Sibo Cheng
Anchored-Branched Steady-state WInd Flow Transformer (AB-SWIFT): a metamodel for 3D atmospheric flow in urban environments
Mar 26, 2026Abstract:Air flow modeling at a local scale is essential for applications such as pollutant dispersion modeling or wind farm modeling. To circumvent costly Computational Fluid Dynamics (CFD) computations, deep learning surrogate models have recently emerged as promising alternatives. However, in the context of urban air flow, deep learning models struggle to adapt to the high variations of the urban geometry and to large mesh sizes. To tackle these challenges, we introduce Anchored Branched Steady-state WInd Flow Transformer (AB-SWIFT), a transformer-based model with an internal branched structure uniquely designed for atmospheric flow modeling. We train our model on a specially designed database of atmospheric simulations around randomised urban geometries and with a mixture of unstable, neutral, and stable atmospheric stratifications. Our model reaches the best accuracy on all predicted fields compared to state-of-the-art transformers and graph-based models. Our code and data is available at https://github.com/cerea-daml/abswift.
Spatiotemporal System Forecasting with Irregular Time Steps via Masked Autoencoder
Mar 26, 2026Abstract:Predicting high-dimensional dynamical systems with irregular time steps presents significant challenges for current data-driven algorithms. These irregularities arise from missing data, sparse observations, or adaptive computational techniques, reducing prediction accuracy. To address these limitations, we propose a novel method: a Physics-Spatiotemporal Masked Autoencoder. This method integrates convolutional autoencoders for spatial feature extraction with masked autoencoders optimised for irregular time series, leveraging attention mechanisms to reconstruct the entire physical sequence in a single prediction pass. The model avoids the need for data imputation while preserving physical integrity of the system. Here, 'physics' refers to high-dimensional fields generated by underlying dynamical systems, rather than the enforcement of explicit physical constraints or PDE residuals. We evaluate this approach on multiple simulated datasets and real-world ocean temperature data. The results demonstrate that our method achieves significant improvements in prediction accuracy, robustness to nonlinearities, and computational efficiency over traditional convolutional and recurrent network methods. The model shows potential for capturing complex spatiotemporal patterns without requiring domain-specific knowledge, with applications in climate modelling, fluid dynamics, ocean forecasting, environmental monitoring, and scientific computing.
Rooftop Wind Field Reconstruction Using Sparse Sensors: From Deterministic to Generative Learning Methods
Mar 13, 2026Abstract:Real-time rooftop wind-speed distribution is important for the safe operation of drones and urban air mobility systems, wind control systems, and rooftop utilization. However, rooftop flows show strong nonlinearity, separation, and cross-direction variability, which make flow field reconstruction from sparse sensors difficult. This study develops a learning-from-observation framework using wind-tunnel experimental data obtained by Particle Image Velocimetry (PIV) and compares Kriging interpolation with three deep learning models: UNet, Vision Transformer Autoencoder (ViTAE), and Conditional Wasserstein GAN (CWGAN). We evaluate two training strategies, single wind-direction training (SDT) and mixed wind-direction training (MDT), across sensor densities from 5 to 30, test robustness under sensor position perturbations of plus or minus 1 grid, and optimize sensor placement via Proper Orthogonal Decomposition with QR decomposition. Results show that deep learning methods can reconstruct rooftop wind fields from sparse sensor data effectively. Compared with Kriging interpolation, the deep learning models improved SSIM by up to 32.7%, FAC2 by 24.2%, and NMSE by 27.8%. Mixed wind-direction training further improved performance, with gains of up to 173.7% in SSIM, 16.7% in FAC2, and 98.3% in MG compared with single-direction training. The results also show that sensor configuration, optimization, and training strategy should be considered jointly for reliable deployment. QR-based optimization improved robustness by up to 27.8% under sensor perturbations, although with metric-dependent trade-offs. Training on experimental rather than simulated data also provides practical guidance for method selection and sensor placement in different scenarios.
Trustworthy Data-Driven Wildfire Risk Prediction and Understanding in Western Canada
Jan 04, 2026Abstract:In recent decades, the intensification of wildfire activity in western Canada has resulted in substantial socio-economic and environmental losses. Accurate wildfire risk prediction is hindered by the intrinsic stochasticity of ignition and spread and by nonlinear interactions among fuel conditions, meteorology, climate variability, topography, and human activities, challenging the reliability and interpretability of purely data-driven models. We propose a trustworthy data-driven wildfire risk prediction framework based on long-sequence, multi-scale temporal modeling, which integrates heterogeneous drivers while explicitly quantifying predictive uncertainty and enabling process-level interpretation. Evaluated over western Canada during the record-breaking 2023 and 2024 fire seasons, the proposed model outperforms existing time-series approaches, achieving an F1 score of 0.90 and a PR-AUC of 0.98 with low computational cost. Uncertainty-aware analysis reveals structured spatial and seasonal patterns in predictive confidence, highlighting increased uncertainty associated with ambiguous predictions and spatiotemporal decision boundaries. SHAP-based interpretation provides mechanistic understanding of wildfire controls, showing that temperature-related drivers dominate wildfire risk in both years, while moisture-related constraints play a stronger role in shaping spatial and land-cover-specific contrasts in 2024 compared to the widespread hot and dry conditions of 2023. Data and code are available at https://github.com/SynUW/mmFire.
Fast-Forward Lattice Boltzmann: Learning Kinetic Behaviour with Physics-Informed Neural Operators
Sep 26, 2025


Abstract:The lattice Boltzmann equation (LBE), rooted in kinetic theory, provides a powerful framework for capturing complex flow behaviour by describing the evolution of single-particle distribution functions (PDFs). Despite its success, solving the LBE numerically remains computationally intensive due to strict time-step restrictions imposed by collision kernels. Here, we introduce a physics-informed neural operator framework for the LBE that enables prediction over large time horizons without step-by-step integration, effectively bypassing the need to explicitly solve the collision kernel. We incorporate intrinsic moment-matching constraints of the LBE, along with global equivariance of the full distribution field, enabling the model to capture the complex dynamics of the underlying kinetic system. Our framework is discretization-invariant, enabling models trained on coarse lattices to generalise to finer ones (kinetic super-resolution). In addition, it is agnostic to the specific form of the underlying collision model, which makes it naturally applicable across different kinetic datasets regardless of the governing dynamics. Our results demonstrate robustness across complex flow scenarios, including von Karman vortex shedding, ligament breakup, and bubble adhesion. This establishes a new data-driven pathway for modelling kinetic systems.
Comparative and Interpretative Analysis of CNN and Transformer Models in Predicting Wildfire Spread Using Remote Sensing Data
Mar 18, 2025



Abstract:Facing the escalating threat of global wildfires, numerous computer vision techniques using remote sensing data have been applied in this area. However, the selection of deep learning methods for wildfire prediction remains uncertain due to the lack of comparative analysis in a quantitative and explainable manner, crucial for improving prevention measures and refining models. This study aims to thoroughly compare the performance, efficiency, and explainability of four prevalent deep learning architectures: Autoencoder, ResNet, UNet, and Transformer-based Swin-UNet. Employing a real-world dataset that includes nearly a decade of remote sensing data from California, U.S., these models predict the spread of wildfires for the following day. Through detailed quantitative comparison analysis, we discovered that Transformer-based Swin-UNet and UNet generally outperform Autoencoder and ResNet, particularly due to the advanced attention mechanisms in Transformer-based Swin-UNet and the efficient use of skip connections in both UNet and Transformer-based Swin-UNet, which contribute to superior predictive accuracy and model interpretability. Then we applied XAI techniques on all four models, this not only enhances the clarity and trustworthiness of models but also promotes focused improvements in wildfire prediction capabilities. The XAI analysis reveals that UNet and Transformer-based Swin-UNet are able to focus on critical features such as 'Previous Fire Mask', 'Drought', and 'Vegetation' more effectively than the other two models, while also maintaining balanced attention to the remaining features, leading to their superior performance. The insights from our thorough comparative analysis offer substantial implications for future model design and also provide guidance for model selection in different scenarios.
Machine learning for modelling unstructured grid data in computational physics: a review
Feb 13, 2025
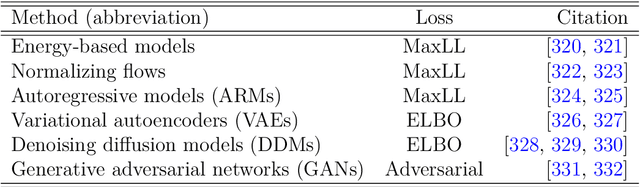
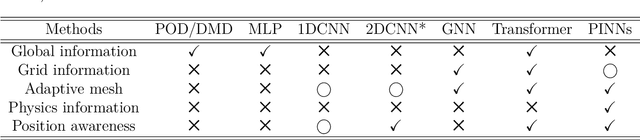
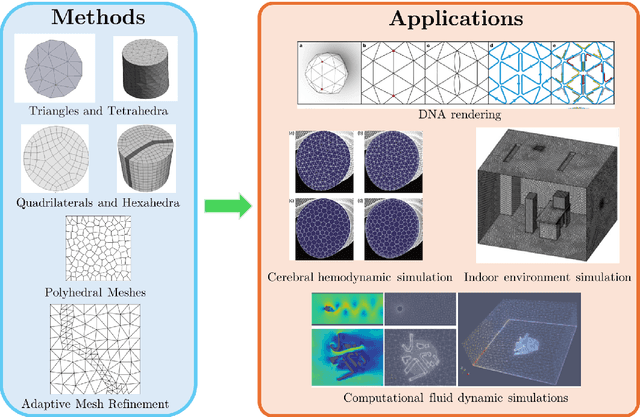

Abstract:Unstructured grid data are essential for modelling complex geometries and dynamics in computational physics. Yet, their inherent irregularity presents significant challenges for conventional machine learning (ML) techniques. This paper provides a comprehensive review of advanced ML methodologies designed to handle unstructured grid data in high-dimensional dynamical systems. Key approaches discussed include graph neural networks, transformer models with spatial attention mechanisms, interpolation-integrated ML methods, and meshless techniques such as physics-informed neural networks. These methodologies have proven effective across diverse fields, including fluid dynamics and environmental simulations. This review is intended as a guidebook for computational scientists seeking to apply ML approaches to unstructured grid data in their domains, as well as for ML researchers looking to address challenges in computational physics. It places special focus on how ML methods can overcome the inherent limitations of traditional numerical techniques and, conversely, how insights from computational physics can inform ML development. To support benchmarking, this review also provides a summary of open-access datasets of unstructured grid data in computational physics. Finally, emerging directions such as generative models with unstructured data, reinforcement learning for mesh generation, and hybrid physics-data-driven paradigms are discussed to inspire future advancements in this evolving field.
Fire-Image-DenseNet (FIDN) for predicting wildfire burnt area using remote sensing data
Dec 02, 2024



Abstract:Predicting the extent of massive wildfires once ignited is essential to reduce the subsequent socioeconomic losses and environmental damage, but challenging because of the complexity of fire behaviour. Existing physics-based models are limited in predicting large or long-duration wildfire events. Here, we develop a deep-learning-based predictive model, Fire-Image-DenseNet (FIDN), that uses spatial features derived from both near real-time and reanalysis data on the environmental and meteorological drivers of wildfire. We trained and tested this model using more than 300 individual wildfires that occurred between 2012 and 2019 in the western US. In contrast to existing models, the performance of FIDN does not degrade with fire size or duration. Furthermore, it predicts final burnt area accurately even in very heterogeneous landscapes in terms of fuel density and flammability. The FIDN model showed higher accuracy, with a mean squared error (MSE) about 82% and 67% lower than those of the predictive models based on cellular automata (CA) and the minimum travel time (MTT) approaches, respectively. Its structural similarity index measure (SSIM) averages 97%, outperforming the CA and FlamMap MTT models by 6% and 2%, respectively. Additionally, FIDN is approximately three orders of magnitude faster than both CA and MTT models. The enhanced computational efficiency and accuracy advancements offer vital insights for strategic planning and resource allocation for firefighting operations.
Dynamical system prediction from sparse observations using deep neural networks with Voronoi tessellation and physics constraint
Aug 31, 2024



Abstract:Despite the success of various methods in addressing the issue of spatial reconstruction of dynamical systems with sparse observations, spatio-temporal prediction for sparse fields remains a challenge. Existing Kriging-based frameworks for spatio-temporal sparse field prediction fail to meet the accuracy and inference time required for nonlinear dynamic prediction problems. In this paper, we introduce the Dynamical System Prediction from Sparse Observations using Voronoi Tessellation (DSOVT) framework, an innovative methodology based on Voronoi tessellation which combines convolutional encoder-decoder (CED) and long short-term memory (LSTM) and utilizing Convolutional Long Short-Term Memory (ConvLSTM). By integrating Voronoi tessellations with spatio-temporal deep learning models, DSOVT is adept at predicting dynamical systems with unstructured, sparse, and time-varying observations. CED-LSTM maps Voronoi tessellations into a low-dimensional representation for time series prediction, while ConvLSTM directly uses these tessellations in an end-to-end predictive model. Furthermore, we incorporate physics constraints during the training process for dynamical systems with explicit formulas. Compared to purely data-driven models, our physics-based approach enables the model to learn physical laws within explicitly formulated dynamics, thereby enhancing the robustness and accuracy of rolling forecasts. Numerical experiments on real sea surface data and shallow water systems clearly demonstrate our framework's accuracy and computational efficiency with sparse and time-varying observations.
TorchDA: A Python package for performing data assimilation with deep learning forward and transformation functions
Aug 30, 2024Abstract:Data assimilation techniques are often confronted with challenges handling complex high dimensional physical systems, because high precision simulation in complex high dimensional physical systems is computationally expensive and the exact observation functions that can be applied in these systems are difficult to obtain. It prompts growing interest in integrating deep learning models within data assimilation workflows, but current software packages for data assimilation cannot handle deep learning models inside. This study presents a novel Python package seamlessly combining data assimilation with deep neural networks to serve as models for state transition and observation functions. The package, named TorchDA, implements Kalman Filter, Ensemble Kalman Filter (EnKF), 3D Variational (3DVar), and 4D Variational (4DVar) algorithms, allowing flexible algorithm selection based on application requirements. Comprehensive experiments conducted on the Lorenz 63 and a two-dimensional shallow water system demonstrate significantly enhanced performance over standalone model predictions without assimilation. The shallow water analysis validates data assimilation capabilities mapping between different physical quantity spaces in either full space or reduced order space. Overall, this innovative software package enables flexible integration of deep learning representations within data assimilation, conferring a versatile tool to tackle complex high dimensional dynamical systems across scientific domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge