Wei-Lun Chao
The Ohio State University
A Study of Failure Modes in Two-Stage Human-Object Interaction Detection
Apr 15, 2026Abstract:Human-object interaction (HOI) detection aims to detect interactions between humans and objects in images. While recent advances have improved performance on existing benchmarks, their evaluations mainly focus on overall prediction accuracy and provide limited insight into the underlying causes of model failures. In particular, modern models often struggle in complex scenes involving multiple people and rare interaction combinations. In this work, we present a study to better understand the failure modes of two-stage HOI models, which form the basis of many current HOI detection approaches. Rather than constructing a large-scale benchmark, we instead decompose HOI detection into multiple interpretable perspectives and analyze model behavior across these dimensions to study different types of failure patterns. We curate a subset of images from an existing HOI dataset organized by human-object-interaction configurations (e.g., multi-person interactions and object sharing), and analyze model behavior under these configurations to examine different failure modes. This design allows us to analyze how these HOI models behave under different scene compositions and why their predictions fail. Importantly, high overall benchmark performance does not necessarily reflect robust visual reasoning about human-object relationships. We hope that this study can provide useful insights into the limitations of HOI models and offer observations for future research in this area.
GS-Surrogate: Deformable Gaussian Splatting for Parameter Space Exploration of Ensemble Simulations
Apr 07, 2026Abstract:Exploring ensemble simulations is increasingly important across many scientific domains. However, supporting flexible post-hoc exploration remains challenging due to the trade-off between storing the expensive raw data and flexibly adjusting visualization settings. Existing visualization surrogate models have improved this workflow, but they either operate in image space without an explicit 3D representation or rely on neural radiance fields that are computationally expensive for interactive exploration and encode all parameter-driven variations within a single implicit field. In this work, we introduce GS-Surrogate, a deformable Gaussian Splatting-based visualization surrogate for parameter-space exploration. Our method first constructs a canonical Gaussian field as a base 3D representation and adapts it through sequential parameter-conditioned deformations. By separating simulation-related variations from visualization-specific changes, this explicit formulation enables efficient and controllable adaptation to different visualization tasks, such as isosurface extraction and transfer function editing. We evaluate our framework on a range of simulation datasets, demonstrating that GS-Surrogate enables real-time and flexible exploration across both simulation and visualization parameter spaces.
TaxaAdapter: Vision Taxonomy Models are Key to Fine-grained Image Generation over the Tree of Life
Mar 27, 2026Abstract:Accurately generating images across the Tree of Life is difficult: there are over 10M distinct species on Earth, many of which differ only by subtle visual traits. Despite the remarkable progress in text-to-image synthesis, existing models often fail to capture the fine-grained visual cues that define species identity, even when their outputs appear photo-realistic. To this end, we propose TaxaAdapter, a simple and lightweight approach that incorporates Vision Taxonomy Models (VTMs) such as BioCLIP to guide fine-grained species generation. Our method injects VTM embeddings into a frozen text-to-image diffusion model, improving species-level fidelity while preserving flexible text control over attributes such as pose, style, and background. Extensive experiments demonstrate that TaxaAdapter consistently improves morphology fidelity and species-identity accuracy over strong baselines, with a cleaner architecture and training recipe. To better evaluate these improvements, we also introduce a multimodal Large Language Model-based metric that summarizes trait-level descriptions from generated and real images, providing a more interpretable measure of morphological consistency. Beyond this, we observe that TaxaAdapter exhibits strong generalization capabilities, enabling species synthesis in challenging regimes such as few-shot species with only a handful of training images and even species unseen during training. Overall, our results highlight that VTMs are a key ingredient for scalable, fine-grained species generation.
Lessons and Open Questions from a Unified Study of Camera-Trap Species Recognition Over Time
Mar 20, 2026Abstract:Camera traps are vital for large-scale biodiversity monitoring, yet accurate automated analysis remains challenging due to diverse deployment environments. While the computer vision community has mostly framed this challenge as cross-domain generalization, this perspective overlooks a primary challenge faced by ecological practitioners: maintaining reliable recognition at the fixed site over time, where the dynamic nature of ecosystems introduces profound temporal shifts in both background and animal distributions. To bridge this gap, we present the first unified study of camera-trap species recognition over time. We introduce a realistic benchmark comprising 546 camera traps with a streaming protocol that evaluates models over chronologically ordered intervals. Our end-user-centric study yields four key findings. (1) Biological foundation models (e.g., BioCLIP 2) underperform at numerous sites even in initial intervals, underscoring the necessity of site-specific adaptation. (2) Adaptation is challenging under realistic evaluation: when models are updated using past data and evaluated on future intervals (mirrors real deployment lifecycles), naive adaptation can even degrade below zero-shot performance. (3) We identify two drivers of this difficulty: severe class imbalance and pronounced temporal shift in both species distribution and backgrounds between consecutive intervals. (4) We find that effective integration of model-update and post-processing techniques can largely improve accuracy, though a gap from the upper bounds remains. Finally, we highlight critical open questions, such as predicting when zero-shot models will succeed at a new site and determining whether/when model updates are necessary. Our benchmark and analysis provide actionable deployment guidelines for ecological practitioners while establishing new directions for future research in vision and machine learning.
When the City Teaches the Car: Label-Free 3D Perception from Infrastructure
Mar 17, 2026Abstract:Building robust 3D perception for self-driving still relies heavily on large-scale data collection and manual annotation, yet this paradigm becomes impractical as deployment expands across diverse cities and regions. Meanwhile, modern cities are increasingly instrumented with roadside units (RSUs), static sensors deployed along roads and at intersections to monitor traffic. This raises a natural question: can the city itself help train the vehicle? We propose infrastructure-taught, label-free 3D perception, a paradigm in which RSUs act as stationary, unsupervised teachers for ego vehicles. Leveraging their fixed viewpoints and repeated observations, RSUs learn local 3D detectors from unlabeled data and broadcast predictions to passing vehicles, which are aggregated as pseudo-label supervision for training a standalone ego detector. The resulting model requires no infrastructure or communication at test time. We instantiate this idea as a fully label-free three-stage pipeline and conduct a concept-and-feasibility study in a CARLA-based multi-agent environment. With CenterPoint, our pipeline achieves 82.3% AP for detecting vehicles, compared to a fully supervised ego upper bound of 94.4%. We further systematically analyze each stage, evaluate its scalability, and demonstrate complementarity with existing ego-centric label-free methods. Together, these results suggest that city infrastructure itself can potentially provide a scalable supervisory signal for autonomous vehicles, positioning infrastructure-taught learning as a promising orthogonal paradigm for reducing annotation cost in 3D perception.
Revisiting Model Stitching In the Foundation Model Era
Mar 16, 2026Abstract:Model stitching, connecting early layers of one model (source) to later layers of another (target) via a light stitch layer, has served as a probe of representational compatibility. Prior work finds that models trained on the same dataset remain stitchable (negligible accuracy drop) despite different initializations or objectives. We revisit stitching for Vision Foundation Models (VFMs) that vary in objectives, data, and modality mix (e.g., CLIP, DINOv2, SigLIP 2) and ask: Are heterogeneous VFMs stitchable? We introduce a systematic protocol spanning the stitch points, stitch layer families, training losses, and downstream tasks. Three findings emerge. (1) Stitch layer training matters: conventional approaches that match the intermediate features at the stitch point or optimize the task loss end-to-end struggle to retain accuracy, especially at shallow stitch points. (2) With a simple feature-matching loss at the target model's penultimate layer, heterogeneous VFMs become reliably stitchable across vision tasks. (3) For deep stitch points, the stitched model can surpass either constituent model at only a small inference overhead (for the stitch layer). Building on these findings, we further propose the VFM Stitch Tree (VST), which shares early layers across VFMs while retaining their later layers, yielding a controllable accuracy-latency trade-off for multimodal LLMs that often leverage multiple VFMs. Taken together, our study elevates stitching from a diagnostic probe to a practical recipe for integrating complementary VFM strengths and pinpointing where their representations align or diverge.
On the Feasibility and Opportunity of Autoregressive 3D Object Detection
Mar 09, 2026Abstract:LiDAR-based 3D object detectors typically rely on proposal heads with hand-crafted components like anchor assignment and non-maximum suppression (NMS), complicating training and limiting extensibility. We present AutoReg3D, an autoregressive 3D detector that casts detection as sequence generation. Given point-cloud features, AutoReg3D emits objects in a range-causal (near-to-far) order and encodes each object as a short, discrete-token sequence consisting of its center, size, orientation, velocity, and class. This near-to-far ordering mirrors LiDAR geometry--near objects occlude far ones but not vice versa--enabling straightforward teacher forcing during training and autoregressive decoding at test time. AutoReg3D is compatible across diverse point-cloud or backbones and attains competitive nuScenes performance without anchors or NMS. Beyond parity, the sequential formulation unlocks language-model advances for 3D perception, including GRPO-style reinforcement learning for task-aligned objectives. These results position autoregressive decoding as a viable, flexible alternative for LiDAR-based detection and open a path to importing modern sequence-modeling tools into 3D perception.
Finer-Personalization Rank: Fine-Grained Retrieval Examines Identity Preservation for Personalized Generation
Dec 22, 2025



Abstract:The rise of personalized generative models raises a central question: how should we evaluate identity preservation? Given a reference image (e.g., one's pet), we expect the generated image to retain precise details attached to the subject's identity. However, current generative evaluation metrics emphasize the overall semantic similarity between the reference and the output, and overlook these fine-grained discriminative details. We introduce Finer-Personalization Rank, an evaluation protocol tailored to identity preservation. Instead of pairwise similarity, Finer-Personalization Rank adopts a ranking view: it treats each generated image as a query against an identity-labeled gallery consisting of visually similar real images. Retrieval metrics (e.g., mean average precision) measure performance, where higher scores indicate that identity-specific details (e.g., a distinctive head spot) are preserved. We assess identity at multiple granularities -- from fine-grained categories (e.g., bird species, car models) to individual instances (e.g., re-identification). Across CUB, Stanford Cars, and animal Re-ID benchmarks, Finer-Personalization Rank more faithfully reflects identity retention than semantic-only metrics and reveals substantial identity drift in several popular personalization methods. These results position the gallery-based protocol as a principled and practical evaluation for personalized generation.
Continual Unlearning for Text-to-Image Diffusion Models: A Regularization Perspective
Nov 11, 2025Abstract:Machine unlearning--the ability to remove designated concepts from a pre-trained model--has advanced rapidly, particularly for text-to-image diffusion models. However, existing methods typically assume that unlearning requests arrive all at once, whereas in practice they often arrive sequentially. We present the first systematic study of continual unlearning in text-to-image diffusion models and show that popular unlearning methods suffer from rapid utility collapse: after only a few requests, models forget retained knowledge and generate degraded images. We trace this failure to cumulative parameter drift from the pre-training weights and argue that regularization is crucial to addressing it. To this end, we study a suite of add-on regularizers that (1) mitigate drift and (2) remain compatible with existing unlearning methods. Beyond generic regularizers, we show that semantic awareness is essential for preserving concepts close to the unlearning target, and propose a gradient-projection method that constrains parameter drift orthogonal to their subspace. This substantially improves continual unlearning performance and is complementary to other regularizers for further gains. Taken together, our study establishes continual unlearning as a fundamental challenge in text-to-image generation and provides insights, baselines, and open directions for advancing safe and accountable generative AI.
AVA-Bench: Atomic Visual Ability Benchmark for Vision Foundation Models
Jun 10, 2025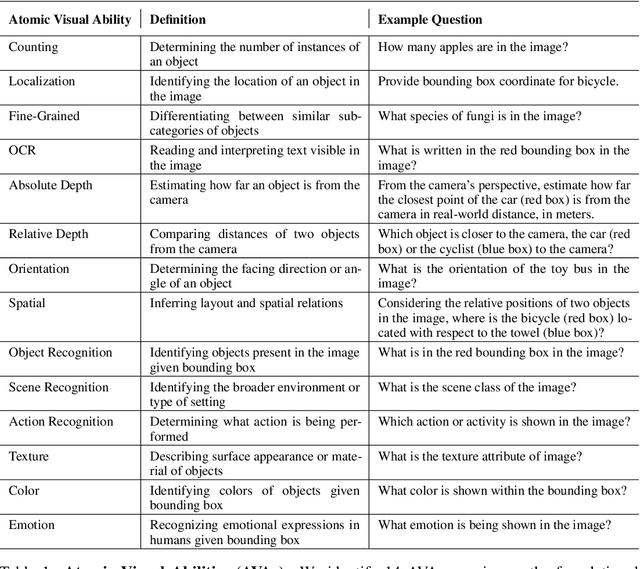
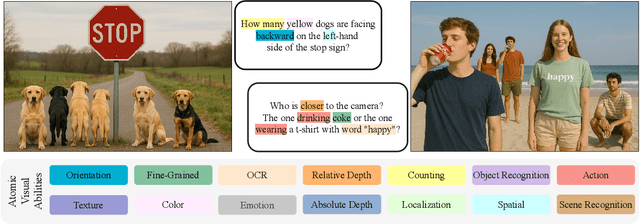
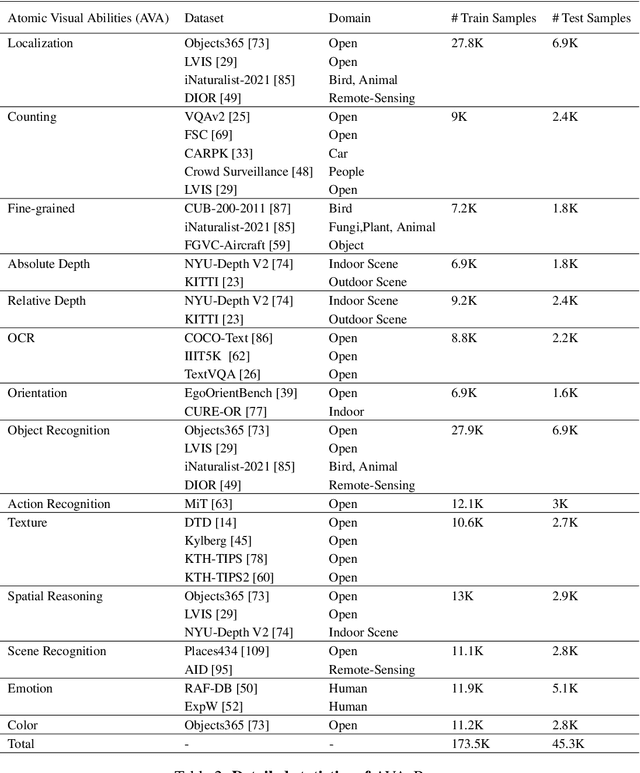

Abstract:The rise of vision foundation models (VFMs) calls for systematic evaluation. A common approach pairs VFMs with large language models (LLMs) as general-purpose heads, followed by evaluation on broad Visual Question Answering (VQA) benchmarks. However, this protocol has two key blind spots: (i) the instruction tuning data may not align with VQA test distributions, meaning a wrong prediction can stem from such data mismatch rather than a VFM' visual shortcomings; (ii) VQA benchmarks often require multiple visual abilities, making it hard to tell whether errors stem from lacking all required abilities or just a single critical one. To address these gaps, we introduce AVA-Bench, the first benchmark that explicitly disentangles 14 Atomic Visual Abilities (AVAs) -- foundational skills like localization, depth estimation, and spatial understanding that collectively support complex visual reasoning tasks. By decoupling AVAs and matching training and test distributions within each, AVA-Bench pinpoints exactly where a VFM excels or falters. Applying AVA-Bench to leading VFMs thus reveals distinctive "ability fingerprints," turning VFM selection from educated guesswork into principled engineering. Notably, we find that a 0.5B LLM yields similar VFM rankings as a 7B LLM while cutting GPU hours by 8x, enabling more efficient evaluation. By offering a comprehensive and transparent benchmark, we hope AVA-Bench lays the foundation for the next generation of VFMs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge