Xin Chen
Univ. California, Santa Barbara
ViBES: A Conversational Agent with Behaviorally-Intelligent 3D Virtual Body
Dec 16, 2025



Abstract:Human communication is inherently multimodal and social: words, prosody, and body language jointly carry intent. Yet most prior systems model human behavior as a translation task co-speech gesture or text-to-motion that maps a fixed utterance to motion clips-without requiring agentic decision-making about when to move, what to do, or how to adapt across multi-turn dialogue. This leads to brittle timing, weak social grounding, and fragmented stacks where speech, text, and motion are trained or inferred in isolation. We introduce ViBES (Voice in Behavioral Expression and Synchrony), a conversational 3D agent that jointly plans language and movement and executes dialogue-conditioned body actions. Concretely, ViBES is a speech-language-behavior (SLB) model with a mixture-of-modality-experts (MoME) backbone: modality-partitioned transformer experts for speech, facial expression, and body motion. The model processes interleaved multimodal token streams with hard routing by modality (parameters are split per expert), while sharing information through cross-expert attention. By leveraging strong pretrained speech-language models, the agent supports mixed-initiative interaction: users can speak, type, or issue body-action directives mid-conversation, and the system exposes controllable behavior hooks for streaming responses. We further benchmark on multi-turn conversation with automatic metrics of dialogue-motion alignment and behavior quality, and observe consistent gains over strong co-speech and text-to-motion baselines. ViBES goes beyond "speech-conditioned motion generation" toward agentic virtual bodies where language, prosody, and movement are jointly generated, enabling controllable, socially competent 3D interaction. Code and data will be made available at: ai.stanford.edu/~juze/ViBES/
SSL-MedSAM2: A Semi-supervised Medical Image Segmentation Framework Powered by Few-shot Learning of SAM2
Dec 12, 2025Abstract:Despite the success of deep learning based models in medical image segmentation, most state-of-the-art (SOTA) methods perform fully-supervised learning, which commonly rely on large scale annotated training datasets. However, medical image annotation is highly time-consuming, hindering its clinical applications. Semi-supervised learning (SSL) has been emerged as an appealing strategy in training with limited annotations, largely reducing the labelling cost. We propose a novel SSL framework SSL-MedSAM2, which contains a training-free few-shot learning branch TFFS-MedSAM2 based on the pretrained large foundation model Segment Anything Model 2 (SAM2) for pseudo label generation, and an iterative fully-supervised learning branch FSL-nnUNet based on nnUNet for pseudo label refinement. The results on MICCAI2025 challenge CARE-LiSeg (Liver Segmentation) demonstrate an outstanding performance of SSL-MedSAM2 among other methods. The average dice scores on the test set in GED4 and T1 MRI are 0.9710 and 0.9648 respectively, and the Hausdorff distances are 20.07 and 21.97 respectively. The code is available via https://github.com/naisops/SSL-MedSAM2/tree/main.
InterAgent: Physics-based Multi-agent Command Execution via Diffusion on Interaction Graphs
Dec 12, 2025Abstract:Humanoid agents are expected to emulate the complex coordination inherent in human social behaviors. However, existing methods are largely confined to single-agent scenarios, overlooking the physically plausible interplay essential for multi-agent interactions. To bridge this gap, we propose InterAgent, the first end-to-end framework for text-driven physics-based multi-agent humanoid control. At its core, we introduce an autoregressive diffusion transformer equipped with multi-stream blocks, which decouples proprioception, exteroception, and action to mitigate cross-modal interference while enabling synergistic coordination. We further propose a novel interaction graph exteroception representation that explicitly captures fine-grained joint-to-joint spatial dependencies to facilitate network learning. Additionally, within it we devise a sparse edge-based attention mechanism that dynamically prunes redundant connections and emphasizes critical inter-agent spatial relations, thereby enhancing the robustness of interaction modeling. Extensive experiments demonstrate that InterAgent consistently outperforms multiple strong baselines, achieving state-of-the-art performance. It enables producing coherent, physically plausible, and semantically faithful multi-agent behaviors from only text prompts. Our code and data will be released to facilitate future research.
MS-BART: Unified Modeling of Mass Spectra and Molecules for Structure Elucidation
Oct 23, 2025Abstract:Mass spectrometry (MS) plays a critical role in molecular identification, significantly advancing scientific discovery. However, structure elucidation from MS data remains challenging due to the scarcity of annotated spectra. While large-scale pretraining has proven effective in addressing data scarcity in other domains, applying this paradigm to mass spectrometry is hindered by the complexity and heterogeneity of raw spectral signals. To address this, we propose MS-BART, a unified modeling framework that maps mass spectra and molecular structures into a shared token vocabulary, enabling cross-modal learning through large-scale pretraining on reliably computed fingerprint-molecule datasets. Multi-task pretraining objectives further enhance MS-BART's generalization by jointly optimizing denoising and translation task. The pretrained model is subsequently transferred to experimental spectra through finetuning on fingerprint predictions generated with MIST, a pre-trained spectral inference model, thereby enhancing robustness to real-world spectral variability. While finetuning alleviates the distributional difference, MS-BART still suffers molecular hallucination and requires further alignment. We therefore introduce a chemical feedback mechanism that guides the model toward generating molecules closer to the reference structure. Extensive evaluations demonstrate that MS-BART achieves SOTA performance across 5/12 key metrics on MassSpecGym and NPLIB1 and is faster by one order of magnitude than competing diffusion-based methods, while comprehensive ablation studies systematically validate the model's effectiveness and robustness.
MARS2 2025 Challenge on Multimodal Reasoning: Datasets, Methods, Results, Discussion, and Outlook
Sep 17, 2025
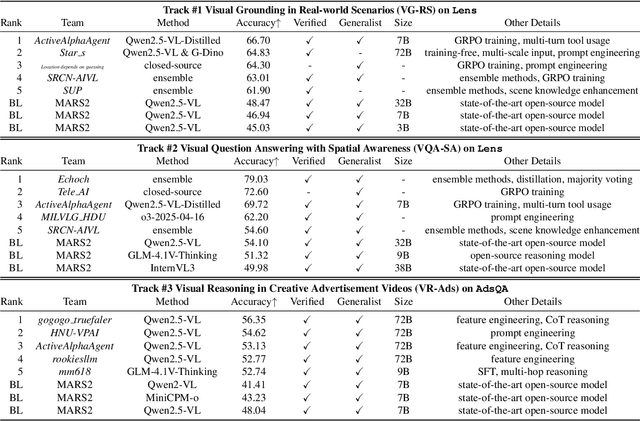

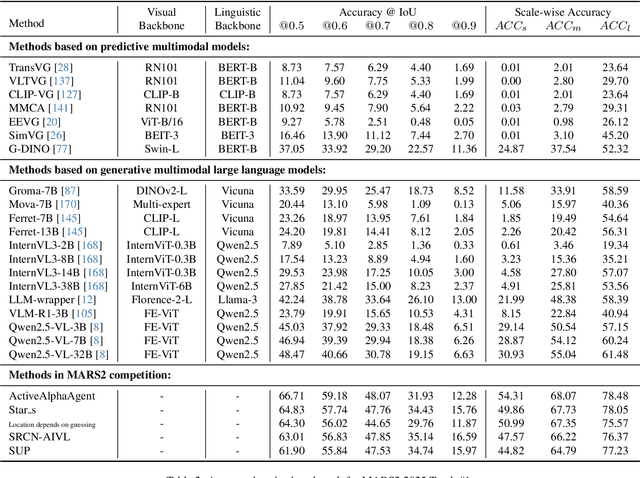
Abstract:This paper reviews the MARS2 2025 Challenge on Multimodal Reasoning. We aim to bring together different approaches in multimodal machine learning and LLMs via a large benchmark. We hope it better allows researchers to follow the state-of-the-art in this very dynamic area. Meanwhile, a growing number of testbeds have boosted the evolution of general-purpose large language models. Thus, this year's MARS2 focuses on real-world and specialized scenarios to broaden the multimodal reasoning applications of MLLMs. Our organizing team released two tailored datasets Lens and AdsQA as test sets, which support general reasoning in 12 daily scenarios and domain-specific reasoning in advertisement videos, respectively. We evaluated 40+ baselines that include both generalist MLLMs and task-specific models, and opened up three competition tracks, i.e., Visual Grounding in Real-world Scenarios (VG-RS), Visual Question Answering with Spatial Awareness (VQA-SA), and Visual Reasoning in Creative Advertisement Videos (VR-Ads). Finally, 76 teams from the renowned academic and industrial institutions have registered and 40+ valid submissions (out of 1200+) have been included in our ranking lists. Our datasets, code sets (40+ baselines and 15+ participants' methods), and rankings are publicly available on the MARS2 workshop website and our GitHub organization page https://github.com/mars2workshop/, where our updates and announcements of upcoming events will be continuously provided.
OneCAT: Decoder-Only Auto-Regressive Model for Unified Understanding and Generation
Sep 03, 2025



Abstract:We introduce OneCAT, a unified multimodal model that seamlessly integrates understanding, generation, and editing within a novel, pure decoder-only transformer architecture. Our framework uniquely eliminates the need for external components such as Vision Transformers (ViT) or vision tokenizer during inference, leading to significant efficiency gains, especially for high-resolution inputs. This is achieved through a modality-specific Mixture-of-Experts (MoE) structure trained with a single autoregressive (AR) objective, which also natively supports dynamic resolutions. Furthermore, we pioneer a multi-scale visual autoregressive mechanism within the Large Language Model (LLM) that drastically reduces decoding steps compared to diffusion-based methods while maintaining state-of-the-art performance. Our findings demonstrate the powerful potential of pure autoregressive modeling as a sufficient and elegant foundation for unified multimodal intelligence. As a result, OneCAT sets a new performance standard, outperforming existing open-source unified multimodal models across benchmarks for multimodal generation, editing, and understanding.
E-BayesSAM: Efficient Bayesian Adaptation of SAM with Self-Optimizing KAN-Based Interpretation for Uncertainty-Aware Ultrasonic Segmentation
Aug 24, 2025Abstract:Although the Segment Anything Model (SAM) has advanced medical image segmentation, its Bayesian adaptation for uncertainty-aware segmentation remains hindered by three key issues: (1) instability in Bayesian fine-tuning of large pre-trained SAMs; (2) high computation cost due to SAM's massive parameters; (3) SAM's black-box design limits interpretability. To overcome these, we propose E-BayesSAM, an efficient framework combining Token-wise Variational Bayesian Inference (T-VBI) for efficienty Bayesian adaptation and Self-Optimizing Kolmogorov-Arnold Network (SO-KAN) for improving interpretability. T-VBI innovatively reinterprets SAM's output tokens as dynamic probabilistic weights and reparameterizes them as latent variables without auxiliary training, enabling training-free VBI for uncertainty estimation. SO-KAN improves token prediction with learnable spline activations via self-supervised learning, providing insight to prune redundant tokens to boost efficiency and accuracy. Experiments on five ultrasound datasets demonstrated that E-BayesSAM achieves: (i) real-time inference (0.03s/image), (ii) superior segmentation accuracy (average DSC: Pruned E-BayesSAM's 89.0\% vs. E-BayesSAM's 88.0% vs. MedSAM's 88.3%), and (iii) identification of four critical tokens governing SAM's decisions. By unifying efficiency, reliability, and interpretability, E-BayesSAM bridges SAM's versatility with clinical needs, advancing deployment in safety-critical medical applications. The source code is available at https://github.com/mp31192/E-BayesSAM.
Motion2Motion: Cross-topology Motion Transfer with Sparse Correspondence
Aug 18, 2025



Abstract:This work studies the challenge of transfer animations between characters whose skeletal topologies differ substantially. While many techniques have advanced retargeting techniques in decades, transfer motions across diverse topologies remains less-explored. The primary obstacle lies in the inherent topological inconsistency between source and target skeletons, which restricts the establishment of straightforward one-to-one bone correspondences. Besides, the current lack of large-scale paired motion datasets spanning different topological structures severely constrains the development of data-driven approaches. To address these limitations, we introduce Motion2Motion, a novel, training-free framework. Simply yet effectively, Motion2Motion works with only one or a few example motions on the target skeleton, by accessing a sparse set of bone correspondences between the source and target skeletons. Through comprehensive qualitative and quantitative evaluations, we demonstrate that Motion2Motion achieves efficient and reliable performance in both similar-skeleton and cross-species skeleton transfer scenarios. The practical utility of our approach is further evidenced by its successful integration in downstream applications and user interfaces, highlighting its potential for industrial applications. Code and data are available at https://lhchen.top/Motion2Motion.
RepreGuard: Detecting LLM-Generated Text by Revealing Hidden Representation Patterns
Aug 18, 2025



Abstract:Detecting content generated by large language models (LLMs) is crucial for preventing misuse and building trustworthy AI systems. Although existing detection methods perform well, their robustness in out-of-distribution (OOD) scenarios is still lacking. In this paper, we hypothesize that, compared to features used by existing detection methods, the internal representations of LLMs contain more comprehensive and raw features that can more effectively capture and distinguish the statistical pattern differences between LLM-generated texts (LGT) and human-written texts (HWT). We validated this hypothesis across different LLMs and observed significant differences in neural activation patterns when processing these two types of texts. Based on this, we propose RepreGuard, an efficient statistics-based detection method. Specifically, we first employ a surrogate model to collect representation of LGT and HWT, and extract the distinct activation feature that can better identify LGT. We can classify the text by calculating the projection score of the text representations along this feature direction and comparing with a precomputed threshold. Experimental results show that RepreGuard outperforms all baselines with average 94.92% AUROC on both in-distribution (ID) and OOD scenarios, while also demonstrating robust resilience to various text sizes and mainstream attacks. Data and code are publicly available at: https://github.com/NLP2CT/RepreGuard
ChemDFM-R: An Chemical Reasoner LLM Enhanced with Atomized Chemical Knowledge
Jul 30, 2025Abstract:While large language models (LLMs) have achieved impressive progress, their application in scientific domains such as chemistry remains hindered by shallow domain understanding and limited reasoning capabilities. In this work, we focus on the specific field of chemistry and develop a Chemical Reasoner LLM, ChemDFM-R. We first construct a comprehensive dataset of atomized knowledge points to enhance the model's understanding of the fundamental principles and logical structure of chemistry. Then, we propose a mix-sourced distillation strategy that integrates expert-curated knowledge with general-domain reasoning skills, followed by domain-specific reinforcement learning to enhance chemical reasoning. Experiments on diverse chemical benchmarks demonstrate that ChemDFM-R achieves cutting-edge performance while providing interpretable, rationale-driven outputs. Further case studies illustrate how explicit reasoning chains significantly improve the reliability, transparency, and practical utility of the model in real-world human-AI collaboration scenarios.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge