Wenjia Bai
Biomedical Image Analysis Group, Department of Computing, Imperial College London
Cardiac Mesh Flow: One-Step Generation of 3D+t Cardiac Four-Chamber Meshes via Flow Matching
May 03, 2026Abstract:Spatio-temporal (3D+t) generative modelling of cardiac shape and motion is crucial for understanding heart structure and function at population scale. Existing generative models for cardiac shape synthesis either adopt volumetric shape representations that lack anatomical correspondence across different time points and subjects, or rely on VAE-based frameworks that suffer from a trade-off between reconstruction fidelity and generative diversity. In this work, we propose Cardiac Mesh Flow, a novel generative flow model for 3D+t cardiac four-chamber mesh generation with anatomical correspondence, temporal coherence, and periodic consistency. Leveraging the flow matching technique, Cardiac Mesh Flow performs efficient one-step generation of multi-scale free-form deformation fields, which warp a template mesh to generate cardiac four-chamber meshes across a cardiac cycle. Furthermore, Cardiac Mesh Flow enables controllable generation conditioned on cardiac chamber volumes, allowing precise control of the synthetic heart. Experimental results demonstrate that Cardiac Mesh Flow achieves high fidelity and diversity on both unconditional and conditional generation, compared to state-of-the-art 3D+t cardiac mesh generation methods.
Dynamic Decision Learning: Test-Time Evolution for Abnormality Grounding in Rare Diseases
Apr 27, 2026Abstract:Clinical abnormality grounding for rare diseases is often hindered by data scarcity, making supervised fine-tuning impractical and single-pass inference highly unstable. We propose Dynamic Decision Learning (DDL), a framework that enables frozen large vision-language models (LVLMs) to refine their decisions across both language and visual spaces by optimizing instructions and consolidating predictions under visual perturbations. This process improves localization quality and produces a consensus-based reliability score that quantifies model confidence. Results on brain imaging benchmarks, including a rare-disease dataset with 281 pathology types across models ranging from 3B to 72B parameters, show that DDL improves mAP@75 by up to 105% on rare-disease cases and outperforms adaptation baselines and supervised fine-tuning. Furthermore, DDL demonstrates stronger calibration between reliability scores and localization accuracy under severe distribution shifts and increasing task difficulty. Code is available at: https://lijunrio.github.io/DDL/
AdamFlow: Adam-based Wasserstein Gradient Flows for Surface Registration in Medical Imaging
Apr 02, 2026Abstract:Surface registration plays an important role for anatomical shape analysis in medical imaging. Existing surface registration methods often face a trade-off between efficiency and robustness. Local point matching methods are computationally efficient, but vulnerable to noise and initialisation. Methods designed for global point set alignment tend to incur a high computational cost. To address the challenge, here we present a fast surface registration method, which formulates surface meshes as probability measures and surface registration as a distributional optimisation problem. The discrepancy between two meshes is measured using an efficient sliced Wasserstein distance with log-linear computational complexity. We propose a novel optimisation method, AdamFlow, which generalises the well-known Adam optimisation method from the Euclidean space to the probability space for minimising the sliced Wasserstein distance. We theoretically analyse the asymptotic convergence of AdamFlow and empirically demonstrate its superior performance in both affine and non-rigid surface registration across various anatomical structures.
Learning a dynamic four-chamber shape model of the human heart for 95,695 UK Biobank participants
Mar 30, 2026Abstract:The human heart is a sophisticated system composed of four cardiac chambers with distinct shapes, which function in a coordinated manner. Existing shape models of the heart mainly focus on the ventricular chambers and they are derived from relatively small datasets. Here, we present a spatio-temporal (3D+t) statistical shape model of all four cardiac chambers, learnt from a large population of nearly 100,000 participants from the UK Biobank. A deep learning-based pipeline is developed to reconstruct 3D+t four-chamber meshes from the cardiac magnetic resonance images of the UK Biobank imaging population. Based on the reconstructed meshes, a 3D+t statistical shape model is learnt to characterise the shape variations and motion patterns of the four cardiac chambers. We reveal the associations of the four-chamber shape model with demographics, anthropometrics, cardiovascular risk factors, and cardiac diseases. Compared to conventional image-derived phenotypes, we validate that the four-chamber shape-derived phenotypes significantly enhance the performance in downstream tasks, including cardiovascular disease classification and heart age prediction. Furthermore, we demonstrate the effectiveness of shape-derived phenotypes in novel applications such as heart shape retrieval and heart re-identification from longitudinal data. To facilitate future research, we will release the learning-based mesh reconstruction pipeline, the four-chamber cardiac shape model, and return all derived four-chamber meshes to the UK Biobank.
Flow Matching-enabled Test-Time Refinement for Unsupervised Cardiac MR Registration
Mar 03, 2026Abstract:Diffusion-based unsupervised image registration has been explored for cardiac cine MR, but expensive multi-step inference limits practical use. We propose FlowReg, a flow-matching framework in displacement field space that achieves strong registration in as few as two steps and supports further refinement with more steps. FlowReg uses warmup-reflow training: a single-step network first acts as a teacher, then a student learns to refine from arbitrary intermediate states, removing the need for a pre-trained model as in existing methods. An Initial Guess strategy feeds back the model prediction as the next starting point, improving refinement from step two onward. On ACDC and MM2 across six tasks (including cross-dataset generalization), FlowReg outperforms the state of the art on five tasks (+0.6% mean Dice score on average), with the largest gain in the left ventricle (+1.09%), and reduces LVEF estimation error on all six tasks (-2.58 percentage points), using only 0.7% extra parameters and no segmentation labels. Code is available at https://github.com/mathpluscode/FlowReg.
Enabling Ultra-Fast Cardiovascular Imaging Across Heterogeneous Clinical Environments with a Generalist Foundation Model and Multimodal Database
Dec 25, 2025Abstract:Multimodal cardiovascular magnetic resonance (CMR) imaging provides comprehensive and non-invasive insights into cardiovascular disease (CVD) diagnosis and underlying mechanisms. Despite decades of advancements, its widespread clinical adoption remains constrained by prolonged scan times and heterogeneity across medical environments. This underscores the urgent need for a generalist reconstruction foundation model for ultra-fast CMR imaging, one capable of adapting across diverse imaging scenarios and serving as the essential substrate for all downstream analyses. To enable this goal, we curate MMCMR-427K, the largest and most comprehensive multimodal CMR k-space database to date, comprising 427,465 multi-coil k-space data paired with structured metadata across 13 international centers, 12 CMR modalities, 15 scanners, and 17 CVD categories in populations across three continents. Building on this unprecedented resource, we introduce CardioMM, a generalist reconstruction foundation model capable of dynamically adapting to heterogeneous fast CMR imaging scenarios. CardioMM unifies semantic contextual understanding with physics-informed data consistency to deliver robust reconstructions across varied scanners, protocols, and patient presentations. Comprehensive evaluations demonstrate that CardioMM achieves state-of-the-art performance in the internal centers and exhibits strong zero-shot generalization to unseen external settings. Even at imaging acceleration up to 24x, CardioMM reliably preserves key cardiac phenotypes, quantitative myocardial biomarkers, and diagnostic image quality, enabling a substantial increase in CMR examination throughput without compromising clinical integrity. Together, our open-access MMCMR-427K database and CardioMM framework establish a scalable pathway toward high-throughput, high-quality, and clinically accessible cardiovascular imaging.
Multimodal Conditional MeshGAN for Personalized Aneurysm Growth Prediction
Aug 27, 2025Abstract:Personalized, accurate prediction of aortic aneurysm progression is essential for timely intervention but remains challenging due to the need to model both subtle local deformations and global anatomical changes within complex 3D geometries. We propose MCMeshGAN, the first multimodal conditional mesh-to-mesh generative adversarial network for 3D aneurysm growth prediction. MCMeshGAN introduces a dual-branch architecture combining a novel local KNN-based convolutional network (KCN) to preserve fine-grained geometric details and a global graph convolutional network (GCN) to capture long-range structural context, overcoming the over-smoothing limitations of deep GCNs. A dedicated condition branch encodes clinical attributes (age, sex) and the target time interval to generate anatomically plausible, temporally controlled predictions, enabling retrospective and prospective modeling. We curated TAAMesh, a new longitudinal thoracic aortic aneurysm mesh dataset consisting of 590 multimodal records (CT scans, 3D meshes, and clinical data) from 208 patients. Extensive experiments demonstrate that MCMeshGAN consistently outperforms state-of-the-art baselines in both geometric accuracy and clinically important diameter estimation. This framework offers a robust step toward clinically deployable, personalized 3D disease trajectory modeling. The source code for MCMeshGAN and the baseline methods is publicly available at https://github.com/ImperialCollegeLondon/MCMeshGAN.
Knowledge to Sight: Reasoning over Visual Attributes via Knowledge Decomposition for Abnormality Grounding
Aug 06, 2025Abstract:In this work, we address the problem of grounding abnormalities in medical images, where the goal is to localize clinical findings based on textual descriptions. While generalist Vision-Language Models (VLMs) excel in natural grounding tasks, they often struggle in the medical domain due to rare, compositional, and domain-specific terms that are poorly aligned with visual patterns. Specialized medical VLMs address this challenge via large-scale domain pretraining, but at the cost of substantial annotation and computational resources. To overcome these limitations, we propose \textbf{Knowledge to Sight (K2Sight)}, a framework that introduces structured semantic supervision by decomposing clinical concepts into interpretable visual attributes, such as shape, density, and anatomical location. These attributes are distilled from domain ontologies and encoded into concise instruction-style prompts, which guide region-text alignment during training. Unlike conventional report-level supervision, our approach explicitly bridges domain knowledge and spatial structure, enabling data-efficient training of compact models. We train compact models with 0.23B and 2B parameters using only 1.5\% of the data required by state-of-the-art medical VLMs. Despite their small size and limited training data, these models achieve performance on par with or better than 7B+ medical VLMs, with up to 9.82\% improvement in $mAP_{50}$. Code and models: \href{https://lijunrio.github.io/K2Sight/}{\textcolor{SOTAPink}{https://lijunrio.github.io/K2Sight/}}.
SAM-aware Test-time Adaptation for Universal Medical Image Segmentation
Jun 05, 2025



Abstract:Universal medical image segmentation using the Segment Anything Model (SAM) remains challenging due to its limited adaptability to medical domains. Existing adaptations, such as MedSAM, enhance SAM's performance in medical imaging but at the cost of reduced generalization to unseen data. Therefore, in this paper, we propose SAM-aware Test-Time Adaptation (SAM-TTA), a fundamentally different pipeline that preserves the generalization of SAM while improving its segmentation performance in medical imaging via a test-time framework. SAM-TTA tackles two key challenges: (1) input-level discrepancies caused by differences in image acquisition between natural and medical images and (2) semantic-level discrepancies due to fundamental differences in object definition between natural and medical domains (e.g., clear boundaries vs. ambiguous structures). Specifically, our SAM-TTA framework comprises (1) Self-adaptive Bezier Curve-based Transformation (SBCT), which adaptively converts single-channel medical images into three-channel SAM-compatible inputs while maintaining structural integrity, to mitigate the input gap between medical and natural images, and (2) Dual-scale Uncertainty-driven Mean Teacher adaptation (DUMT), which employs consistency learning to align SAM's internal representations to medical semantics, enabling efficient adaptation without auxiliary supervision or expensive retraining. Extensive experiments on five public datasets demonstrate that our SAM-TTA outperforms existing TTA approaches and even surpasses fully fine-tuned models such as MedSAM in certain scenarios, establishing a new paradigm for universal medical image segmentation. Code can be found at https://github.com/JianghaoWu/SAM-TTA.
Beyond Distillation: Pushing the Limits of Medical LLM Reasoning with Minimalist Rule-Based RL
May 23, 2025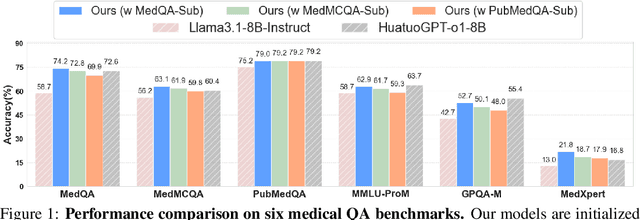
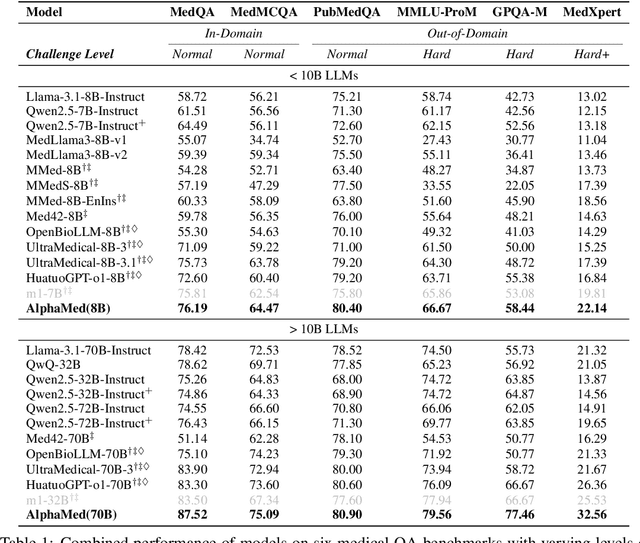
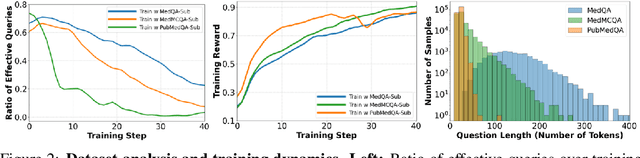
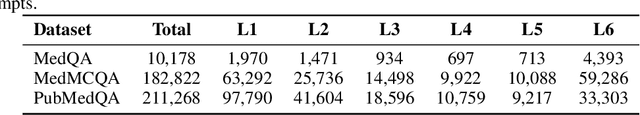
Abstract:Improving performance on complex tasks and enabling interpretable decision making in large language models (LLMs), especially for clinical applications, requires effective reasoning. Yet this remains challenging without supervised fine-tuning (SFT) on costly chain-of-thought (CoT) data distilled from closed-source models (e.g., GPT-4o). In this work, we present AlphaMed, the first medical LLM to show that reasoning capability can emerge purely through reinforcement learning (RL), using minimalist rule-based rewards on public multiple-choice QA datasets, without relying on SFT or distilled CoT data. AlphaMed achieves state-of-the-art results on six medical QA benchmarks, outperforming models trained with conventional SFT+RL pipelines. On challenging benchmarks (e.g., MedXpert), AlphaMed even surpasses larger or closed-source models such as DeepSeek-V3-671B and Claude-3.5-Sonnet. To understand the factors behind this success, we conduct a comprehensive data-centric analysis guided by three questions: (i) Can minimalist rule-based RL incentivize reasoning without distilled CoT supervision? (ii) How do dataset quantity and diversity impact reasoning? (iii) How does question difficulty shape the emergence and generalization of reasoning? Our findings show that dataset informativeness is a key driver of reasoning performance, and that minimalist RL on informative, multiple-choice QA data is effective at inducing reasoning without CoT supervision. We also observe divergent trends across benchmarks, underscoring limitations in current evaluation and the need for more challenging, reasoning-oriented medical QA benchmarks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge