Ling Zhang
Missouri S&T EMC Laboratory, Rolla, MO, USA
CT-FineBench: A Diagnostic Fidelity Benchmark for Fine-Grained Evaluation of CT Report Generation
Apr 27, 2026Abstract:The evaluation of generated reports remains a critical challenge in Computed Tomography (CT) report generation, due to the large volume of text, the diversity and complexity of findings, and the presence of fine-grained, disease-oriented attributes. Conventional evaluation metrics offer only coarse measures of lexical overlap or entity matching and fail to reflect the granular diagnostic accuracy required for clinical use. To address this gap, we propose CT-FineBench, a benchmark built from CT-RATE and Merlin to evaluate the fine-grained factual consistency of CT reports, constructed from CT-RATE and Merlin. Our benchmark is constructed through a meticulous, Question-Answering (QA) based process: first, we identify and structure key, finding-specific clinical attributes (like location, size, margin). Second, we systematically transform these attributes into a QA dataset, where questions probe for specific clinical details grounded in gold-standard reports. The evaluation protocol for CT-FineBench involves using this QA dataset to query a machine-generated report and scoring the correctness of the answers. This allows for a comprehensive, interpretable, and clinically-relevant assessment, moving beyond superficial lexical overlap to pinpoint specific clinical errors. Experiments show that CT-FineBench correlates better with expert clinical assessment and is substantially more sensitive to fine-grained factual errors than prior metrics.
Dictionary-based Pathology Mining with Hard-instance-assisted Classifier Debiasing for Genetic Biomarker Prediction from WSIs
Mar 26, 2026Abstract:Prediction of genetic biomarkers, e.g., microsatellite instability in colorectal cancer is crucial for clinical decision making. But, two primary challenges hamper accurate prediction: (1) It is difficult to construct a pathology-aware representation involving the complex interconnections among pathological components. (2) WSIs contain a large proportion of areas unrelated to genetic biomarkers, which make the model easily overfit simple but irrelative instances. We hereby propose a Dictionary-based hierarchical pathology mining with hard-instance-assisted classifier Debiasing framework to address these challenges, dubbed as D2Bio. Our first module, dictionary-based hierarchical pathology mining, is able to mine diverse and very fine-grained pathological contextual interaction without the limit to the distances between patches. The second module, hard-instance-assisted classfier debiasing, learns a debiased classifier via focusing on hard but task-related features, without any additional annotations. Experimental results on five cohorts show the superiority of our method, with over 4% improvement in AUROC compared with the second best on the TCGA-CRC-MSI cohort. Our analysis further shows the clinical interpretability of D2Bio in genetic biomarker diagnosis and potential clinical utility in survival analysis. Code will be available at https://github.com/DeepMed-Lab-ECNU/D2Bio.
Rethinking the Efficiency and Effectiveness of Reinforcement Learning for Radiology Report Generation
Mar 04, 2026Abstract:Radiologists highly desire fully automated AI for radiology report generation (R2G), yet existing approaches fall short in clinical utility. Reinforcement learning (RL) holds potential to address these shortcomings, but its adoption in this task remains underexplored. In this paper, we revisit RL in terms of data efficiency and optimization effectiveness for R2G tasks. First, we explore the impact of data quantity and quality on the performance of RL in medical contexts, revealing that data quality plays a more critical role than quantity. To this end, we propose a diagnostic diversity-based data sampling strategy that enables comparable performance with fewer samples. Second, we observe that the majority of tokens in radiology reports are template-like and diagnostically uninformative, whereas the low frequency of clinically critical tokens heightens the risk of being overlooked during optimization. To tackle this, we introduce Diagnostic Token-weighted Policy Optimization (DiTPO), which directly optimizes for clinical accuracy by using a diagnostic F1 score as the reward signal. Unlike standard RL approaches that treat all tokens equally, DiTPO explicitly models the varying importance of different tokens through rule- or gradient-based mechanisms to prioritize clinically relevant content. Extensive experiments on the MIMIC-CXR, IU-Xray, and CheXpert Plus datasets demonstrate that our framework achieves state-of-the-art (SOTA) performance while requiring substantially fewer training samples in RL. Notably, on MIMIC-CXR, our framework attains an F1 score of 0.516 using only 20% of the RL training samples.
OmniCT: Towards a Unified Slice-Volume LVLM for Comprehensive CT Analysis
Feb 18, 2026Abstract:Computed Tomography (CT) is one of the most widely used and diagnostically information-dense imaging modalities, covering critical organs such as the heart, lungs, liver, and colon. Clinical interpretation relies on both slice-driven local features (e.g., sub-centimeter nodules, lesion boundaries) and volume-driven spatial representations (e.g., tumor infiltration, inter-organ anatomical relations). However, existing Large Vision-Language Models (LVLMs) remain fragmented in CT slice versus volumetric understanding: slice-driven LVLMs show strong generalization but lack cross-slice spatial consistency, while volume-driven LVLMs explicitly capture volumetric semantics but suffer from coarse granularity and poor compatibility with slice inputs. The absence of a unified modeling paradigm constitutes a major bottleneck for the clinical translation of medical LVLMs. We present OmniCT, a powerful unified slice-volume LVLM for CT scenarios, which makes three contributions: (i) Spatial Consistency Enhancement (SCE): volumetric slice composition combined with tri-axial positional embedding that introduces volumetric consistency, and an MoE hybrid projection enables efficient slice-volume adaptation; (ii) Organ-level Semantic Enhancement (OSE): segmentation and ROI localization explicitly align anatomical regions, emphasizing lesion- and organ-level semantics; (iii) MedEval-CT: the largest slice-volume CT dataset and hybrid benchmark integrates comprehensive metrics for unified evaluation. OmniCT consistently outperforms existing methods with a substantial margin across diverse clinical tasks and satisfies both micro-level detail sensitivity and macro-level spatial reasoning. More importantly, it establishes a new paradigm for cross-modal medical imaging understanding.
Non-Contrast CT Esophageal Varices Grading through Clinical Prior-Enhanced Multi-Organ Analysis
Dec 22, 2025Abstract:Esophageal varices (EV) represent a critical complication of portal hypertension, affecting approximately 60% of cirrhosis patients with a significant bleeding risk of ~30%. While traditionally diagnosed through invasive endoscopy, non-contrast computed tomography (NCCT) presents a potential non-invasive alternative that has yet to be fully utilized in clinical practice. We present Multi-Organ-COhesion Network++ (MOON++), a novel multimodal framework that enhances EV assessment through comprehensive analysis of NCCT scans. Inspired by clinical evidence correlating organ volumetric relationships with liver disease severity, MOON++ synthesizes imaging characteristics of the esophagus, liver, and spleen through multimodal learning. We evaluated our approach using 1,631 patients, those with endoscopically confirmed EV were classified into four severity grades. Validation in 239 patient cases and independent testing in 289 cases demonstrate superior performance compared to conventional single organ methods, achieving an AUC of 0.894 versus 0.803 for the severe grade EV classification (G3 versus <G3) and 0.921 versus 0.793 for the differentiation of moderate to severe grades (>=G2 versus <G2). We conducted a reader study involving experienced radiologists to further validate the performance of MOON++. To our knowledge, MOON++ represents the first comprehensive multi-organ NCCT analysis framework incorporating clinical knowledge priors for EV assessment, potentially offering a promising non-invasive diagnostic alternative.
Holdout-Loss-Based Data Selection for LLM Finetuning via In-Context Learning
Oct 16, 2025Abstract:Fine-tuning large pretrained language models is a common approach for aligning them with human preferences, but noisy or off-target examples can dilute supervision. While small, well-chosen datasets often match the performance of much larger ones, systematic and efficient ways to identify high-value training data remain underexplored. Many current methods rely on heuristics or expensive retraining. We present a theoretically grounded, resource-efficient framework for data selection and reweighting. At its core is an In-Context Approximation (ICA) that estimates the holdout loss a model would incur after training on a candidate example by conditioning on a small, curated holdout set in context. ICA requires no reference model and no additional finetuning. Under a local linearization, ICA is equivalent to a first-order update toward the holdout optimum, motivating its use as a proxy for data value. We derive per-example weights from ICA scores, dynamically reweighting gradient updates as model parameters evolve. Across SFT, DPO, and SimPO, and over diverse backbones and datasets, ICA-based reweighting consistently improves model alignment with minimal overhead. We analyze sensitivity to score update frequency and the choice of $k$ holdout examples for in-context demonstrations, and note limitations for rapidly drifting on-policy updates, highlighting directions for future work. Code and prompts will be released.
HeurAgenix: Leveraging LLMs for Solving Complex Combinatorial Optimization Challenges
Jun 18, 2025Abstract:Heuristic algorithms play a vital role in solving combinatorial optimization (CO) problems, yet traditional designs depend heavily on manual expertise and struggle to generalize across diverse instances. We introduce \textbf{HeurAgenix}, a two-stage hyper-heuristic framework powered by large language models (LLMs) that first evolves heuristics and then selects among them automatically. In the heuristic evolution phase, HeurAgenix leverages an LLM to compare seed heuristic solutions with higher-quality solutions and extract reusable evolution strategies. During problem solving, it dynamically picks the most promising heuristic for each problem state, guided by the LLM's perception ability. For flexibility, this selector can be either a state-of-the-art LLM or a fine-tuned lightweight model with lower inference cost. To mitigate the scarcity of reliable supervision caused by CO complexity, we fine-tune the lightweight heuristic selector with a dual-reward mechanism that jointly exploits singals from selection preferences and state perception, enabling robust selection under noisy annotations. Extensive experiments on canonical benchmarks show that HeurAgenix not only outperforms existing LLM-based hyper-heuristics but also matches or exceeds specialized solvers. Code is available at https://github.com/microsoft/HeurAgenix.
Leveraging Interview-Informed LLMs to Model Survey Responses: Comparative Insights from AI-Generated and Human Data
May 28, 2025
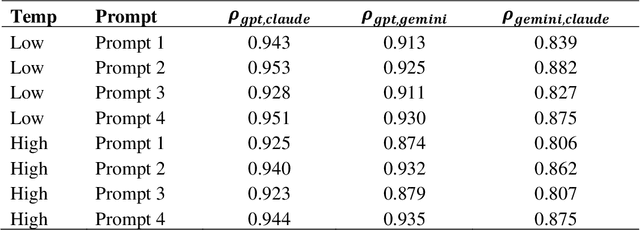


Abstract:Mixed methods research integrates quantitative and qualitative data but faces challenges in aligning their distinct structures, particularly in examining measurement characteristics and individual response patterns. Advances in large language models (LLMs) offer promising solutions by generating synthetic survey responses informed by qualitative data. This study investigates whether LLMs, guided by personal interviews, can reliably predict human survey responses, using the Behavioral Regulations in Exercise Questionnaire (BREQ) and interviews from after-school program staff as a case study. Results indicate that LLMs capture overall response patterns but exhibit lower variability than humans. Incorporating interview data improves response diversity for some models (e.g., Claude, GPT), while well-crafted prompts and low-temperature settings enhance alignment between LLM and human responses. Demographic information had less impact than interview content on alignment accuracy. These findings underscore the potential of interview-informed LLMs to bridge qualitative and quantitative methodologies while revealing limitations in response variability, emotional interpretation, and psychometric fidelity. Future research should refine prompt design, explore bias mitigation, and optimize model settings to enhance the validity of LLM-generated survey data in social science research.
Large-scale and Fine-grained Vision-language Pre-training for Enhanced CT Image Understanding
Jan 24, 2025



Abstract:Artificial intelligence (AI) shows great potential in assisting radiologists to improve the efficiency and accuracy of medical image interpretation and diagnosis. However, a versatile AI model requires large-scale data and comprehensive annotations, which are often impractical in medical settings. Recent studies leverage radiology reports as a naturally high-quality supervision for medical images, using contrastive language-image pre-training (CLIP) to develop language-informed models for radiological image interpretation. Nonetheless, these approaches typically contrast entire images with reports, neglecting the local associations between imaging regions and report sentences, which may undermine model performance and interoperability. In this paper, we propose a fine-grained vision-language model (fVLM) for anatomy-level CT image interpretation. Specifically, we explicitly match anatomical regions of CT images with corresponding descriptions in radiology reports and perform contrastive pre-training for each anatomy individually. Fine-grained alignment, however, faces considerable false-negative challenges, mainly from the abundance of anatomy-level healthy samples and similarly diseased abnormalities. To tackle this issue, we propose identifying false negatives of both normal and abnormal samples and calibrating contrastive learning from patient-level to disease-aware pairing. We curated the largest CT dataset to date, comprising imaging and report data from 69,086 patients, and conducted a comprehensive evaluation of 54 major and important disease diagnosis tasks across 15 main anatomies. Experimental results demonstrate the substantial potential of fVLM in versatile medical image interpretation. In the zero-shot classification task, we achieved an average AUC of 81.3% on 54 diagnosis tasks, surpassing CLIP and supervised methods by 12.9% and 8.0%, respectively.
Doc-Guided Sent2Sent++: A Sent2Sent++ Agent with Doc-Guided memory for Document-level Machine Translation
Jan 15, 2025



Abstract:The field of artificial intelligence has witnessed significant advancements in natural language processing, largely attributed to the capabilities of Large Language Models (LLMs). These models form the backbone of Agents designed to address long-context dependencies, particularly in Document-level Machine Translation (DocMT). DocMT presents unique challenges, with quality, consistency, and fluency being the key metrics for evaluation. Existing approaches, such as Doc2Doc and Doc2Sent, either omit sentences or compromise fluency. This paper introduces Doc-Guided Sent2Sent++, an Agent that employs an incremental sentence-level forced decoding strategy \textbf{to ensure every sentence is translated while enhancing the fluency of adjacent sentences.} Our Agent leverages a Doc-Guided Memory, focusing solely on the summary and its translation, which we find to be an efficient approach to maintaining consistency. Through extensive testing across multiple languages and domains, we demonstrate that Sent2Sent++ outperforms other methods in terms of quality, consistency, and fluency. The results indicate that, our approach has achieved significant improvements in metrics such as s-COMET, d-COMET, LTCR-$1_f$, and document-level perplexity (d-ppl). The contributions of this paper include a detailed analysis of current DocMT research, the introduction of the Sent2Sent++ decoding method, the Doc-Guided Memory mechanism, and validation of its effectiveness across languages and domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge