Mitko Veta
Deep Learning-Based Grading of Ductal Carcinoma In Situ in Breast Histopathology Images
Oct 07, 2020
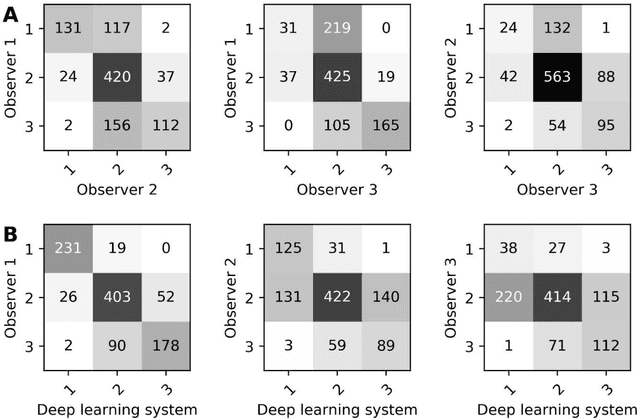
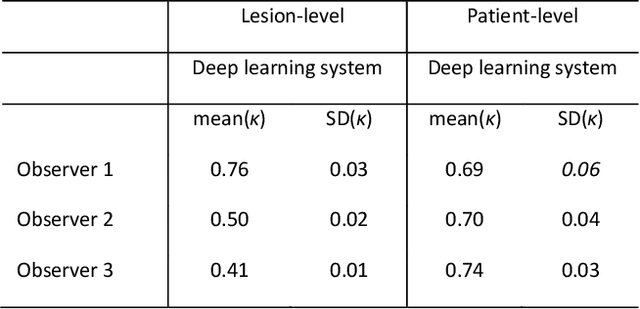
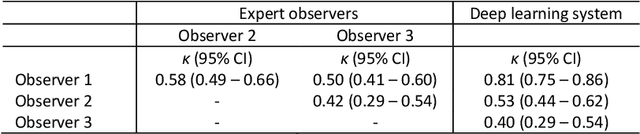
Abstract:Ductal carcinoma in situ (DCIS) is a non-invasive breast cancer that can progress into invasive ductal carcinoma (IDC). Studies suggest DCIS is often overtreated since a considerable part of DCIS lesions may never progress into IDC. Lower grade lesions have a lower progression speed and risk, possibly allowing treatment de-escalation. However, studies show significant inter-observer variation in DCIS grading. Automated image analysis may provide an objective solution to address high subjectivity of DCIS grading by pathologists. In this study, we developed a deep learning-based DCIS grading system. It was developed using the consensus DCIS grade of three expert observers on a dataset of 1186 DCIS lesions from 59 patients. The inter-observer agreement, measured by quadratic weighted Cohen's kappa, was used to evaluate the system and compare its performance to that of expert observers. We present an analysis of the lesion-level and patient-level inter-observer agreement on an independent test set of 1001 lesions from 50 patients. The deep learning system (dl) achieved on average slightly higher inter-observer agreement to the observers (o1, o2 and o3) ($\kappa_{o1,dl}=0.81, \kappa_{o2,dl}=0.53, \kappa_{o3,dl}=0.40$) than the observers amongst each other ($\kappa_{o1,o2}=0.58, \kappa_{o1,o3}=0.50, \kappa_{o2,o3}=0.42$) at the lesion-level. At the patient-level, the deep learning system achieved similar agreement to the observers ($\kappa_{o1,dl}=0.77, \kappa_{o2,dl}=0.75, \kappa_{o3,dl}=0.70$) as the observers amongst each other ($\kappa_{o1,o2}=0.77, \kappa_{o1,o3}=0.75, \kappa_{o2,o3}=0.72$). In conclusion, we developed a deep learning-based DCIS grading system that achieved a performance similar to expert observers. We believe this is the first automated system that could assist pathologists by providing robust and reproducible second opinions on DCIS grade.
Domain-Adversarial Learning for Multi-Centre, Multi-Vendor, and Multi-Disease Cardiac MR Image Segmentation
Aug 26, 2020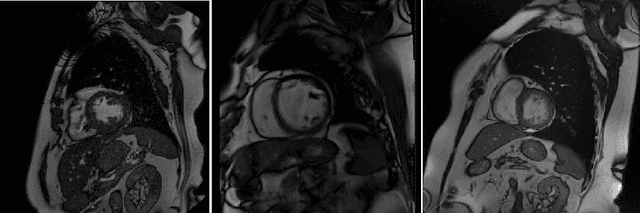
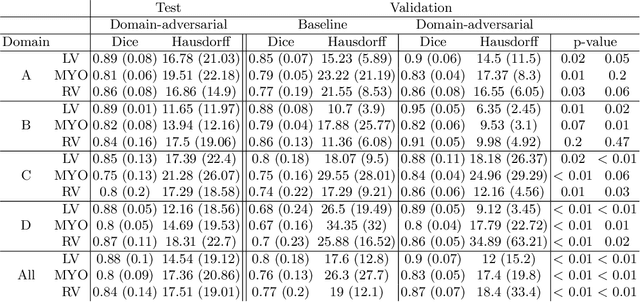
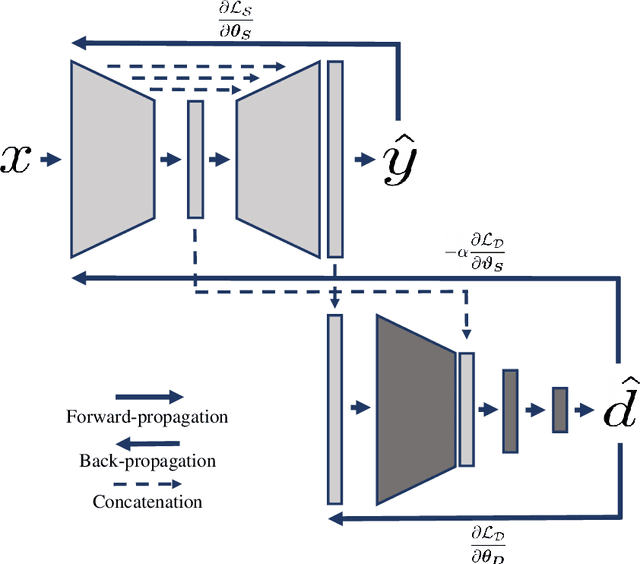
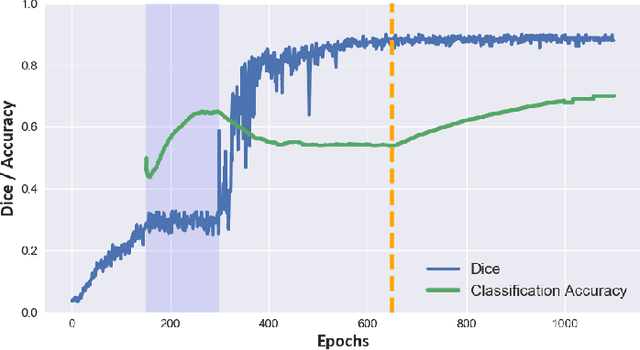
Abstract:Cine cardiac magnetic resonance (CMR) has become the gold standard for the non-invasive evaluation of cardiac function. In particular, it allows the accurate quantification of functional parameters including the chamber volumes and ejection fraction. Deep learning has shown the potential to automate the requisite cardiac structure segmentation. However, the lack of robustness of deep learning models has hindered their widespread clinical adoption. Due to differences in the data characteristics, neural networks trained on data from a specific scanner are not guaranteed to generalise well to data acquired at a different centre or with a different scanner. In this work, we propose a principled solution to the problem of this domain shift. Domain-adversarial learning is used to train a domain-invariant 2D U-Net using labelled and unlabelled data. This approach is evaluated on both seen and unseen domains from the M\&Ms challenge dataset and the domain-adversarial approach shows improved performance as compared to standard training. Additionally, we show that the domain information cannot be recovered from the learned features.
Orientation-Disentangled Unsupervised Representation Learning for Computational Pathology
Aug 26, 2020
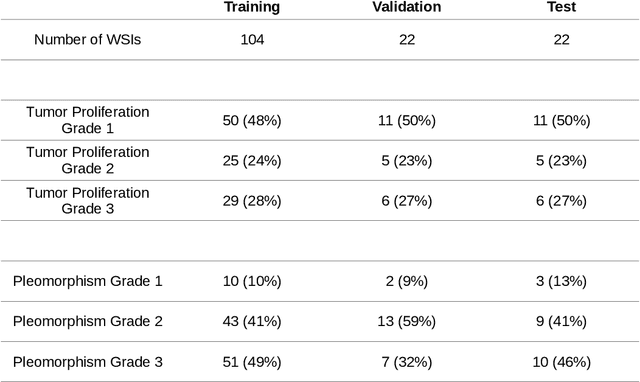
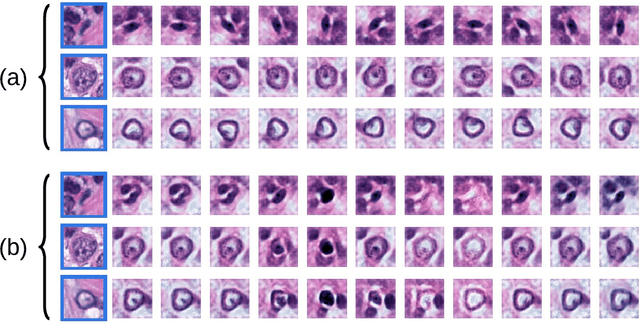
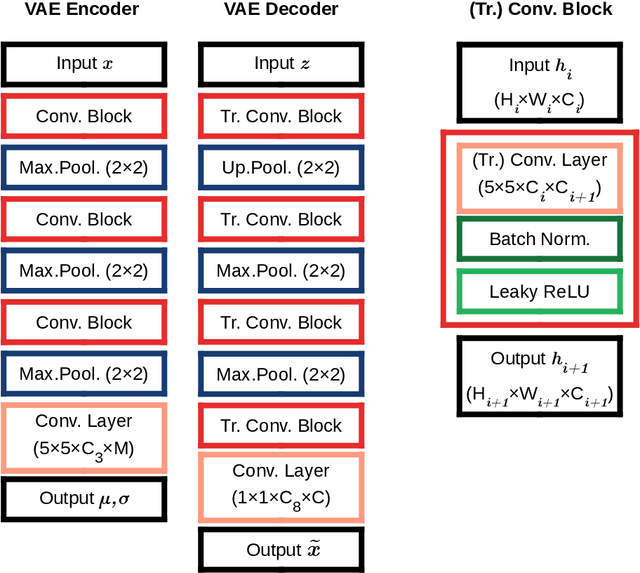
Abstract:Unsupervised learning enables modeling complex images without the need for annotations. The representation learned by such models can facilitate any subsequent analysis of large image datasets. However, some generative factors that cause irrelevant variations in images can potentially get entangled in such a learned representation causing the risk of negatively affecting any subsequent use. The orientation of imaged objects, for instance, is often arbitrary/irrelevant, thus it can be desired to learn a representation in which the orientation information is disentangled from all other factors. Here, we propose to extend the Variational Auto-Encoder framework by leveraging the group structure of rotation-equivariant convolutional networks to learn orientation-wise disentangled generative factors of histopathology images. This way, we enforce a novel partitioning of the latent space, such that oriented and isotropic components get separated. We evaluated this structured representation on a dataset that consists of tissue regions for which nuclear pleomorphism and mitotic activity was assessed by expert pathologists. We show that the trained models efficiently disentangle the inherent orientation information of single-cell images. In comparison to classical approaches, the resulting aggregated representation of sub-populations of cells produces higher performances in subsequent tasks.
Are pathologist-defined labels reproducible? Comparison of the TUPAC16 mitotic figure dataset with an alternative set of labels
Jul 10, 2020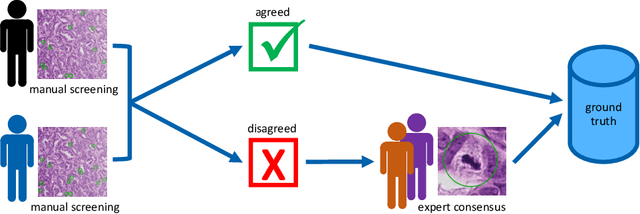
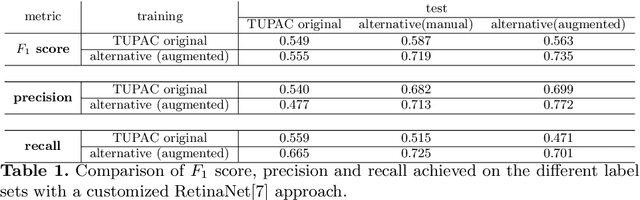
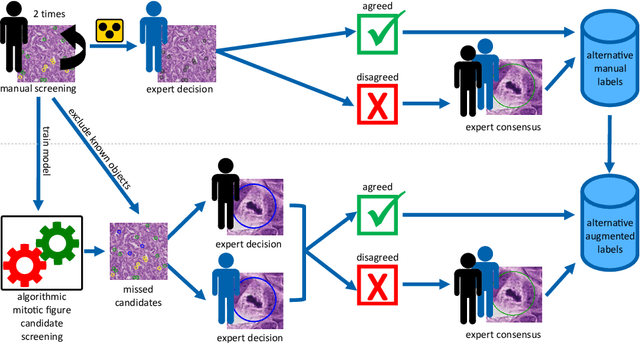
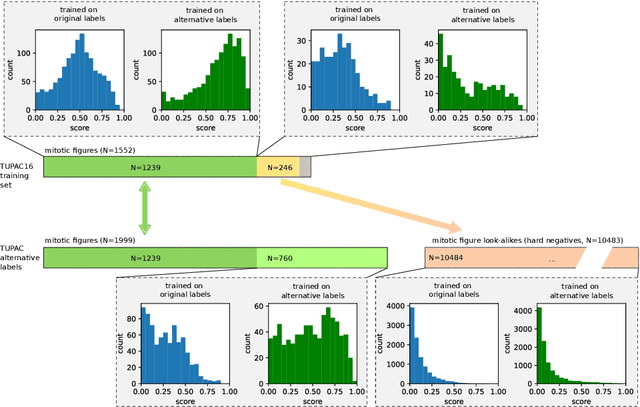
Abstract:Pathologist-defined labels are the gold standard for histopathological data sets, regardless of well-known limitations in consistency for some tasks. To date, some datasets on mitotic figures are available and were used for development of promising deep learning-based algorithms. In order to assess robustness of those algorithms and reproducibility of their methods it is necessary to test on several independent datasets. The influence of different labeling methods of these available datasets is currently unknown. To tackle this, we present an alternative set of labels for the images of the auxiliary mitosis dataset of the TUPAC16 challenge. Additional to manual mitotic figure screening, we used a novel, algorithm-aided labeling process, that allowed to minimize the risk of missing rare mitotic figures in the images. All potential mitotic figures were independently assessed by two pathologists. The novel, publicly available set of labels contains 1,999 mitotic figures (+28.80%) and additionally includes 10,483 labels of cells with high similarities to mitotic figures (hard examples). We found significant difference comparing F_1 scores between the original label set (0.549) and the new alternative label set (0.735) using a standard deep learning object detection architecture. The models trained on the alternative set showed higher overall confidence values, suggesting a higher overall label consistency. Findings of the present study show that pathologists-defined labels may vary significantly resulting in notable difference in the model performance. Comparison of deep learning-based algorithms between independent datasets with different labeling methods should be done with caution.
Adversarial Attack Vulnerability of Medical Image Analysis Systems: Unexplored Factors
Jun 12, 2020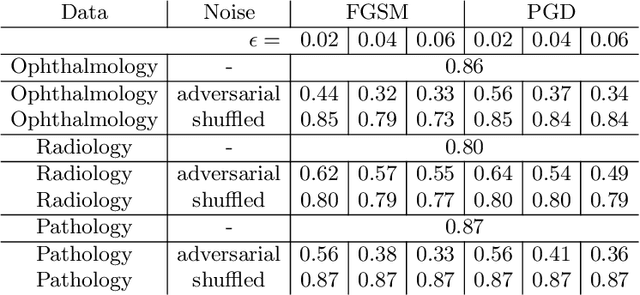
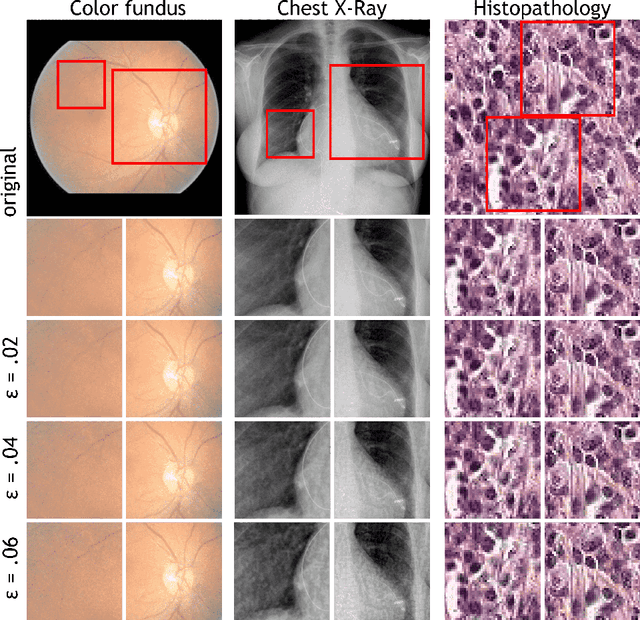
Abstract:Adversarial attacks are considered a potentially serious security threat for machine learning systems. Medical image analysis (MedIA) systems have recently been argued to be particularly vulnerable to adversarial attacks due to strong financial incentives. In this paper, we study several previously unexplored factors affecting adversarial attack vulnerability of deep learning MedIA systems in three medical domains: ophthalmology, radiology and pathology. Firstly, we study the effect of varying the degree of adversarial perturbation on the attack performance and its visual perceptibility. Secondly, we study how pre-training on a public dataset (ImageNet) affects the models' vulnerability to attacks. Thirdly, we study the influence of data and model architecture disparity between target and attacker models. Our experiments show that the degree of perturbation significantly affects both performance and human perceptibility of attacks. Pre-training may dramatically increase the transfer of adversarial examples; the larger the performance gain achieved by pre-training, the larger the transfer. Finally, disparity in data and/or model architecture between target and attacker models substantially decreases the success of attacks. We believe that these factors should be considered when designing cybersecurity-critical MedIA systems, as well as kept in mind when evaluating their vulnerability to adversarial attacks.
A Global Benchmark of Algorithms for Segmenting Late Gadolinium-Enhanced Cardiac Magnetic Resonance Imaging
May 07, 2020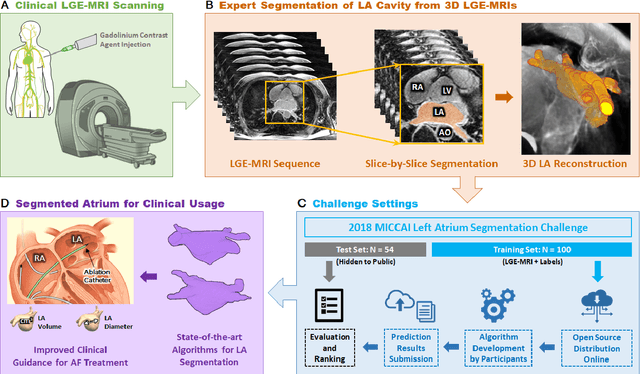
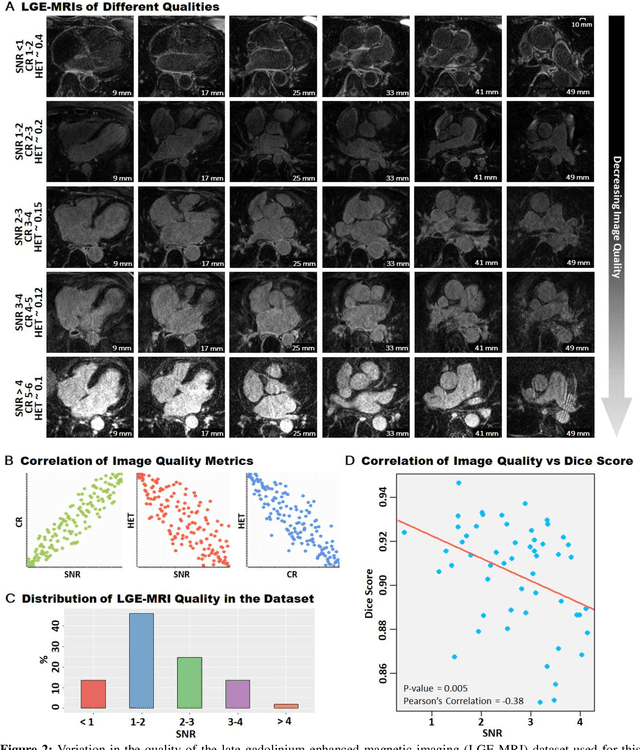
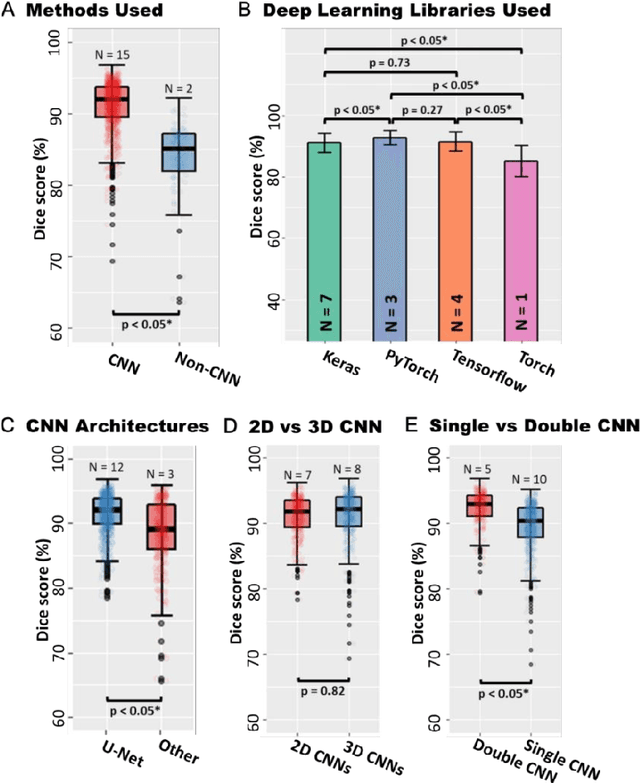
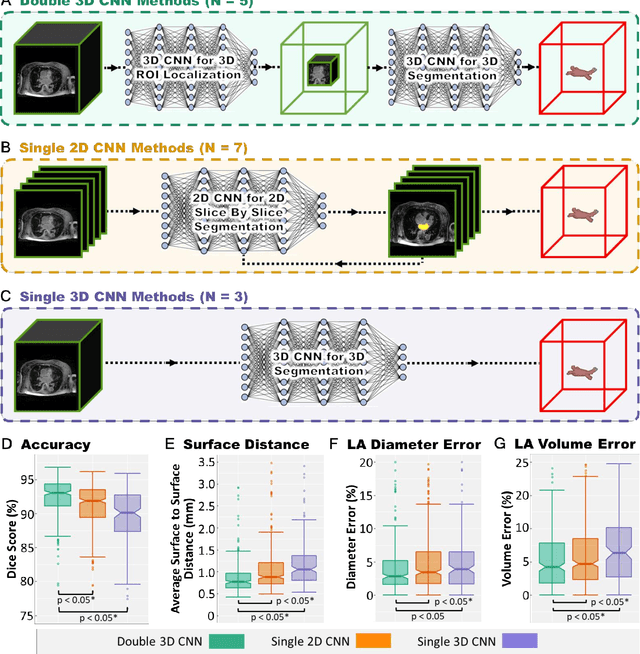
Abstract:Segmentation of cardiac images, particularly late gadolinium-enhanced magnetic resonance imaging (LGE-MRI) widely used for visualizing diseased cardiac structures, is a crucial first step for clinical diagnosis and treatment. However, direct segmentation of LGE-MRIs is challenging due to its attenuated contrast. Since most clinical studies have relied on manual and labor-intensive approaches, automatic methods are of high interest, particularly optimized machine learning approaches. To address this, we organized the "2018 Left Atrium Segmentation Challenge" using 154 3D LGE-MRIs, currently the world's largest cardiac LGE-MRI dataset, and associated labels of the left atrium segmented by three medical experts, ultimately attracting the participation of 27 international teams. In this paper, extensive analysis of the submitted algorithms using technical and biological metrics was performed by undergoing subgroup analysis and conducting hyper-parameter analysis, offering an overall picture of the major design choices of convolutional neural networks (CNNs) and practical considerations for achieving state-of-the-art left atrium segmentation. Results show the top method achieved a dice score of 93.2% and a mean surface to a surface distance of 0.7 mm, significantly outperforming prior state-of-the-art. Particularly, our analysis demonstrated that double, sequentially used CNNs, in which a first CNN is used for automatic region-of-interest localization and a subsequent CNN is used for refined regional segmentation, achieved far superior results than traditional methods and pipelines containing single CNNs. This large-scale benchmarking study makes a significant step towards much-improved segmentation methods for cardiac LGE-MRIs, and will serve as an important benchmark for evaluating and comparing the future works in the field.
Quantifying Graft Detachment after Descemet's Membrane Endothelial Keratoplasty with Deep Convolutional Neural Networks
Apr 24, 2020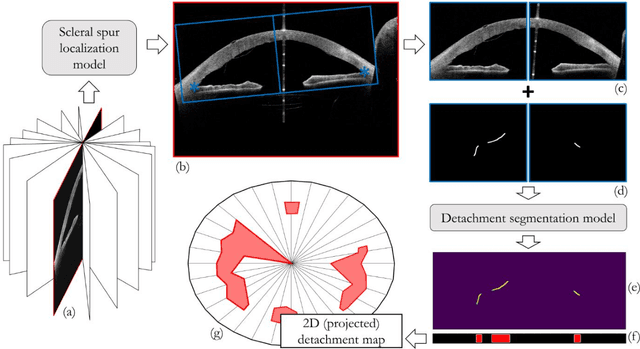
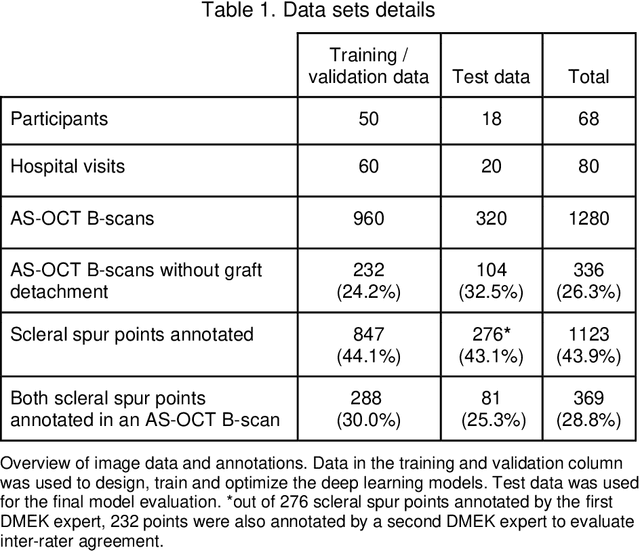
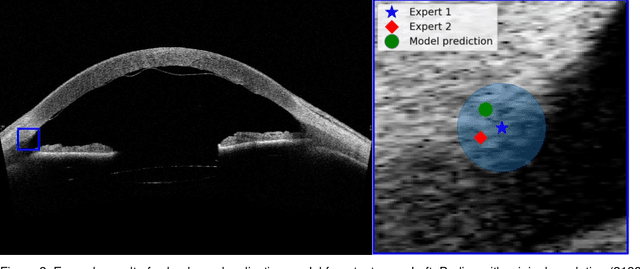
Abstract:Purpose: We developed a method to automatically locate and quantify graft detachment after Descemet's Membrane Endothelial Keratoplasty (DMEK) in Anterior Segment Optical Coherence Tomography (AS-OCT) scans. Methods: 1280 AS-OCT B-scans were annotated by a DMEK expert. Using the annotations, a deep learning pipeline was developed to localize scleral spur, center the AS-OCT B-scans and segment the detached graft sections. Detachment segmentation model performance was evaluated per B-scan by comparing (1) length of detachment and (2) horizontal projection of the detached sections with the expert annotations. Horizontal projections were used to construct graft detachment maps. All final evaluations were done on a test set that was set apart during training of the models. A second DMEK expert annotated the test set to determine inter-rater performance. Results: Mean scleral spur localization error was 0.155 mm, whereas the inter-rater difference was 0.090 mm. The estimated graft detachment lengths were in 69% of the cases within a 10-pixel (~150{\mu}m) difference from the ground truth (77% for the second DMEK expert). Dice scores for the horizontal projections of all B-scans with detachments were 0.896 and 0.880 for our model and the second DMEK expert respectively. Conclusion: Our deep learning model can be used to automatically and instantly localize graft detachment in AS-OCT B-scans. Horizontal detachment projections can be determined with the same accuracy as a human DMEK expert, allowing for the construction of accurate graft detachment maps. Translational Relevance: Automated localization and quantification of graft detachment can support DMEK research and standardize clinical decision making.
Roto-Translation Equivariant Convolutional Networks: Application to Histopathology Image Analysis
Feb 20, 2020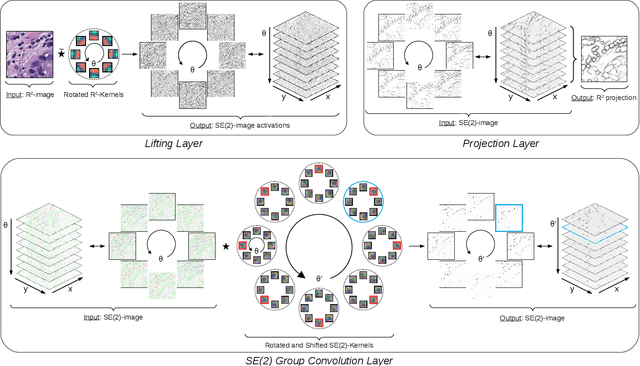
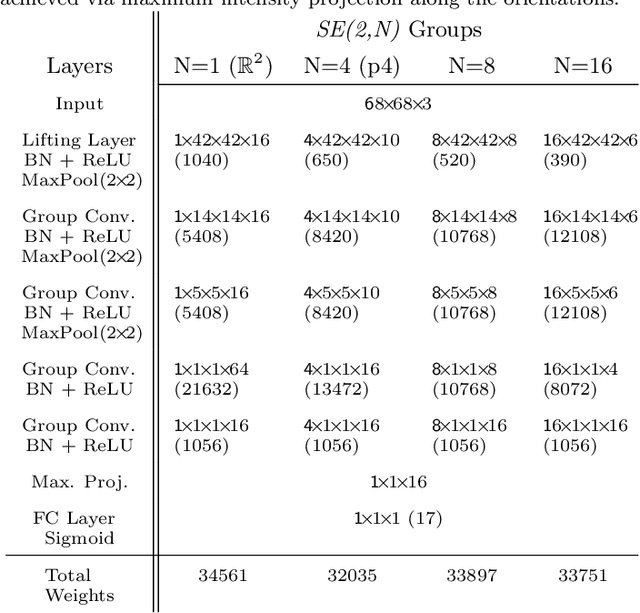
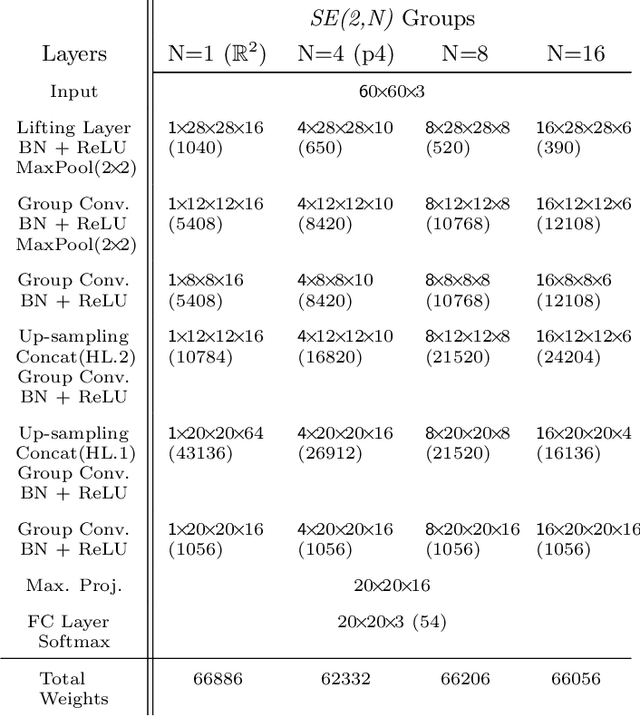
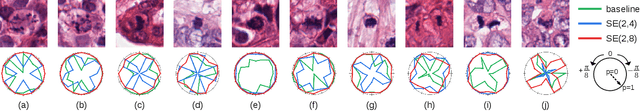
Abstract:Rotation-invariance is a desired property of machine-learning models for medical image analysis and in particular for computational pathology applications. We propose a framework to encode the geometric structure of the special Euclidean motion group SE(2) in convolutional networks to yield translation and rotation equivariance via the introduction of SE(2)-group convolution layers. This structure enables models to learn feature representations with a discretized orientation dimension that guarantees that their outputs are invariant under a discrete set of rotations. Conventional approaches for rotation invariance rely mostly on data augmentation, but this does not guarantee the robustness of the output when the input is rotated. At that, trained conventional CNNs may require test-time rotation augmentation to reach their full capability. This study is focused on histopathology image analysis applications for which it is desirable that the arbitrary global orientation information of the imaged tissues is not captured by the machine learning models. The proposed framework is evaluated on three different histopathology image analysis tasks (mitosis detection, nuclei segmentation and tumor classification). We present a comparative analysis for each problem and show that consistent increase of performances can be achieved when using the proposed framework.
Direct Classification of Type 2 Diabetes From Retinal Fundus Images in a Population-based Sample From The Maastricht Study
Nov 22, 2019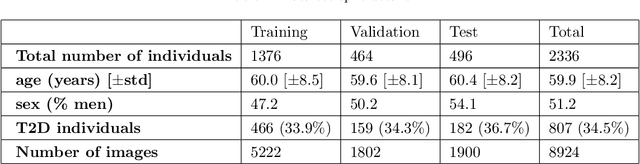
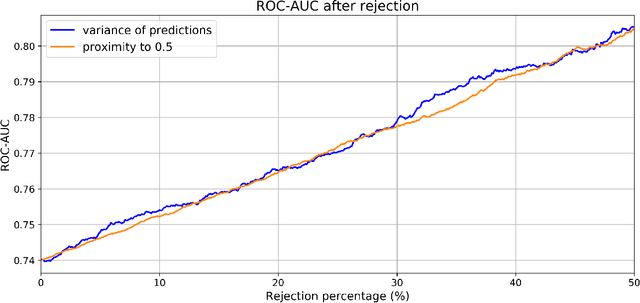
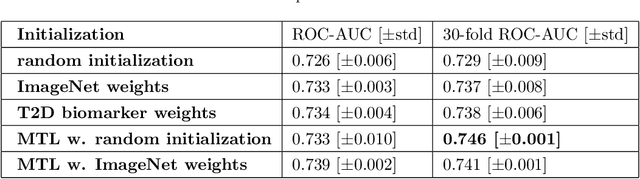

Abstract:Type 2 Diabetes (T2D) is a chronic metabolic disorder that can lead to blindness and cardiovascular disease. Information about early stage T2D might be present in retinal fundus images, but to what extent these images can be used for a screening setting is still unknown. In this study, deep neural networks were employed to differentiate between fundus images from individuals with and without T2D. We investigated three methods to achieve high classification performance, measured by the area under the receiver operating curve (ROC-AUC). A multi-target learning approach to simultaneously output retinal biomarkers as well as T2D works best (AUC = 0.746 [$\pm$0.001]). Furthermore, the classification performance can be improved when images with high prediction uncertainty are referred to a specialist. We also show that the combination of images of the left and right eye per individual can further improve the classification performance (AUC = 0.758 [$\pm$0.003]), using a simple averaging approach. The results are promising, suggesting the feasibility of screening for T2D from retinal fundus images.
Deep learning assessment of breast terminal duct lobular unit involution: towards automated prediction of breast cancer risk
Oct 31, 2019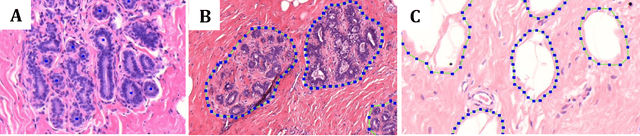
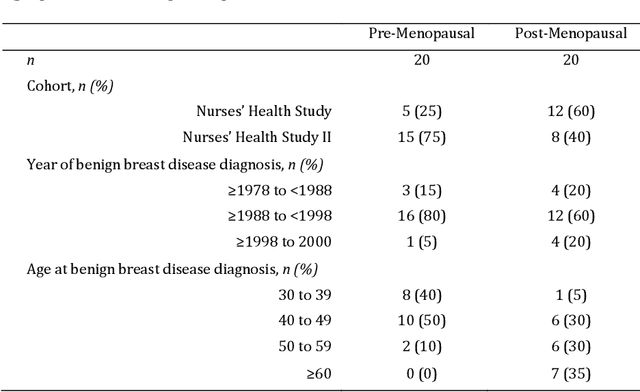
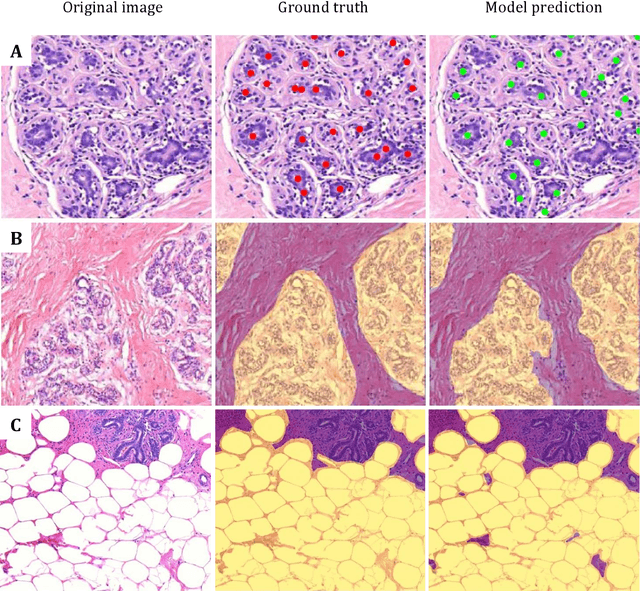
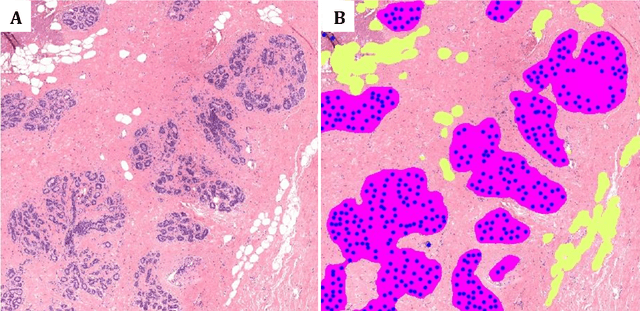
Abstract:Terminal ductal lobular unit (TDLU) involution is the regression of milk-producing structures in the breast. Women with less TDLU involution are more likely to develop breast cancer. A major bottleneck in studying TDLU involution in large cohort studies is the need for labor-intensive manual assessment of TDLUs. We developed a computational pathology solution to automatically capture TDLU involution measures. Whole slide images (WSIs) of benign breast biopsies were obtained from the Nurses' Health Study (NHS). A first set of 92 WSIs was annotated for TDLUs, acini and adipose tissue to train deep convolutional neural network (CNN) models for detection of acini, and segmentation of TDLUs and adipose tissue. These networks were integrated into a single computational method to capture TDLU involution measures including number of TDLUs per tissue area, median TDLU span and median number of acini per TDLU. We validated our method on 40 additional WSIs by comparing with manually acquired measures. Our CNN models detected acini with an F1 score of 0.73$\pm$0.09, and segmented TDLUs and adipose tissue with Dice scores of 0.86$\pm$0.11 and 0.86$\pm$0.04, respectively. The inter-observer ICC scores for manual assessments on 40 WSIs of number of TDLUs per tissue area, median TDLU span, and median acini count per TDLU were 0.71, 95% CI [0.51, 0.83], 0.81, 95% CI [0.67, 0.90], and 0.73, 95% CI [0.54, 0.85], respectively. Intra-observer reliability was evaluated on 10/40 WSIs with ICC scores of >0.8. Inter-observer ICC scores between automated results and the mean of the two observers were: 0.80, 95% CI [0.63, 0.90] for number of TDLUs per tissue area, 0.57, 95% CI [0.19, 0.77] for median TDLU span, and 0.80, 95% CI [0.62, 0.89] for median acini count per TDLU. TDLU involution measures evaluated by manual and automated assessment were inversely associated with age and menopausal status.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge