Zaiyi Liu
Treatment Outcome Prediction for Intracerebral Hemorrhage via Generative Prognostic Model with Imaging and Tabular Data
Jul 24, 2023


Abstract:Intracerebral hemorrhage (ICH) is the second most common and deadliest form of stroke. Despite medical advances, predicting treat ment outcomes for ICH remains a challenge. This paper proposes a novel prognostic model that utilizes both imaging and tabular data to predict treatment outcome for ICH. Our model is trained on observational data collected from non-randomized controlled trials, providing reliable predictions of treatment success. Specifically, we propose to employ a variational autoencoder model to generate a low-dimensional prognostic score, which can effectively address the selection bias resulting from the non-randomized controlled trials. Importantly, we develop a variational distributions combination module that combines the information from imaging data, non-imaging clinical data, and treatment assignment to accurately generate the prognostic score. We conducted extensive experiments on a real-world clinical dataset of intracerebral hemorrhage. Our proposed method demonstrates a substantial improvement in treatment outcome prediction compared to existing state-of-the-art approaches. Code is available at https://github.com/med-air/TOP-GPM
Parse and Recall: Towards Accurate Lung Nodule Malignancy Prediction like Radiologists
Jul 20, 2023



Abstract:Lung cancer is a leading cause of death worldwide and early screening is critical for improving survival outcomes. In clinical practice, the contextual structure of nodules and the accumulated experience of radiologists are the two core elements related to the accuracy of identification of benign and malignant nodules. Contextual information provides comprehensive information about nodules such as location, shape, and peripheral vessels, and experienced radiologists can search for clues from previous cases as a reference to enrich the basis of decision-making. In this paper, we propose a radiologist-inspired method to simulate the diagnostic process of radiologists, which is composed of context parsing and prototype recalling modules. The context parsing module first segments the context structure of nodules and then aggregates contextual information for a more comprehensive understanding of the nodule. The prototype recalling module utilizes prototype-based learning to condense previously learned cases as prototypes for comparative analysis, which is updated online in a momentum way during training. Building on the two modules, our method leverages both the intrinsic characteristics of the nodules and the external knowledge accumulated from other nodules to achieve a sound diagnosis. To meet the needs of both low-dose and noncontrast screening, we collect a large-scale dataset of 12,852 and 4,029 nodules from low-dose and noncontrast CTs respectively, each with pathology- or follow-up-confirmed labels. Experiments on several datasets demonstrate that our method achieves advanced screening performance on both low-dose and noncontrast scenarios.
Cluster-Induced Mask Transformers for Effective Opportunistic Gastric Cancer Screening on Non-contrast CT Scans
Jul 16, 2023



Abstract:Gastric cancer is the third leading cause of cancer-related mortality worldwide, but no guideline-recommended screening test exists. Existing methods can be invasive, expensive, and lack sensitivity to identify early-stage gastric cancer. In this study, we explore the feasibility of using a deep learning approach on non-contrast CT scans for gastric cancer detection. We propose a novel cluster-induced Mask Transformer that jointly segments the tumor and classifies abnormality in a multi-task manner. Our model incorporates learnable clusters that encode the texture and shape prototypes of gastric cancer, utilizing self- and cross-attention to interact with convolutional features. In our experiments, the proposed method achieves a sensitivity of 85.0% and specificity of 92.6% for detecting gastric tumors on a hold-out test set consisting of 100 patients with cancer and 148 normal. In comparison, two radiologists have an average sensitivity of 73.5% and specificity of 84.3%. We also obtain a specificity of 97.7% on an external test set with 903 normal cases. Our approach performs comparably to established state-of-the-art gastric cancer screening tools like blood testing and endoscopy, while also being more sensitive in detecting early-stage cancer. This demonstrates the potential of our approach as a novel, non-invasive, low-cost, and accurate method for opportunistic gastric cancer screening.
Rethinking Mitosis Detection: Towards Diverse Data and Feature Representation
Jul 12, 2023Abstract:Mitosis detection is one of the fundamental tasks in computational pathology, which is extremely challenging due to the heterogeneity of mitotic cell. Most of the current studies solve the heterogeneity in the technical aspect by increasing the model complexity. However, lacking consideration of the biological knowledge and the complex model design may lead to the overfitting problem while limited the generalizability of the detection model. In this paper, we systematically study the morphological appearances in different mitotic phases as well as the ambiguous non-mitotic cells and identify that balancing the data and feature diversity can achieve better generalizability. Based on this observation, we propose a novel generalizable framework (MitDet) for mitosis detection. The data diversity is considered by the proposed diversity-guided sample balancing (DGSB). And the feature diversity is preserved by inter- and intra- class feature diversity-preserved module (InCDP). Stain enhancement (SE) module is introduced to enhance the domain-relevant diversity of both data and features simultaneously. Extensive experiments have demonstrated that our proposed model outperforms all the SOTA approaches in several popular mitosis detection datasets in both internal and external test sets using minimal annotation efforts with point annotations only. Comprehensive ablation studies have also proven the effectiveness of the rethinking of data and feature diversity balancing. By analyzing the results quantitatively and qualitatively, we believe that our proposed model not only achieves SOTA performance but also might inspire the future studies in new perspectives. Source code is at https://github.com/Onehour0108/MitDet.
UOD: Universal One-shot Detection of Anatomical Landmarks
Jun 14, 2023Abstract:One-shot medical landmark detection gains much attention and achieves great success for its label-efficient training process. However, existing one-shot learning methods are highly specialized in a single domain and suffer domain preference heavily in the situation of multi-domain unlabeled data. Moreover, one-shot learning is not robust that it faces performance drop when annotating a sub-optimal image. To tackle these issues, we resort to developing a domain-adaptive one-shot landmark detection framework for handling multi-domain medical images, named Universal One-shot Detection (UOD). UOD consists of two stages and two corresponding universal models which are designed as combinations of domain-specific modules and domain-shared modules. In the first stage, a domain-adaptive convolution model is self-supervised learned to generate pseudo landmark labels. In the second stage, we design a domain-adaptive transformer to eliminate domain preference and build the global context for multi-domain data. Even though only one annotated sample from each domain is available for training, the domain-shared modules help UOD aggregate all one-shot samples to detect more robust and accurate landmarks. We investigated both qualitatively and quantitatively the proposed UOD on three widely-used public X-ray datasets in different anatomical domains (i.e., head, hand, chest) and obtained state-of-the-art performances in each domain.
Domain Generalization for Mammographic Image Analysis via Contrastive Learning
Apr 20, 2023Abstract:Mammographic image analysis is a fundamental problem in the computer-aided diagnosis scheme, which has recently made remarkable progress with the advance of deep learning. However, the construction of a deep learning model requires training data that are large and sufficiently diverse in terms of image style and quality. In particular, the diversity of image style may be majorly attributed to the vendor factor. However, mammogram collection from vendors as many as possible is very expensive and sometimes impractical for laboratory-scale studies. Accordingly, to further augment the generalization capability of deep learning models to various vendors with limited resources, a new contrastive learning scheme is developed. Specifically, the backbone network is firstly trained with a multi-style and multi-view unsupervised self-learning scheme for the embedding of invariant features to various vendor styles. Afterward, the backbone network is then recalibrated to the downstream tasks of mass detection, multi-view mass matching, BI-RADS classification and breast density classification with specific supervised learning. The proposed method is evaluated with mammograms from four vendors and two unseen public datasets. The experimental results suggest that our approach can effectively improve analysis performance on both seen and unseen domains, and outperforms many state-of-the-art (SOTA) generalization methods.
Devil is in the Queries: Advancing Mask Transformers for Real-world Medical Image Segmentation and Out-of-Distribution Localization
Apr 01, 2023



Abstract:Real-world medical image segmentation has tremendous long-tailed complexity of objects, among which tail conditions correlate with relatively rare diseases and are clinically significant. A trustworthy medical AI algorithm should demonstrate its effectiveness on tail conditions to avoid clinically dangerous damage in these out-of-distribution (OOD) cases. In this paper, we adopt the concept of object queries in Mask Transformers to formulate semantic segmentation as a soft cluster assignment. The queries fit the feature-level cluster centers of inliers during training. Therefore, when performing inference on a medical image in real-world scenarios, the similarity between pixels and the queries detects and localizes OOD regions. We term this OOD localization as MaxQuery. Furthermore, the foregrounds of real-world medical images, whether OOD objects or inliers, are lesions. The difference between them is less than that between the foreground and background, possibly misleading the object queries to focus redundantly on the background. Thus, we propose a query-distribution (QD) loss to enforce clear boundaries between segmentation targets and other regions at the query level, improving the inlier segmentation and OOD indication. Our proposed framework is tested on two real-world segmentation tasks, i.e., segmentation of pancreatic and liver tumors, outperforming previous state-of-the-art algorithms by an average of 7.39% on AUROC, 14.69% on AUPR, and 13.79% on FPR95 for OOD localization. On the other hand, our framework improves the performance of inlier segmentation by an average of 5.27% DSC when compared with the leading baseline nnUNet.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
FedDBL: Communication and Data Efficient Federated Deep-Broad Learning for Histopathological Tissue Classification
Feb 24, 2023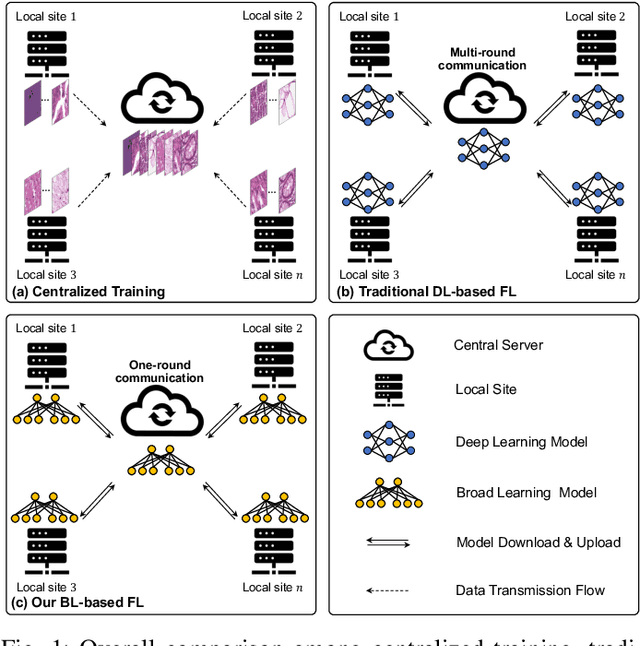
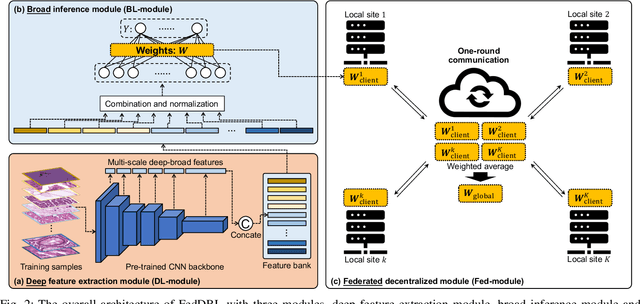
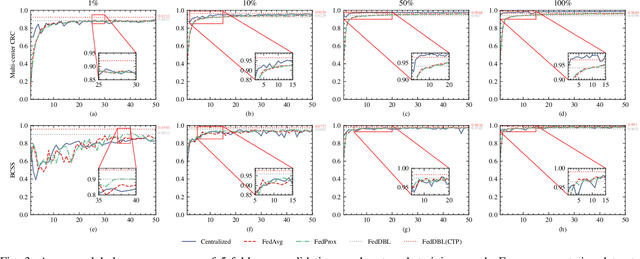
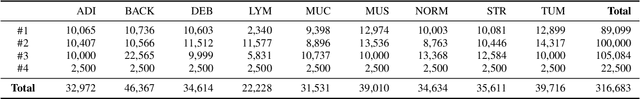
Abstract:Histopathological tissue classification is a fundamental task in computational pathology. Deep learning-based models have achieved superior performance but centralized training with data centralization suffers from the privacy leakage problem. Federated learning (FL) can safeguard privacy by keeping training samples locally, but existing FL-based frameworks require a large number of well-annotated training samples and numerous rounds of communication which hinder their practicability in the real-world clinical scenario. In this paper, we propose a universal and lightweight federated learning framework, named Federated Deep-Broad Learning (FedDBL), to achieve superior classification performance with limited training samples and only one-round communication. By simply associating a pre-trained deep learning feature extractor, a fast and lightweight broad learning inference system and a classical federated aggregation approach, FedDBL can dramatically reduce data dependency and improve communication efficiency. Five-fold cross-validation demonstrates that FedDBL greatly outperforms the competitors with only one-round communication and limited training samples, while it even achieves comparable performance with the ones under multiple-round communications. Furthermore, due to the lightweight design and one-round communication, FedDBL reduces the communication burden from 4.6GB to only 276.5KB per client using the ResNet-50 backbone at 50-round training. Since no data or deep model sharing across different clients, the privacy issue is well-solved and the model security is guaranteed with no model inversion attack risk. Code is available at https://github.com/tianpeng-deng/FedDBL.
Towards a Single Unified Model for Effective Detection, Segmentation, and Diagnosis of Eight Major Cancers Using a Large Collection of CT Scans
Jan 28, 2023



Abstract:Human readers or radiologists routinely perform full-body multi-organ multi-disease detection and diagnosis in clinical practice, while most medical AI systems are built to focus on single organs with a narrow list of a few diseases. This might severely limit AI's clinical adoption. A certain number of AI models need to be assembled non-trivially to match the diagnostic process of a human reading a CT scan. In this paper, we construct a Unified Tumor Transformer (UniT) model to detect (tumor existence and location) and diagnose (tumor characteristics) eight major cancer-prevalent organs in CT scans. UniT is a query-based Mask Transformer model with the output of multi-organ and multi-tumor semantic segmentation. We decouple the object queries into organ queries, detection queries and diagnosis queries, and further establish hierarchical relationships among the three groups. This clinically-inspired architecture effectively assists inter- and intra-organ representation learning of tumors and facilitates the resolution of these complex, anatomically related multi-organ cancer image reading tasks. UniT is trained end-to-end using a curated large-scale CT images of 10,042 patients including eight major types of cancers and occurring non-cancer tumors (all are pathology-confirmed with 3D tumor masks annotated by radiologists). On the test set of 631 patients, UniT has demonstrated strong performance under a set of clinically relevant evaluation metrics, substantially outperforming both multi-organ segmentation methods and an assembly of eight single-organ expert models in tumor detection, segmentation, and diagnosis. Such a unified multi-cancer image reading model (UniT) can significantly reduce the number of false positives produced by combined multi-system models. This moves one step closer towards a universal high-performance cancer screening tool.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge