Yike Guo
Conservation-informed Graph Learning for Spatiotemporal Dynamics Prediction
Dec 30, 2024



Abstract:Data-centric methods have shown great potential in understanding and predicting spatiotemporal dynamics, enabling better design and control of the object system. However, pure deep learning models often lack interpretability, fail to obey intrinsic physics, and struggle to cope with the various domains. While geometry-based methods, e.g., graph neural networks (GNNs), have been proposed to further tackle these challenges, they still need to find the implicit physical laws from large datasets and rely excessively on rich labeled data. In this paper, we herein introduce the conservation-informed GNN (CiGNN), an end-to-end explainable learning framework, to learn spatiotemporal dynamics based on limited training data. The network is designed to conform to the general conservation law via symmetry, where conservative and non-conservative information passes over a multiscale space enhanced by a latent temporal marching strategy. The efficacy of our model has been verified in various spatiotemporal systems based on synthetic and real-world datasets, showing superiority over baseline models. Results demonstrate that CiGNN exhibits remarkable accuracy and generalization ability, and is readily applicable to learning for prediction of various spatiotemporal dynamics in a spatial domain with complex geometry.
SINGER: Vivid Audio-driven Singing Video Generation with Multi-scale Spectral Diffusion Model
Dec 04, 2024Abstract:Recent advancements in generative models have significantly enhanced talking face video generation, yet singing video generation remains underexplored. The differences between human talking and singing limit the performance of existing talking face video generation models when applied to singing. The fundamental differences between talking and singing-specifically in audio characteristics and behavioral expressions-limit the effectiveness of existing models. We observe that the differences between singing and talking audios manifest in terms of frequency and amplitude. To address this, we have designed a multi-scale spectral module to help the model learn singing patterns in the spectral domain. Additionally, we develop a spectral-filtering module that aids the model in learning the human behaviors associated with singing audio. These two modules are integrated into the diffusion model to enhance singing video generation performance, resulting in our proposed model, SINGER. Furthermore, the lack of high-quality real-world singing face videos has hindered the development of the singing video generation community. To address this gap, we have collected an in-the-wild audio-visual singing dataset to facilitate research in this area. Our experiments demonstrate that SINGER is capable of generating vivid singing videos and outperforms state-of-the-art methods in both objective and subjective evaluations.
Fire-Image-DenseNet (FIDN) for predicting wildfire burnt area using remote sensing data
Dec 02, 2024



Abstract:Predicting the extent of massive wildfires once ignited is essential to reduce the subsequent socioeconomic losses and environmental damage, but challenging because of the complexity of fire behaviour. Existing physics-based models are limited in predicting large or long-duration wildfire events. Here, we develop a deep-learning-based predictive model, Fire-Image-DenseNet (FIDN), that uses spatial features derived from both near real-time and reanalysis data on the environmental and meteorological drivers of wildfire. We trained and tested this model using more than 300 individual wildfires that occurred between 2012 and 2019 in the western US. In contrast to existing models, the performance of FIDN does not degrade with fire size or duration. Furthermore, it predicts final burnt area accurately even in very heterogeneous landscapes in terms of fuel density and flammability. The FIDN model showed higher accuracy, with a mean squared error (MSE) about 82% and 67% lower than those of the predictive models based on cellular automata (CA) and the minimum travel time (MTT) approaches, respectively. Its structural similarity index measure (SSIM) averages 97%, outperforming the CA and FlamMap MTT models by 6% and 2%, respectively. Additionally, FIDN is approximately three orders of magnitude faster than both CA and MTT models. The enhanced computational efficiency and accuracy advancements offer vital insights for strategic planning and resource allocation for firefighting operations.
Foundation Cures Personalization: Recovering Facial Personalized Models' Prompt Consistency
Nov 22, 2024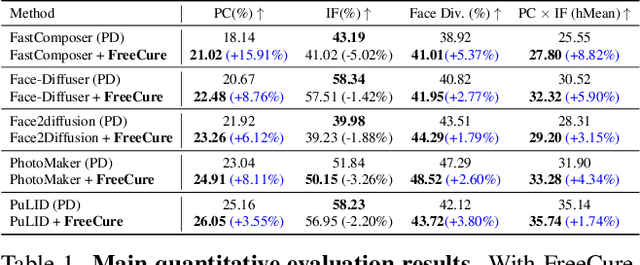
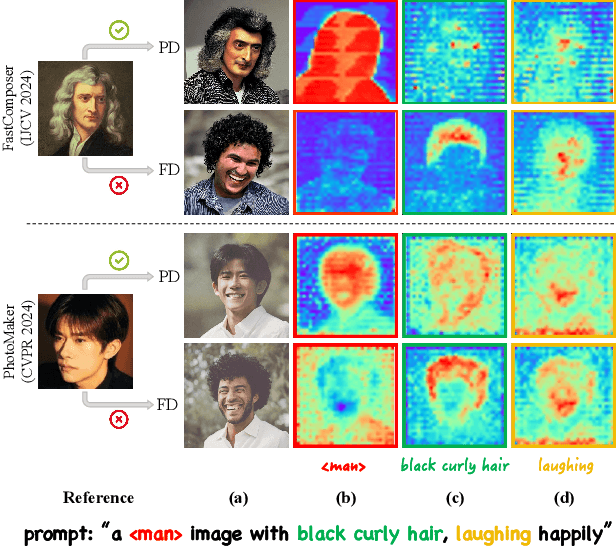

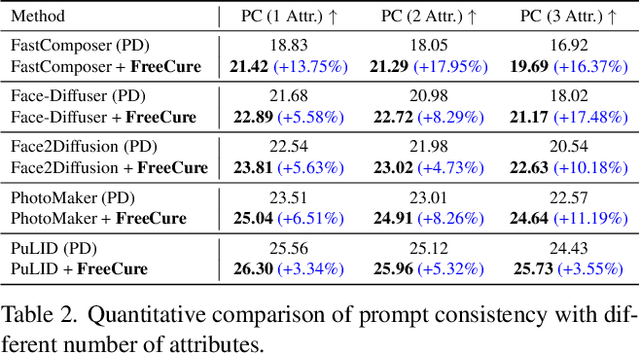

Abstract:Facial personalization represents a crucial downstream task in the domain of text-to-image generation. To preserve identity fidelity while ensuring alignment with user-defined prompts, current mainstream frameworks for facial personalization predominantly employ identity embedding mechanisms to associate identity information with textual embeddings. However, our experiments show that identity embeddings compromise the effectiveness of other tokens within the prompt, thereby hindering high prompt consistency, particularly when prompts involve multiple facial attributes. Moreover, previous works overlook the fact that their corresponding foundation models hold great potential to generate faces aligning to prompts well and can be easily leveraged to cure these ill-aligned attributes in personalized models. Building upon these insights, we propose FreeCure, a training-free framework that harnesses the intrinsic knowledge from the foundation models themselves to improve the prompt consistency of personalization models. First, by extracting cross-attention and semantic maps from the denoising process of foundation models, we identify easily localized attributes (e.g., hair, accessories, etc). Second, we enhance multiple attributes in the outputs of personalization models through a novel noise-blending strategy coupled with an inversion-based process. Our approach offers several advantages: it eliminates the need for training; it effectively facilitates the enhancement for a wide array of facial attributes in a non-intrusive manner; and it can be seamlessly integrated into existing popular personalization models. FreeCure has demonstrated significant improvements in prompt consistency across a diverse set of state-of-the-art facial personalization models while maintaining the integrity of original identity fidelity.
EVA: An Embodied World Model for Future Video Anticipation
Oct 20, 2024



Abstract:World models integrate raw data from various modalities, such as images and language to simulate comprehensive interactions in the world, thereby displaying crucial roles in fields like mixed reality and robotics. Yet, applying the world model for accurate video prediction is quite challenging due to the complex and dynamic intentions of the various scenes in practice. In this paper, inspired by the human rethinking process, we decompose the complex video prediction into four meta-tasks that enable the world model to handle this issue in a more fine-grained manner. Alongside these tasks, we introduce a new benchmark named Embodied Video Anticipation Benchmark (EVA-Bench) to provide a well-rounded evaluation. EVA-Bench focused on evaluating the video prediction ability of human and robot actions, presenting significant challenges for both the language model and the generation model. Targeting embodied video prediction, we propose the Embodied Video Anticipator (EVA), a unified framework aiming at video understanding and generation. EVA integrates a video generation model with a visual language model, effectively combining reasoning capabilities with high-quality generation. Moreover, to enhance the generalization of our framework, we tailor-designed a multi-stage pretraining paradigm that adaptatively ensembles LoRA to produce high-fidelity results. Extensive experiments on EVA-Bench highlight the potential of EVA to significantly improve performance in embodied scenes, paving the way for large-scale pre-trained models in real-world prediction tasks.
Both Ears Wide Open: Towards Language-Driven Spatial Audio Generation
Oct 14, 2024



Abstract:Recently, diffusion models have achieved great success in mono-channel audio generation. However, when it comes to stereo audio generation, the soundscapes often have a complex scene of multiple objects and directions. Controlling stereo audio with spatial contexts remains challenging due to high data costs and unstable generative models. To the best of our knowledge, this work represents the first attempt to address these issues. We first construct a large-scale, simulation-based, and GPT-assisted dataset, BEWO-1M, with abundant soundscapes and descriptions even including moving and multiple sources. Beyond text modality, we have also acquired a set of images and rationally paired stereo audios through retrieval to advance multimodal generation. Existing audio generation models tend to generate rather random and indistinct spatial audio. To provide accurate guidance for latent diffusion models, we introduce the SpatialSonic model utilizing spatial-aware encoders and azimuth state matrices to reveal reasonable spatial guidance. By leveraging spatial guidance, our unified model not only achieves the objective of generating immersive and controllable spatial audio from text and image but also enables interactive audio generation during inference. Finally, under fair settings, we conduct subjective and objective evaluations on simulated and real-world data to compare our approach with prevailing methods. The results demonstrate the effectiveness of our method, highlighting its capability to generate spatial audio that adheres to physical rules.
You Know What I'm Saying -- Jailbreak Attack via Implicit Reference
Oct 04, 2024



Abstract:While recent advancements in large language model (LLM) alignment have enabled the effective identification of malicious objectives involving scene nesting and keyword rewriting, our study reveals that these methods remain inadequate at detecting malicious objectives expressed through context within nested harmless objectives. This study identifies a previously overlooked vulnerability, which we term Attack via Implicit Reference (AIR). AIR decomposes a malicious objective into permissible objectives and links them through implicit references within the context. This method employs multiple related harmless objectives to generate malicious content without triggering refusal responses, thereby effectively bypassing existing detection techniques.Our experiments demonstrate AIR's effectiveness across state-of-the-art LLMs, achieving an attack success rate (ASR) exceeding 90% on most models, including GPT-4o, Claude-3.5-Sonnet, and Qwen-2-72B. Notably, we observe an inverse scaling phenomenon, where larger models are more vulnerable to this attack method. These findings underscore the urgent need for defense mechanisms capable of understanding and preventing contextual attacks. Furthermore, we introduce a cross-model attack strategy that leverages less secure models to generate malicious contexts, thereby further increasing the ASR when targeting other models.Our code and jailbreak artifacts can be found at https://github.com/Lucas-TY/llm_Implicit_reference.
PSHuman: Photorealistic Single-view Human Reconstruction using Cross-Scale Diffusion
Sep 16, 2024



Abstract:Detailed and photorealistic 3D human modeling is essential for various applications and has seen tremendous progress. However, full-body reconstruction from a monocular RGB image remains challenging due to the ill-posed nature of the problem and sophisticated clothing topology with self-occlusions. In this paper, we propose PSHuman, a novel framework that explicitly reconstructs human meshes utilizing priors from the multiview diffusion model. It is found that directly applying multiview diffusion on single-view human images leads to severe geometric distortions, especially on generated faces. To address it, we propose a cross-scale diffusion that models the joint probability distribution of global full-body shape and local facial characteristics, enabling detailed and identity-preserved novel-view generation without any geometric distortion. Moreover, to enhance cross-view body shape consistency of varied human poses, we condition the generative model on parametric models like SMPL-X, which provide body priors and prevent unnatural views inconsistent with human anatomy. Leveraging the generated multi-view normal and color images, we present SMPLX-initialized explicit human carving to recover realistic textured human meshes efficiently. Extensive experimental results and quantitative evaluations on CAPE and THuman2.1 datasets demonstrate PSHumans superiority in geometry details, texture fidelity, and generalization capability.
HiPrompt: Tuning-free Higher-Resolution Generation with Hierarchical MLLM Prompts
Sep 04, 2024



Abstract:The potential for higher-resolution image generation using pretrained diffusion models is immense, yet these models often struggle with issues of object repetition and structural artifacts especially when scaling to 4K resolution and higher. We figure out that the problem is caused by that, a single prompt for the generation of multiple scales provides insufficient efficacy. In response, we propose HiPrompt, a new tuning-free solution that tackles the above problems by introducing hierarchical prompts. The hierarchical prompts offer both global and local guidance. Specifically, the global guidance comes from the user input that describes the overall content, while the local guidance utilizes patch-wise descriptions from MLLMs to elaborately guide the regional structure and texture generation. Furthermore, during the inverse denoising process, the generated noise is decomposed into low- and high-frequency spatial components. These components are conditioned on multiple prompt levels, including detailed patch-wise descriptions and broader image-level prompts, facilitating prompt-guided denoising under hierarchical semantic guidance. It further allows the generation to focus more on local spatial regions and ensures the generated images maintain coherent local and global semantics, structures, and textures with high definition. Extensive experiments demonstrate that HiPrompt outperforms state-of-the-art works in higher-resolution image generation, significantly reducing object repetition and enhancing structural quality.
Deep learning surrogate models of JULES-INFERNO for wildfire prediction on a global scale
Aug 30, 2024



Abstract:Global wildfire models play a crucial role in anticipating and responding to changing wildfire regimes. JULES-INFERNO is a global vegetation and fire model simulating wildfire emissions and area burnt on a global scale. However, because of the high data dimensionality and system complexity, JULES-INFERNO's computational costs make it challenging to apply to fire risk forecasting with unseen initial conditions. Typically, running JULES-INFERNO for 30 years of prediction will take several hours on High Performance Computing (HPC) clusters. To tackle this bottleneck, two data-driven models are built in this work based on Deep Learning techniques to surrogate the JULES-INFERNO model and speed up global wildfire forecasting. More precisely, these machine learning models take global temperature, vegetation density, soil moisture and previous forecasts as inputs to predict the subsequent global area burnt on an iterative basis. Average Error per Pixel (AEP) and Structural Similarity Index Measure (SSIM) are used as metrics to evaluate the performance of the proposed surrogate models. A fine tuning strategy is also proposed in this work to improve the algorithm performance for unseen scenarios. Numerical results show a strong performance of the proposed models, in terms of both computational efficiency (less than 20 seconds for 30 years of prediction on a laptop CPU) and prediction accuracy (with AEP under 0.3\% and SSIM over 98\% compared to the outputs of JULES-INFERNO).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge