Minfeng Xu
MedVR: Annotation-Free Medical Visual Reasoning via Agentic Reinforcement Learning
Apr 09, 2026Abstract:Medical Vision-Language Models (VLMs) hold immense promise for complex clinical tasks, but their reasoning capabilities are often constrained by text-only paradigms that fail to ground inferences in visual evidence. This limitation not only curtails performance on tasks requiring fine-grained visual analysis but also introduces risks of visual hallucination in safety-critical applications. Thus, we introduce MedVR, a novel reinforcement learning framework that enables annotation-free visual reasoning for medical VLMs. Its core innovation lies in two synergistic mechanisms: Entropy-guided Visual Regrounding (EVR) uses model uncertainty to direct exploration, while Consensus-based Credit Assignment (CCA) distills pseudo-supervision from rollout agreement. Without any human annotations for intermediate steps, MedVR achieves state-of-the-art performance on diverse public medical VQA benchmarks, significantly outperforming existing models. By learning to reason directly with visual evidence, MedVR promotes the robustness and transparency essential for accelerating the clinical deployment of medical AI.
Photon: Speedup Volume Understanding with Efficient Multimodal Large Language Models
Mar 26, 2026Abstract:Multimodal large language models are promising for clinical visual question answering tasks, but scaling to 3D imaging is hindered by high computational costs. Prior methods often rely on 2D slices or fixed-length token compression, disrupting volumetric continuity and obscuring subtle findings. We present Photon, a framework that represents 3D medical volumes with token sequences of variable length. Photon introduces instruction-conditioned token scheduling and surrogate gradient propagation to adaptively reduce tokens during both training and inference, which lowers computational cost while mitigating the attention dilution caused by redundant tokens. It incorporates a custom backpropagation rule with gradient restoration to enable differentiable optimization despite discrete token drop. To stabilize token compression and ensure reliable use of visual evidence, Photon further applies regularization objectives that mitigate language-only bias and improve reliability. Experiments on diverse medical visual question answering tasks show that Photon achieves state-of-the-art accuracy while reducing resource usage and accelerating both training and inference.
MUSE: Multi-Scale Dense Self-Distillation for Nucleus Detection and Classification
Nov 07, 2025



Abstract:Nucleus detection and classification (NDC) in histopathology analysis is a fundamental task that underpins a wide range of high-level pathology applications. However, existing methods heavily rely on labor-intensive nucleus-level annotations and struggle to fully exploit large-scale unlabeled data for learning discriminative nucleus representations. In this work, we propose MUSE (MUlti-scale denSE self-distillation), a novel self-supervised learning method tailored for NDC. At its core is NuLo (Nucleus-based Local self-distillation), a coordinate-guided mechanism that enables flexible local self-distillation based on predicted nucleus positions. By removing the need for strict spatial alignment between augmented views, NuLo allows critical cross-scale alignment, thus unlocking the capacity of models for fine-grained nucleus-level representation. To support MUSE, we design a simple yet effective encoder-decoder architecture and a large field-of-view semi-supervised fine-tuning strategy that together maximize the value of unlabeled pathology images. Extensive experiments on three widely used benchmarks demonstrate that MUSE effectively addresses the core challenges of histopathological NDC. The resulting models not only surpass state-of-the-art supervised baselines but also outperform generic pathology foundation models.
Anatomy-Aware Low-Dose CT Denoising via Pretrained Vision Models and Semantic-Guided Contrastive Learning
Aug 11, 2025Abstract:To reduce radiation exposure and improve the diagnostic efficacy of low-dose computed tomography (LDCT), numerous deep learning-based denoising methods have been developed to mitigate noise and artifacts. However, most of these approaches ignore the anatomical semantics of human tissues, which may potentially result in suboptimal denoising outcomes. To address this problem, we propose ALDEN, an anatomy-aware LDCT denoising method that integrates semantic features of pretrained vision models (PVMs) with adversarial and contrastive learning. Specifically, we introduce an anatomy-aware discriminator that dynamically fuses hierarchical semantic features from reference normal-dose CT (NDCT) via cross-attention mechanisms, enabling tissue-specific realism evaluation in the discriminator. In addition, we propose a semantic-guided contrastive learning module that enforces anatomical consistency by contrasting PVM-derived features from LDCT, denoised CT and NDCT, preserving tissue-specific patterns through positive pairs and suppressing artifacts via dual negative pairs. Extensive experiments conducted on two LDCT denoising datasets reveal that ALDEN achieves the state-of-the-art performance, offering superior anatomy preservation and substantially reducing over-smoothing issue of previous work. Further validation on a downstream multi-organ segmentation task (encompassing 117 anatomical structures) affirms the model's ability to maintain anatomical awareness.
A Continual Learning-driven Model for Accurate and Generalizable Segmentation of Clinically Comprehensive and Fine-grained Whole-body Anatomies in CT
Mar 16, 2025Abstract:Precision medicine in the quantitative management of chronic diseases and oncology would be greatly improved if the Computed Tomography (CT) scan of any patient could be segmented, parsed and analyzed in a precise and detailed way. However, there is no such fully annotated CT dataset with all anatomies delineated for training because of the exceptionally high manual cost, the need for specialized clinical expertise, and the time required to finish the task. To this end, we proposed a novel continual learning-driven CT model that can segment complete anatomies presented using dozens of previously partially labeled datasets, dynamically expanding its capacity to segment new ones without compromising previously learned organ knowledge. Existing multi-dataset approaches are not able to dynamically segment new anatomies without catastrophic forgetting and would encounter optimization difficulty or infeasibility when segmenting hundreds of anatomies across the whole range of body regions. Our single unified CT segmentation model, CL-Net, can highly accurately segment a clinically comprehensive set of 235 fine-grained whole-body anatomies. Composed of a universal encoder, multiple optimized and pruned decoders, CL-Net is developed using 13,952 CT scans from 20 public and 16 private high-quality partially labeled CT datasets of various vendors, different contrast phases, and pathologies. Extensive evaluation demonstrates that CL-Net consistently outperforms the upper limit of an ensemble of 36 specialist nnUNets trained per dataset with the complexity of 5% model size and significantly surpasses the segmentation accuracy of recent leading Segment Anything-style medical image foundation models by large margins. Our continual learning-driven CL-Net model would lay a solid foundation to facilitate many downstream tasks of oncology and chronic diseases using the most widely adopted CT imaging.
MRS: A Fast Sampler for Mean Reverting Diffusion based on ODE and SDE Solvers
Feb 11, 2025Abstract:In applications of diffusion models, controllable generation is of practical significance, but is also challenging. Current methods for controllable generation primarily focus on modifying the score function of diffusion models, while Mean Reverting (MR) Diffusion directly modifies the structure of the stochastic differential equation (SDE), making the incorporation of image conditions simpler and more natural. However, current training-free fast samplers are not directly applicable to MR Diffusion. And thus MR Diffusion requires hundreds of NFEs (number of function evaluations) to obtain high-quality samples. In this paper, we propose a new algorithm named MRS (MR Sampler) to reduce the sampling NFEs of MR Diffusion. We solve the reverse-time SDE and the probability flow ordinary differential equation (PF-ODE) associated with MR Diffusion, and derive semi-analytical solutions. The solutions consist of an analytical function and an integral parameterized by a neural network. Based on this solution, we can generate high-quality samples in fewer steps. Our approach does not require training and supports all mainstream parameterizations, including noise prediction, data prediction and velocity prediction. Extensive experiments demonstrate that MR Sampler maintains high sampling quality with a speedup of 10 to 20 times across ten different image restoration tasks. Our algorithm accelerates the sampling procedure of MR Diffusion, making it more practical in controllable generation.
A Joint Representation Using Continuous and Discrete Features for Cardiovascular Diseases Risk Prediction on Chest CT Scans
Oct 24, 2024



Abstract:Cardiovascular diseases (CVD) remain a leading health concern and contribute significantly to global mortality rates. While clinical advancements have led to a decline in CVD mortality, accurately identifying individuals who could benefit from preventive interventions remains an unsolved challenge in preventive cardiology. Current CVD risk prediction models, recommended by guidelines, are based on limited traditional risk factors or use CT imaging to acquire quantitative biomarkers, and still have limitations in predictive accuracy and applicability. On the other hand, end-to-end trained CVD risk prediction methods leveraging deep learning on CT images often fail to provide transparent and explainable decision grounds for assisting physicians. In this work, we proposed a novel joint representation that integrates discrete quantitative biomarkers and continuous deep features extracted from chest CT scans. Our approach initiated with a deep CVD risk classification model by capturing comprehensive continuous deep learning features while jointly obtaining currently clinical-established quantitative biomarkers via segmentation models. In the feature joint representation stage, we use an instance-wise feature-gated mechanism to align the continuous and discrete features, followed by a soft instance-wise feature interaction mechanism fostering independent and effective feature interaction for the final CVD risk prediction. Our method substantially improves CVD risk predictive performance and offers individual contribution analysis of each biomarker, which is important in assisting physicians' decision-making processes. We validated our method on a public chest low-dose CT dataset and a private external chest standard-dose CT patient cohort of 17,207 CT volumes from 6,393 unique subjects, and demonstrated superior predictive performance, achieving AUCs of 0.875 and 0.843, respectively.
Cross-Phase Mutual Learning Framework for Pulmonary Embolism Identification on Non-Contrast CT Scans
Jul 16, 2024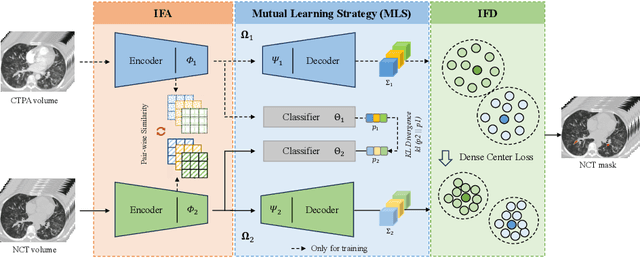
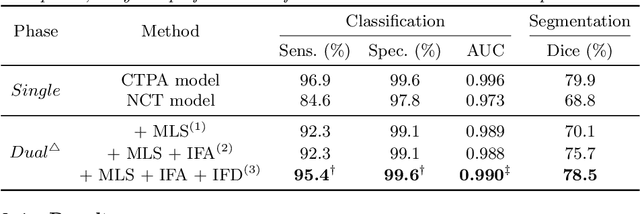
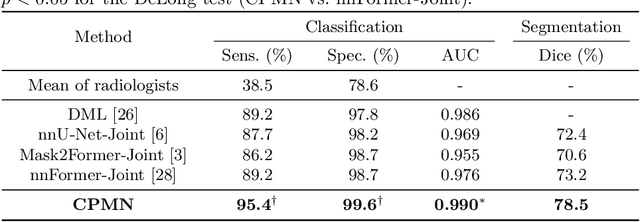
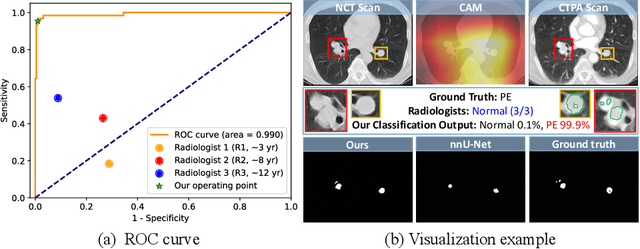
Abstract:Pulmonary embolism (PE) is a life-threatening condition where rapid and accurate diagnosis is imperative yet difficult due to predominantly atypical symptomatology. Computed tomography pulmonary angiography (CTPA) is acknowledged as the gold standard imaging tool in clinics, yet it can be contraindicated for emergency department (ED) patients and represents an onerous procedure, thus necessitating PE identification through non-contrast CT (NCT) scans. In this work, we explore the feasibility of applying a deep-learning approach to NCT scans for PE identification. We propose a novel Cross-Phase Mutual learNing framework (CPMN) that fosters knowledge transfer from CTPA to NCT, while concurrently conducting embolism segmentation and abnormality classification in a multi-task manner. The proposed CPMN leverages the Inter-Feature Alignment (IFA) strategy that enhances spatial contiguity and mutual learning between the dual-pathway network, while the Intra-Feature Discrepancy (IFD) strategy can facilitate precise segmentation of PE against complex backgrounds for single-pathway networks. For a comprehensive assessment of the proposed approach, a large-scale dual-phase dataset containing 334 PE patients and 1,105 normal subjects has been established. Experimental results demonstrate that CPMN achieves the leading identification performance, which is 95.4\% and 99.6\% in patient-level sensitivity and specificity on NCT scans, indicating the potential of our approach as an economical, accessible, and precise tool for PE identification in clinical practice.
Rapid and Accurate Diagnosis of Acute Aortic Syndrome using Non-contrast CT: A Large-scale, Retrospective, Multi-center and AI-based Study
Jun 25, 2024



Abstract:Chest pain symptoms are highly prevalent in emergency departments (EDs), where acute aortic syndrome (AAS) is a catastrophic cardiovascular emergency with a high fatality rate, especially when timely and accurate treatment is not administered. However, current triage practices in the ED can cause up to approximately half of patients with AAS to have an initially missed diagnosis or be misdiagnosed as having other acute chest pain conditions. Subsequently, these AAS patients will undergo clinically inaccurate or suboptimal differential diagnosis. Fortunately, even under these suboptimal protocols, nearly all these patients underwent non-contrast CT covering the aorta anatomy at the early stage of differential diagnosis. In this study, we developed an artificial intelligence model (DeepAAS) using non-contrast CT, which is highly accurate for identifying AAS and provides interpretable results to assist in clinical decision-making. Performance was assessed in two major phases: a multi-center retrospective study (n = 20,750) and an exploration in real-world emergency scenarios (n = 137,525). In the multi-center cohort, DeepAAS achieved a mean area under the receiver operating characteristic curve of 0.958 (95% CI 0.950-0.967). In the real-world cohort, DeepAAS detected 109 AAS patients with misguided initial suspicion, achieving 92.6% (95% CI 76.2%-97.5%) in mean sensitivity and 99.2% (95% CI 99.1%-99.3%) in mean specificity. Our AI model performed well on non-contrast CT at all applicable early stages of differential diagnosis workflows, effectively reduced the overall missed diagnosis and misdiagnosis rate from 48.8% to 4.8% and shortened the diagnosis time for patients with misguided initial suspicion from an average of 681.8 (74-11,820) mins to 68.5 (23-195) mins. DeepAAS could effectively fill the gap in the current clinical workflow without requiring additional tests.
CycleINR: Cycle Implicit Neural Representation for Arbitrary-Scale Volumetric Super-Resolution of Medical Data
Apr 07, 2024



Abstract:In the realm of medical 3D data, such as CT and MRI images, prevalent anisotropic resolution is characterized by high intra-slice but diminished inter-slice resolution. The lowered resolution between adjacent slices poses challenges, hindering optimal viewing experiences and impeding the development of robust downstream analysis algorithms. Various volumetric super-resolution algorithms aim to surmount these challenges, enhancing inter-slice resolution and overall 3D medical imaging quality. However, existing approaches confront inherent challenges: 1) often tailored to specific upsampling factors, lacking flexibility for diverse clinical scenarios; 2) newly generated slices frequently suffer from over-smoothing, degrading fine details, and leading to inter-slice inconsistency. In response, this study presents CycleINR, a novel enhanced Implicit Neural Representation model for 3D medical data volumetric super-resolution. Leveraging the continuity of the learned implicit function, the CycleINR model can achieve results with arbitrary up-sampling rates, eliminating the need for separate training. Additionally, we enhance the grid sampling in CycleINR with a local attention mechanism and mitigate over-smoothing by integrating cycle-consistent loss. We introduce a new metric, Slice-wise Noise Level Inconsistency (SNLI), to quantitatively assess inter-slice noise level inconsistency. The effectiveness of our approach is demonstrated through image quality evaluations on an in-house dataset and a downstream task analysis on the Medical Segmentation Decathlon liver tumor dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge