WARDEN to have separate transcription and translation models: WARDEN first turns a Wardaman audio input into phonemic transcription, and then the transcription into English translation. Further, we propose two useful techniques to enhance performance. For transcription, we initialize the Wardaman token from Sundanese, a language that shares similar phonemes with Wardaman, to accelerate fine-tuning of the transcription model. For translation, we compile a Wardaman-English dictionary from expert annotations, and provide this domain-specific knowledge to a large language model (LLM) to reason and decide the final output. We empirically demonstrate that this two-stage design works better than data-hungry unified approaches in extremely low data settings. Using a mere 6 hours of annotated data, WARDEN outperforms larger open-source and proprietary models and establishes a strong baseline. Data and code are available.
GS-Playground: A High-Throughput Photorealistic Simulator for Vision-Informed Robot Learning
Apr 28, 2026Abstract:Embodied AI research is undergoing a shift toward vision-centric perceptual paradigms. While massively parallel simulators have catalyzed breakthroughs in proprioception-based locomotion, their potential remains largely untapped for vision-informed tasks due to the prohibitive computational overhead of large-scale photorealistic rendering. Furthermore, the creation of simulation-ready 3D assets heavily relies on labor-intensive manual modeling, while the significant sim-to-real physical gap hinders the transfer of contact-rich manipulation policies. To address these bottlenecks, we propose GS-Playground, a multi-modal simulation framework designed to accelerate end-to-end perceptual learning. We develop a novel high-performance parallel physics engine, specifically designed to integrate with a batch 3D Gaussian Splatting (3DGS) rendering pipeline to ensure high-fidelity synchronization. Our system achieves a breakthrough throughput of 10^4 FPS at 640x480 resolution, significantly lowering the barrier for large-scale visual RL. Additionally, we introduce an automated Real2Sim workflow that reconstructs photorealistic, physically consistent, and memory-efficient environments, streamlining the generation of complex simulation-ready scenes. Extensive experiments on locomotion, navigation, and manipulation demonstrate that GS-Playground effectively bridges the perceptual and physical gaps across diverse embodied tasks. Project homepage: https://gsplayground.github.io.
Agentization of Digital Assets for the Agentic Web: Concepts, Techniques, and Benchmark
Apr 05, 2026Abstract:Agentic Web, as a new paradigm that redefines the internet through autonomous, goal-driven interactions, plays an important role in group intelligence. As the foundational semantic primitives of the Agentic Web, digital assets encapsulate interactive web elements into agents, which expand the capacities and coverage of agents in agentic web. The lack of automated methodologies for agent generation limits the wider usage of digital assets and the advancement of the Agentic Web. In this paper, we first formalize these challenges by strictly defining the A2A-Agentization process, decomposing it into critical stages and identifying key technical hurdles on top of the A2A protocol. Based on this framework, we develop an Agentization Agent to agentize digital assets for the Agentic Web. To rigorously evaluate this capability, we propose A2A-Agentization Bench, the first benchmark explicitly designed to evaluate agentization quality in terms of fidelity and interoperability. Our experiments demonstrate that our approach effectively activates the functional capabilities of digital assets and enables interoperable A2A multi-agent collaboration. We believe this work will further facilitate scalable and standardized integration of digital assets into the Agentic Web ecosystem.
MMSpec: Benchmarking Speculative Decoding for Vision-Language Models
Mar 16, 2026Abstract:Vision-language models (VLMs) achieve strong performance on multimodal tasks but suffer from high inference latency due to large model sizes and long multimodal contexts. Speculative decoding has recently emerged as an effective acceleration technique, yet its behavior in VLMs remains insufficiently understood. We introduce MMSpec, the first benchmark for evaluating speculative decoding in vision-language models. MMSpec contains 600 multimodal samples across six task categories and integrates ten representative speculative decoding algorithms under a unified evaluation framework. Our study reveals three key findings: (1) methods designed for text-only LLMs degrade in multimodal scenarios, (2) vision awareness becomes increasingly important at larger batch sizes, and (3) throughput speedup alone does not reliably reflect latency performance. Motivated by these findings, we propose ViSkip, a plug-and-play speculative decoding method that dynamically adapts speculation to vision tokens and achieves state-of-the-art performance.
DM0: An Embodied-Native Vision-Language-Action Model towards Physical AI
Feb 16, 2026Abstract:Moving beyond the traditional paradigm of adapting internet-pretrained models to physical tasks, we present DM0, an Embodied-Native Vision-Language-Action (VLA) framework designed for Physical AI. Unlike approaches that treat physical grounding as a fine-tuning afterthought, DM0 unifies embodied manipulation and navigation by learning from heterogeneous data sources from the onset. Our methodology follows a comprehensive three-stage pipeline: Pretraining, Mid-Training, and Post-Training. First, we conduct large-scale unified pretraining on the Vision-Language Model (VLM) using diverse corpora--seamlessly integrating web text, autonomous driving scenarios, and embodied interaction logs-to jointly acquire semantic knowledge and physical priors. Subsequently, we build a flow-matching action expert atop the VLM. To reconcile high-level reasoning with low-level control, DM0 employs a hybrid training strategy: for embodied data, gradients from the action expert are not backpropagated to the VLM to preserve generalized representations, while the VLM remains trainable on non-embodied data. Furthermore, we introduce an Embodied Spatial Scaffolding strategy to construct spatial Chain-of-Thought (CoT) reasoning, effectively constraining the action solution space. Experiments on the RoboChallenge benchmark demonstrate that DM0 achieves state-of-the-art performance in both Specialist and Generalist settings on Table30.
Intelligent Reflecting Surfaces for Integrated Sensing and Communications: A Survey
Nov 14, 2025



Abstract:The rapid development of sixth-generation (6G) wireless networks requires seamless integration of communication and sensing to support ubiquitous intelligence and real-time, high-reliability applications. Integrated sensing and communication (ISAC) has emerged as a key solution for achieving this convergence, offering joint utilization of spectral, hardware, and computing resources. However, realizing high-performance ISAC remains challenging due to environmental line-of-sight (LoS) blockage, limited spatial resolution, and the inherent coverage asymmetry and resource coupling between sensing and communication. Intelligent reflecting surfaces (IRSs), featuring low-cost, energy-efficient, and programmable electromagnetic reconfiguration, provide a promising solution to overcome these limitations. This article presents a comprehensive overview of IRS-aided wireless sensing and ISAC technologies, including IRS architectures, target detection and estimation techniques, beamforming designs, and performance metrics. It further explores IRS-enabled new opportunities for more efficient performance balancing, coexistence, and networking in ISAC systems, focuses on current design bottlenecks, and outlines future research directions. This article aims to offer a unified design framework that guides the development of practical and scalable IRS-aided ISAC systems for the next-generation wireless network.
Reconfigurable Airspace: Synergizing Movable Antenna and Intelligent Surface for Low-Altitude ISAC Networks
Nov 13, 2025Abstract:Low-altitude unmanned aerial vehicle (UAV) networks are integral to future 6G integrated sensing and communication (ISAC) systems. However, their deployment is hindered by challenges stemming from high mobility of UAVs, complex propagation environments, and the inherent trade-offs between coexisting sensing and communication functions. This article proposes a novel framework that leverages movable antennas (MAs) and intelligent reflecting surfaces (IRSs) as dual enablers to overcome these limitations. MAs, through active transceiver reconfiguration, and IRSs, via passive channel reconstruction, can work in synergy to significantly enhance system performance. Our analysis first elaborates on the fundamental gains offered by MAs and IRSs, and provides simulation results that validate the immense potential of the MA-IRS-enabled ISAC architecture. Two core UAV deployment scenarios are then investigated: (i) UAVs as ISAC users, where we focus on achieving high-precision tracking and aerial safety, and (ii) UAVs as aerial network nodes, where we address robust design and complex coupled resource optimization. Finally, key technical challenges and research opportunities are identified and analyzed for each scenario, charting a clear course for the future design of advanced low-altitude ISAC networks.
Why Do Open-Source LLMs Struggle with Data Analysis? A Systematic Empirical Study
Jun 24, 2025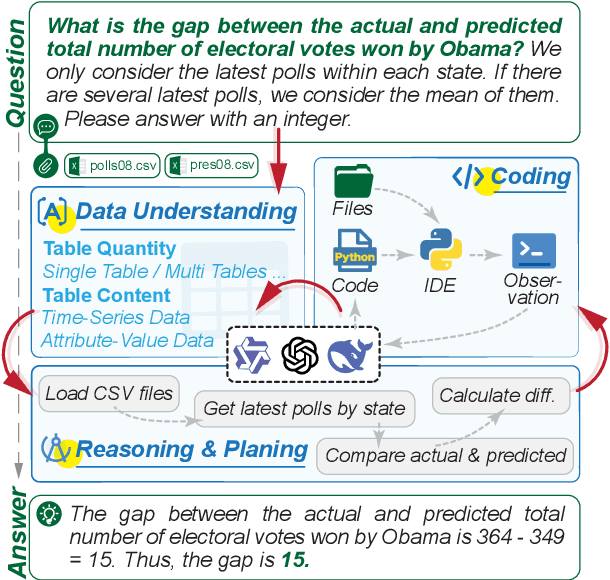
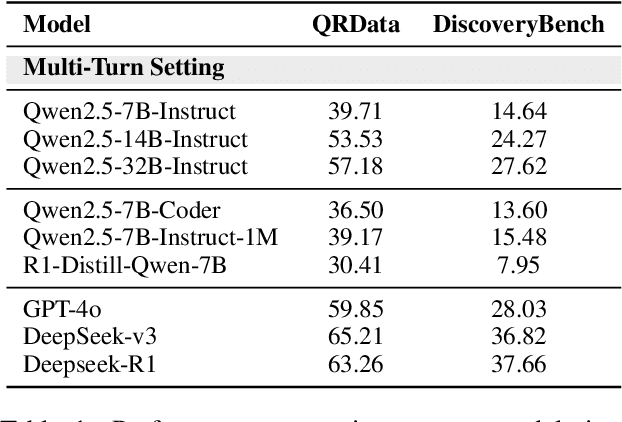
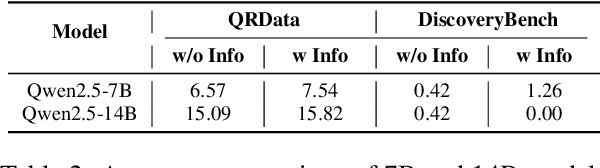
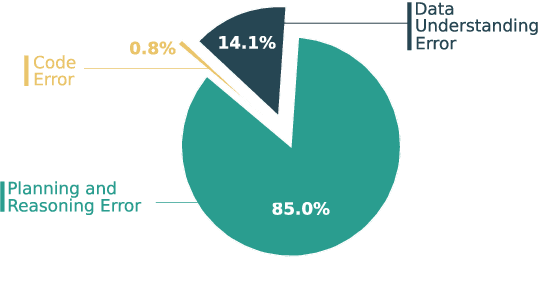
Abstract:Large Language Models (LLMs) hold promise in automating data analysis tasks, yet open-source models face significant limitations in these kinds of reasoning-intensive scenarios. In this work, we investigate strategies to enhance the data analysis capabilities of open-source LLMs. By curating a seed dataset of diverse, realistic scenarios, we evaluate models across three dimensions: data understanding, code generation, and strategic planning. Our analysis reveals three key findings: (1) Strategic planning quality serves as the primary determinant of model performance; (2) Interaction design and task complexity significantly influence reasoning capabilities; (3) Data quality demonstrates a greater impact than diversity in achieving optimal performance. We leverage these insights to develop a data synthesis methodology, demonstrating significant improvements in open-source LLMs' analytical reasoning capabilities.
PADriver: Towards Personalized Autonomous Driving
May 08, 2025Abstract:In this paper, we propose PADriver, a novel closed-loop framework for personalized autonomous driving (PAD). Built upon Multi-modal Large Language Model (MLLM), PADriver takes streaming frames and personalized textual prompts as inputs. It autoaggressively performs scene understanding, danger level estimation and action decision. The predicted danger level reflects the risk of the potential action and provides an explicit reference for the final action, which corresponds to the preset personalized prompt. Moreover, we construct a closed-loop benchmark named PAD-Highway based on Highway-Env simulator to comprehensively evaluate the decision performance under traffic rules. The dataset contains 250 hours videos with high-quality annotation to facilitate the development of PAD behavior analysis. Experimental results on the constructed benchmark show that PADriver outperforms state-of-the-art approaches on different evaluation metrics, and enables various driving modes.
Not All Parameters Matter: Masking Diffusion Models for Enhancing Generation Ability
May 06, 2025



Abstract:The diffusion models, in early stages focus on constructing basic image structures, while the refined details, including local features and textures, are generated in later stages. Thus the same network layers are forced to learn both structural and textural information simultaneously, significantly differing from the traditional deep learning architectures (e.g., ResNet or GANs) which captures or generates the image semantic information at different layers. This difference inspires us to explore the time-wise diffusion models. We initially investigate the key contributions of the U-Net parameters to the denoising process and identify that properly zeroing out certain parameters (including large parameters) contributes to denoising, substantially improving the generation quality on the fly. Capitalizing on this discovery, we propose a simple yet effective method-termed ``MaskUNet''- that enhances generation quality with negligible parameter numbers. Our method fully leverages timestep- and sample-dependent effective U-Net parameters. To optimize MaskUNet, we offer two fine-tuning strategies: a training-based approach and a training-free approach, including tailored networks and optimization functions. In zero-shot inference on the COCO dataset, MaskUNet achieves the best FID score and further demonstrates its effectiveness in downstream task evaluations. Project page: https://gudaochangsheng.github.io/MaskUnet-Page/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge