Ruizhe Guo
2L-LSH: A Locality-Sensitive Hash Function-Based Method For Rapid Point Cloud Indexing
Apr 23, 2026Abstract:The development of 3D scanning technology has enabled the acquisition of massive point cloud models with diverse structures and large scales, thereby presenting significant challenges in point cloud processing. Fast neighboring points search is one of the most common problems, which is frequently used in model reconstruction, classification, retrieval and feature visualization. Hash function is well known for its high-speed and accurate performance in searching high-dimensional data, which is also the core of the proposed 2L-LSH. Specifically, the 2L-LSH algorithm adopts a two-step hash function strategy, in which the popular step divides the bounding box of the point cloud model and the second step constructs a generalized table-based data structure. The proposed 2L-LSH offers a highly efficient and accurate solution for fast neighboring points search in large-scale 3D point cloud models, making it a promising technique for various applications in the field. The proposed algorithm is compared with the well-known methods including Kd-tree and Octree; the obtained results demonstrated that the proposed method outperforms Kd-tree and Octree in terms of speed, i.e. the time consumption of kNN search can be 51.111% and 94.159% lower than Kd-tree and Octree, respectively. And the RN search time can be 54.519% and 41.840% lower than Kd-tree and Octree, respectively.
* 13 pages, 13 figures. Published in The Computer Journal
Towards Effective and Efficient Context-aware Nucleus Detection in Histopathology Whole Slide Images
Mar 04, 2025
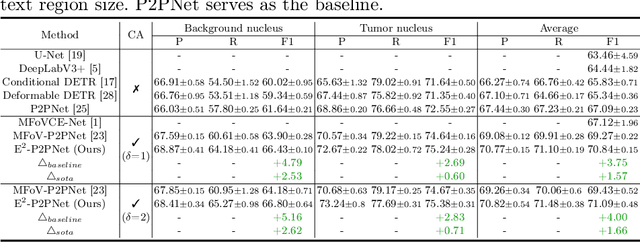
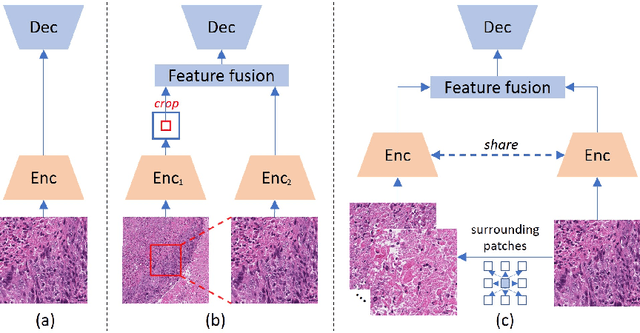
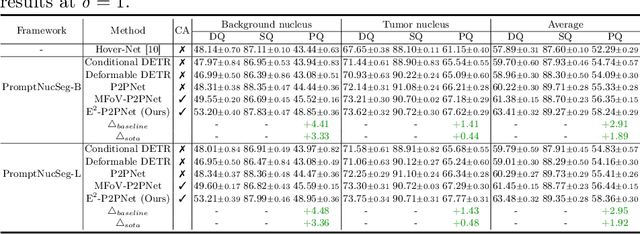
Abstract:Nucleus detection in histopathology whole slide images (WSIs) is crucial for a broad spectrum of clinical applications. Current approaches for nucleus detection in gigapixel WSIs utilize a sliding window methodology, which overlooks boarder contextual information (eg, tissue structure) and easily leads to inaccurate predictions. To address this problem, recent studies additionally crops a large Filed-of-View (FoV) region around each sliding window to extract contextual features. However, such methods substantially increases the inference latency. In this paper, we propose an effective and efficient context-aware nucleus detection algorithm. Specifically, instead of leveraging large FoV regions, we aggregate contextual clues from off-the-shelf features of historically visited sliding windows. This design greatly reduces computational overhead. Moreover, compared to large FoV regions at a low magnification, the sliding window patches have higher magnification and provide finer-grained tissue details, thereby enhancing the detection accuracy. To further improve the efficiency, we propose a grid pooling technique to compress dense feature maps of each patch into a few contextual tokens. Finally, we craft OCELOT-seg, the first benchmark dedicated to context-aware nucleus instance segmentation. Code, dataset, and model checkpoints will be available at https://github.com/windygoo/PathContext.
Large-scale and Fine-grained Vision-language Pre-training for Enhanced CT Image Understanding
Jan 24, 2025



Abstract:Artificial intelligence (AI) shows great potential in assisting radiologists to improve the efficiency and accuracy of medical image interpretation and diagnosis. However, a versatile AI model requires large-scale data and comprehensive annotations, which are often impractical in medical settings. Recent studies leverage radiology reports as a naturally high-quality supervision for medical images, using contrastive language-image pre-training (CLIP) to develop language-informed models for radiological image interpretation. Nonetheless, these approaches typically contrast entire images with reports, neglecting the local associations between imaging regions and report sentences, which may undermine model performance and interoperability. In this paper, we propose a fine-grained vision-language model (fVLM) for anatomy-level CT image interpretation. Specifically, we explicitly match anatomical regions of CT images with corresponding descriptions in radiology reports and perform contrastive pre-training for each anatomy individually. Fine-grained alignment, however, faces considerable false-negative challenges, mainly from the abundance of anatomy-level healthy samples and similarly diseased abnormalities. To tackle this issue, we propose identifying false negatives of both normal and abnormal samples and calibrating contrastive learning from patient-level to disease-aware pairing. We curated the largest CT dataset to date, comprising imaging and report data from 69,086 patients, and conducted a comprehensive evaluation of 54 major and important disease diagnosis tasks across 15 main anatomies. Experimental results demonstrate the substantial potential of fVLM in versatile medical image interpretation. In the zero-shot classification task, we achieved an average AUC of 81.3% on 54 diagnosis tasks, surpassing CLIP and supervised methods by 12.9% and 8.0%, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge