"Image": models, code, and papers
IRGen: Generative Modeling for Image Retrieval
Mar 17, 2023
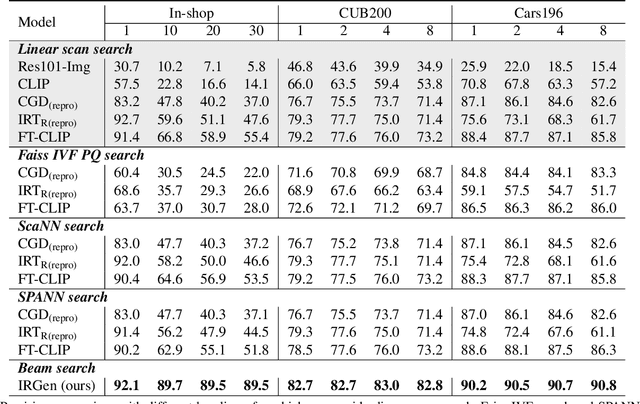
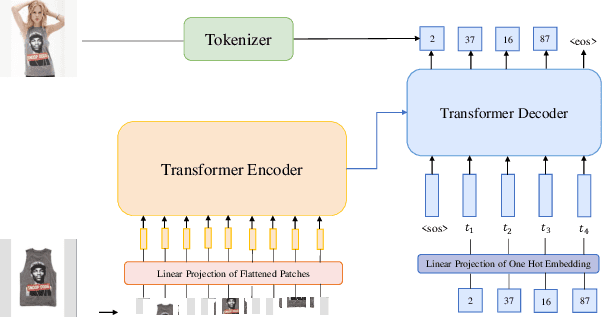
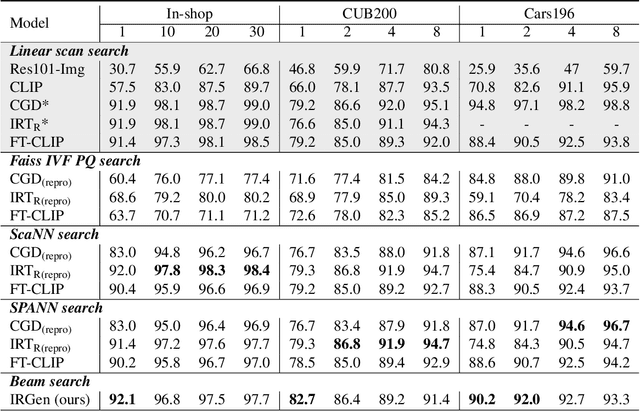
While generative modeling has been ubiquitous in natural language processing and computer vision, its application to image retrieval remains unexplored. In this paper, we recast image retrieval as a form of generative modeling by employing a sequence-to-sequence model, contributing to the current unified theme. Our framework, IRGen, is a unified model that enables end-to-end differentiable search, thus achieving superior performance thanks to direct optimization. While developing IRGen we tackle the key technical challenge of converting an image into quite a short sequence of semantic units in order to enable efficient and effective retrieval. Empirical experiments demonstrate that our model yields significant improvement over three commonly used benchmarks, for example, 22.9\% higher than the best baseline method in precision@10 on In-shop dataset with comparable recall@10 score.
Differentiable Blocks World: Qualitative 3D Decomposition by Rendering Primitives
Jul 11, 2023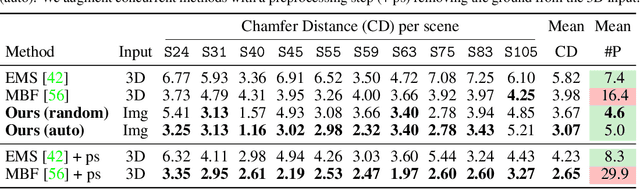
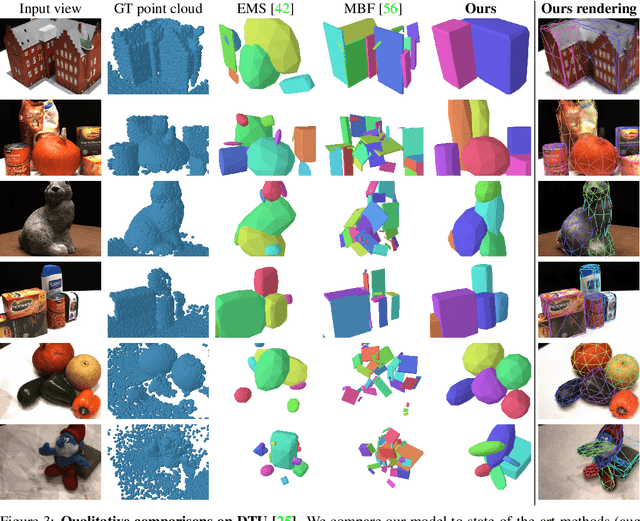
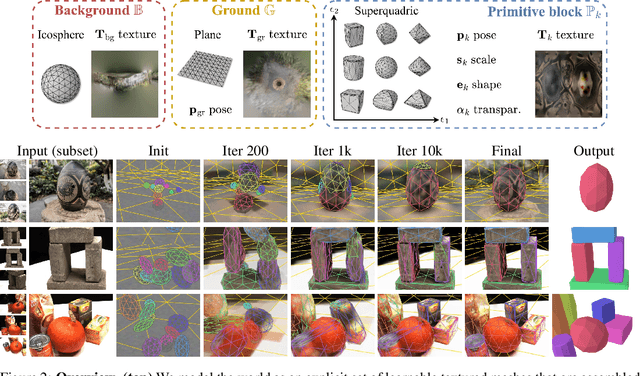
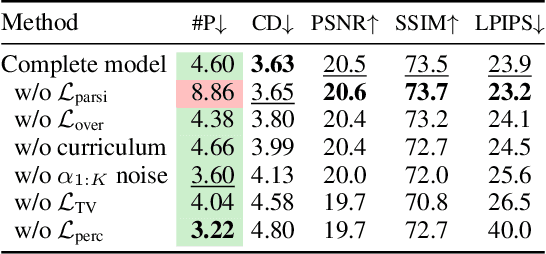

Given a set of calibrated images of a scene, we present an approach that produces a simple, compact, and actionable 3D world representation by means of 3D primitives. While many approaches focus on recovering high-fidelity 3D scenes, we focus on parsing a scene into mid-level 3D representations made of a small set of textured primitives. Such representations are interpretable, easy to manipulate and suited for physics-based simulations. Moreover, unlike existing primitive decomposition methods that rely on 3D input data, our approach operates directly on images through differentiable rendering. Specifically, we model primitives as textured superquadric meshes and optimize their parameters from scratch with an image rendering loss. We highlight the importance of modeling transparency for each primitive, which is critical for optimization and also enables handling varying numbers of primitives. We show that the resulting textured primitives faithfully reconstruct the input images and accurately model the visible 3D points, while providing amodal shape completions of unseen object regions. We compare our approach to the state of the art on diverse scenes from DTU, and demonstrate its robustness on real-life captures from BlendedMVS and Nerfstudio. We also showcase how our results can be used to effortlessly edit a scene or perform physical simulations. Code and video results are available at https://www.tmonnier.com/DBW .
Efficient 3D Articulated Human Generation with Layered Surface Volumes
Jul 11, 2023
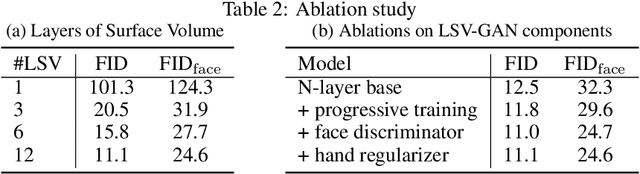
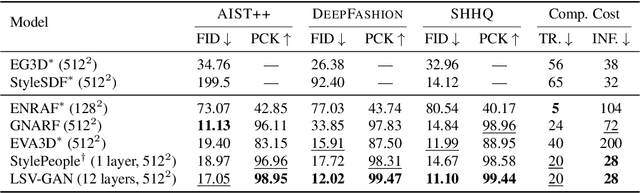
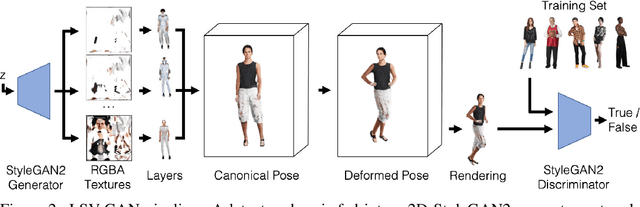

Access to high-quality and diverse 3D articulated digital human assets is crucial in various applications, ranging from virtual reality to social platforms. Generative approaches, such as 3D generative adversarial networks (GANs), are rapidly replacing laborious manual content creation tools. However, existing 3D GAN frameworks typically rely on scene representations that leverage either template meshes, which are fast but offer limited quality, or volumes, which offer high capacity but are slow to render, thereby limiting the 3D fidelity in GAN settings. In this work, we introduce layered surface volumes (LSVs) as a new 3D object representation for articulated digital humans. LSVs represent a human body using multiple textured mesh layers around a conventional template. These layers are rendered using alpha compositing with fast differentiable rasterization, and they can be interpreted as a volumetric representation that allocates its capacity to a manifold of finite thickness around the template. Unlike conventional single-layer templates that struggle with representing fine off-surface details like hair or accessories, our surface volumes naturally capture such details. LSVs can be articulated, and they exhibit exceptional efficiency in GAN settings, where a 2D generator learns to synthesize the RGBA textures for the individual layers. Trained on unstructured, single-view 2D image datasets, our LSV-GAN generates high-quality and view-consistent 3D articulated digital humans without the need for view-inconsistent 2D upsampling networks.
Continual Diffusion: Continual Customization of Text-to-Image Diffusion with C-LoRA
Apr 12, 2023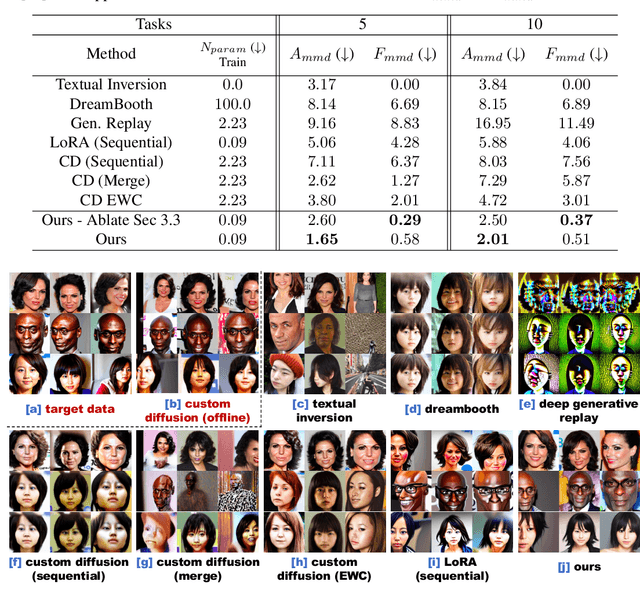
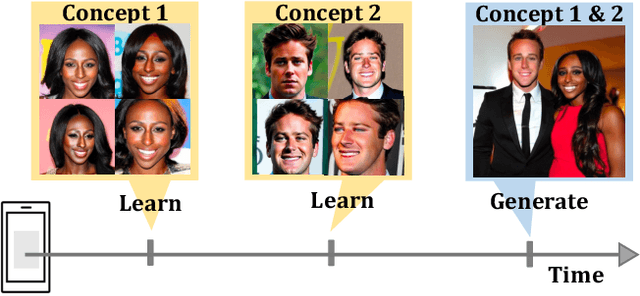
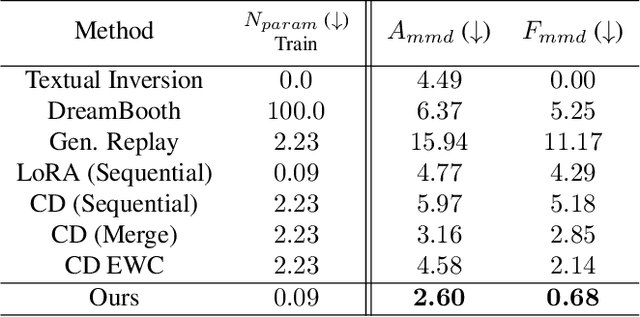
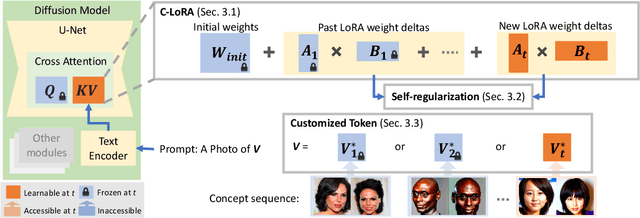
Recent works demonstrate a remarkable ability to customize text-to-image diffusion models while only providing a few example images. What happens if you try to customize such models using multiple, fine-grained concepts in a sequential (i.e., continual) manner? In our work, we show that recent state-of-the-art customization of text-to-image models suffer from catastrophic forgetting when new concepts arrive sequentially. Specifically, when adding a new concept, the ability to generate high quality images of past, similar concepts degrade. To circumvent this forgetting, we propose a new method, C-LoRA, composed of a continually self-regularized low-rank adaptation in cross attention layers of the popular Stable Diffusion model. Furthermore, we use customization prompts which do not include the word of the customized object (i.e., "person" for a human face dataset) and are initialized as completely random embeddings. Importantly, our method induces only marginal additional parameter costs and requires no storage of user data for replay. We show that C-LoRA not only outperforms several baselines for our proposed setting of text-to-image continual customization, which we refer to as Continual Diffusion, but that we achieve a new state-of-the-art in the well-established rehearsal-free continual learning setting for image classification. The high achieving performance of C-LoRA in two separate domains positions it as a compelling solution for a wide range of applications, and we believe it has significant potential for practical impact.
Deep Learning of Crystalline Defects from TEM images: A Solution for the Problem of "Never Enough Training Data"
Jul 12, 2023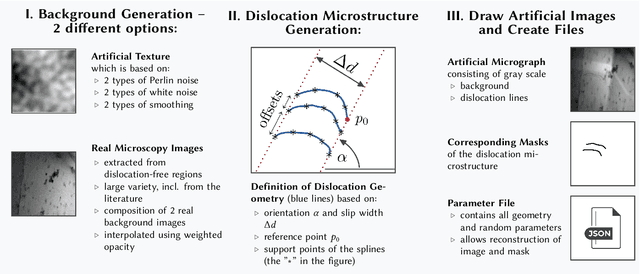
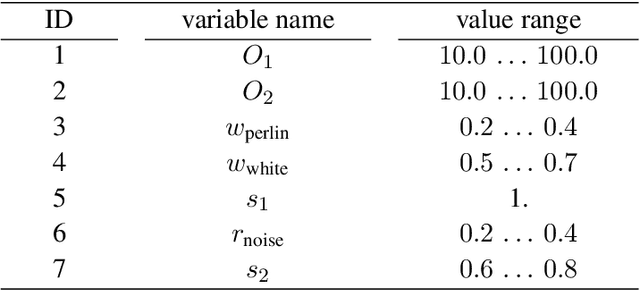
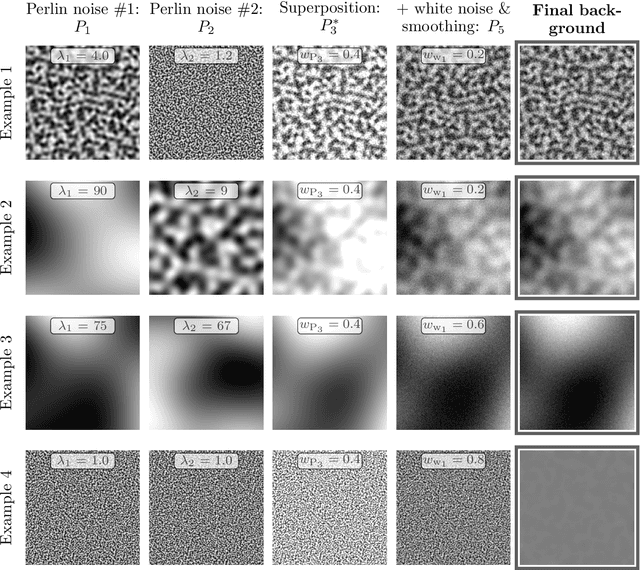
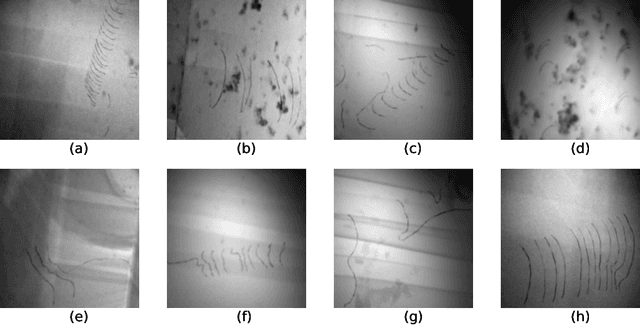



Crystalline defects, such as line-like dislocations, play an important role for the performance and reliability of many metallic devices. Their interaction and evolution still poses a multitude of open questions to materials science and materials physics. In-situ TEM experiments can provide important insights into how dislocations behave and move. During such experiments, the dislocation microstructure is captured in form of videos. The analysis of individual video frames can provide useful insights but is limited by the capabilities of automated identification, digitization, and quantitative extraction of the dislocations as curved objects. The vast amount of data also makes manual annotation very time consuming, thereby limiting the use of Deep Learning-based, automated image analysis and segmentation of the dislocation microstructure. In this work, a parametric model for generating synthetic training data for segmentation of dislocations is developed. Even though domain scientists might dismiss synthetic training images sometimes as too artificial, our findings show that they can result in superior performance, particularly regarding the generalizing of the Deep Learning models with respect to different microstructures and imaging conditions. Additionally, we propose an enhanced deep learning method optimized for segmenting overlapping or intersecting dislocation lines. Upon testing this framework on four distinct real datasets, we find that our synthetic training data are able to yield high-quality results also on real images-even more so if fine-tune on a few real images was done.
DCN-T: Dual Context Network with Transformer for Hyperspectral Image Classification
Apr 19, 2023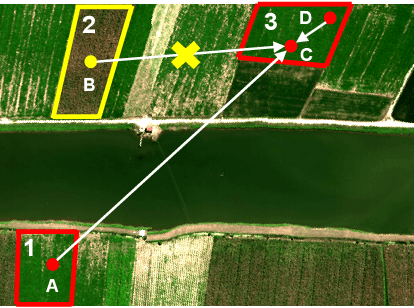
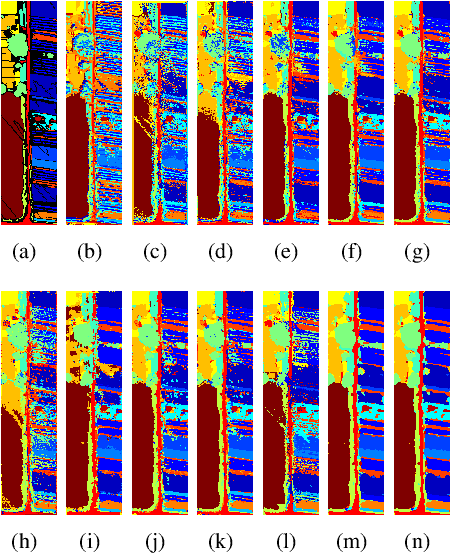
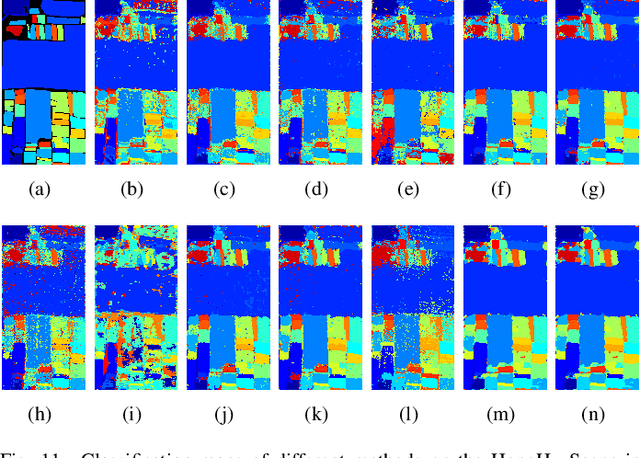
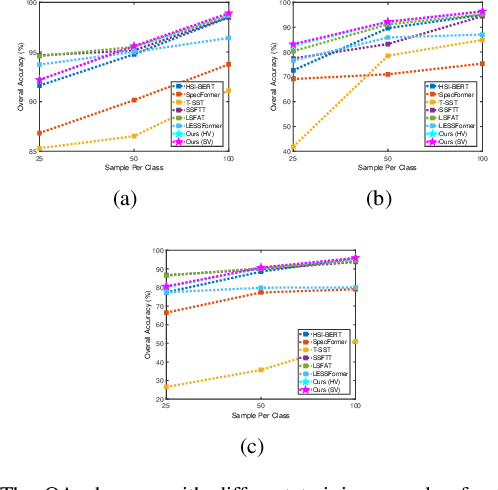
Hyperspectral image (HSI) classification is challenging due to spatial variability caused by complex imaging conditions. Prior methods suffer from limited representation ability, as they train specially designed networks from scratch on limited annotated data. We propose a tri-spectral image generation pipeline that transforms HSI into high-quality tri-spectral images, enabling the use of off-the-shelf ImageNet pretrained backbone networks for feature extraction. Motivated by the observation that there are many homogeneous areas with distinguished semantic and geometric properties in HSIs, which can be used to extract useful contexts, we propose an end-to-end segmentation network named DCN-T. It adopts transformers to effectively encode regional adaptation and global aggregation spatial contexts within and between the homogeneous areas discovered by similarity-based clustering. To fully exploit the rich spectrums of the HSI, we adopt an ensemble approach where all segmentation results of the tri-spectral images are integrated into the final prediction through a voting scheme. Extensive experiments on three public benchmarks show that our proposed method outperforms state-of-the-art methods for HSI classification.
Multimodal Image-Text Matching Improves Retrieval-based Chest X-Ray Report Generation
Mar 29, 2023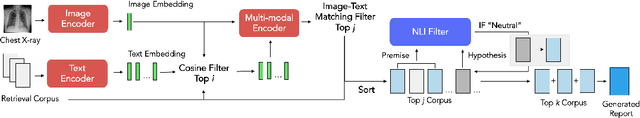
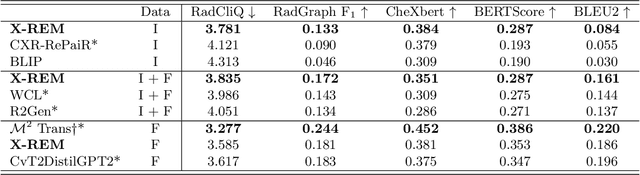
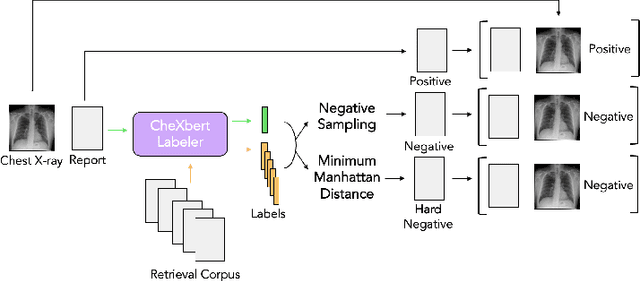
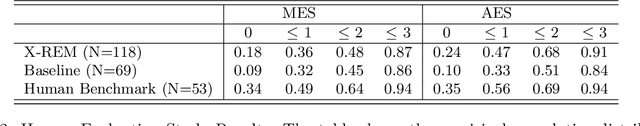
Automated generation of clinically accurate radiology reports can improve patient care. Previous report generation methods that rely on image captioning models often generate incoherent and incorrect text due to their lack of relevant domain knowledge, while retrieval-based attempts frequently retrieve reports that are irrelevant to the input image. In this work, we propose Contrastive X-Ray REport Match (X-REM), a novel retrieval-based radiology report generation module that uses an image-text matching score to measure the similarity of a chest X-ray image and radiology report for report retrieval. We observe that computing the image-text matching score with a language-image model can effectively capture the fine-grained interaction between image and text that is often lost when using cosine similarity. X-REM outperforms multiple prior radiology report generation modules in terms of both natural language and clinical metrics. Human evaluation of the generated reports suggests that X-REM increased the number of zero-error reports and decreased the average error severity compared to the baseline retrieval approach. Our code is available at: https://github.com/rajpurkarlab/X-REM
MedLSAM: Localize and Segment Anything Model for 3D Medical Images
Jun 30, 2023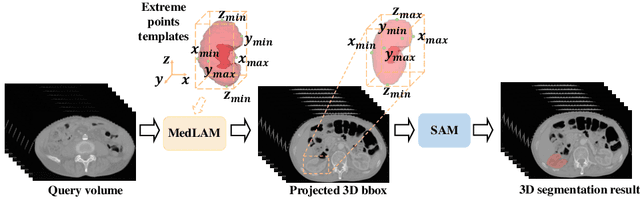
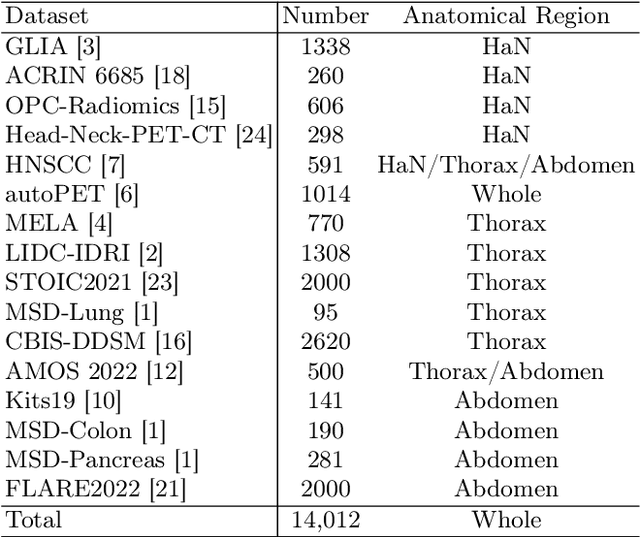
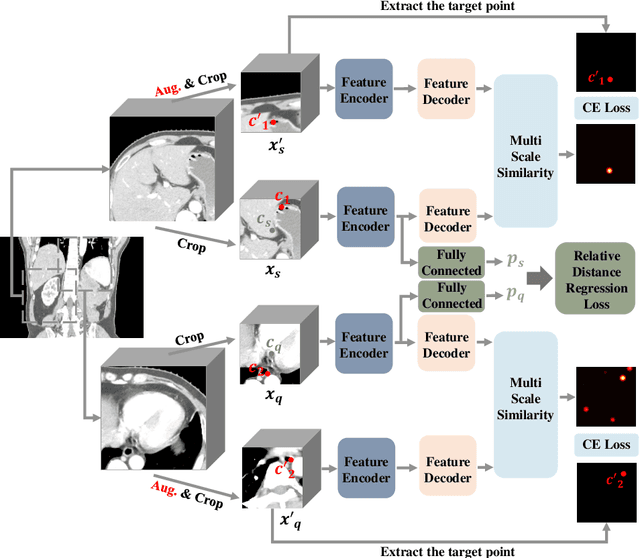
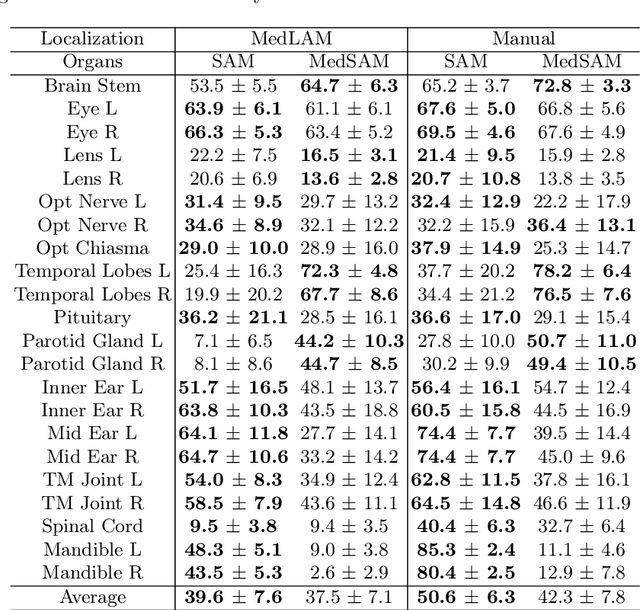
The Segment Anything Model (SAM) has recently emerged as a groundbreaking model in the field of image segmentation. Nevertheless, both the original SAM and its medical adaptations necessitate slice-by-slice annotations, which directly increase the annotation workload with the size of the dataset. We propose MedLSAM to address this issue, ensuring a constant annotation workload irrespective of dataset size and thereby simplifying the annotation process. Our model introduces a few-shot localization framework capable of localizing any target anatomical part within the body. To achieve this, we develop a Localize Anything Model for 3D Medical Images (MedLAM), utilizing two self-supervision tasks: relative distance regression (RDR) and multi-scale similarity (MSS) across a comprehensive dataset of 14,012 CT scans. We then establish a methodology for accurate segmentation by integrating MedLAM with SAM. By annotating only six extreme points across three directions on a few templates, our model can autonomously identify the target anatomical region on all data scheduled for annotation. This allows our framework to generate a 2D bounding box for every slice of the image, which are then leveraged by SAM to carry out segmentations. We conducted experiments on two 3D datasets covering 38 organs and found that MedLSAM matches the performance of SAM and its medical adaptations while requiring only minimal extreme point annotations for the entire dataset. Furthermore, MedLAM has the potential to be seamlessly integrated with future 3D SAM models, paving the way for enhanced performance. Our code is public at https://github.com/openmedlab/MedLSAM.
Designing a Better Asymmetric VQGAN for StableDiffusion
Jun 07, 2023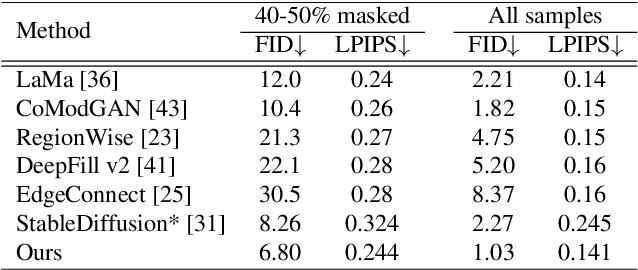
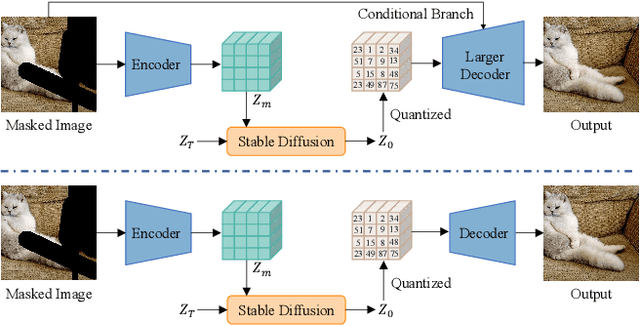
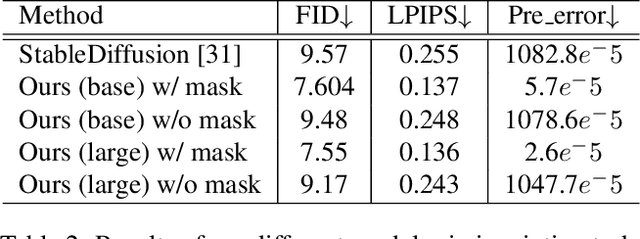
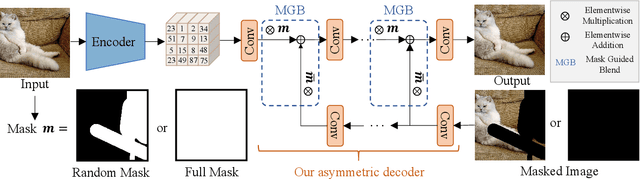
StableDiffusion is a revolutionary text-to-image generator that is causing a stir in the world of image generation and editing. Unlike traditional methods that learn a diffusion model in pixel space, StableDiffusion learns a diffusion model in the latent space via a VQGAN, ensuring both efficiency and quality. It not only supports image generation tasks, but also enables image editing for real images, such as image inpainting and local editing. However, we have observed that the vanilla VQGAN used in StableDiffusion leads to significant information loss, causing distortion artifacts even in non-edited image regions. To this end, we propose a new asymmetric VQGAN with two simple designs. Firstly, in addition to the input from the encoder, the decoder contains a conditional branch that incorporates information from task-specific priors, such as the unmasked image region in inpainting. Secondly, the decoder is much heavier than the encoder, allowing for more detailed recovery while only slightly increasing the total inference cost. The training cost of our asymmetric VQGAN is cheap, and we only need to retrain a new asymmetric decoder while keeping the vanilla VQGAN encoder and StableDiffusion unchanged. Our asymmetric VQGAN can be widely used in StableDiffusion-based inpainting and local editing methods. Extensive experiments demonstrate that it can significantly improve the inpainting and editing performance, while maintaining the original text-to-image capability. The code is available at \url{https://github.com/buxiangzhiren/Asymmetric_VQGAN}.
Self-training with dual uncertainty for semi-supervised medical image segmentation
Apr 10, 2023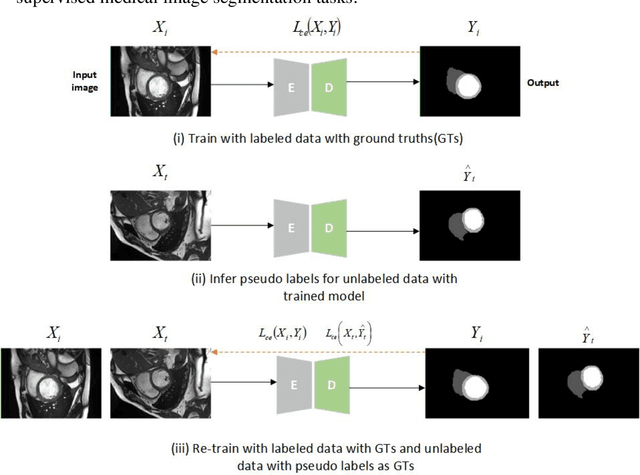
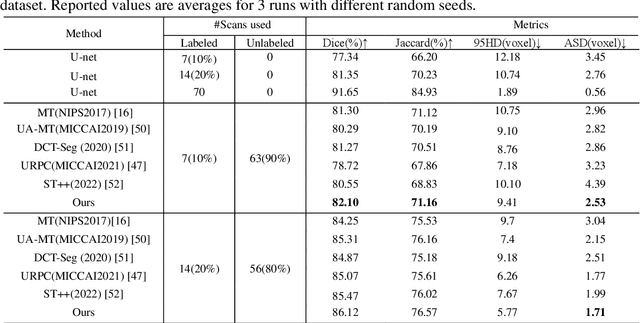
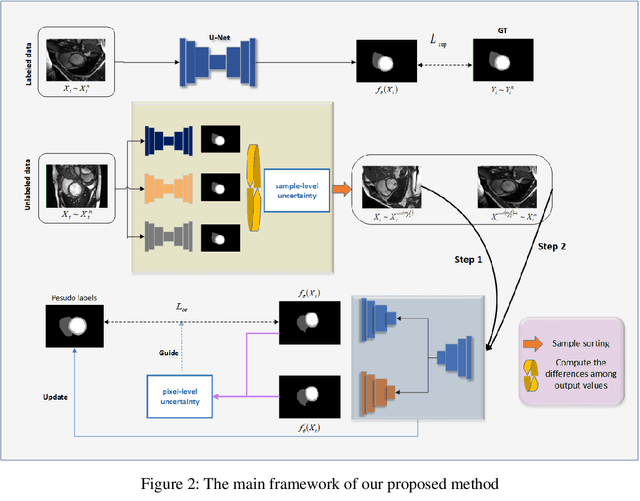
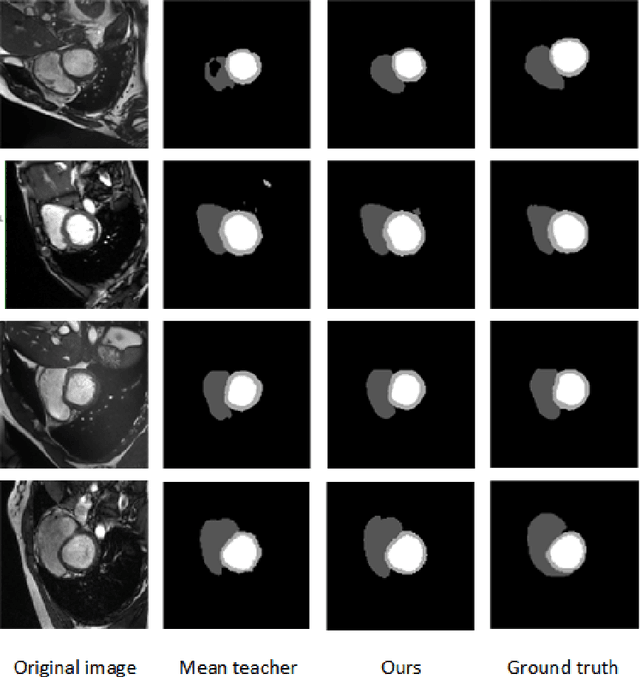
In the field of semi-supervised medical image segmentation, the shortage of labeled data is the fundamental problem. How to effectively learn image features from unlabeled images to improve segmentation accuracy is the main research direction in this field. Traditional self-training methods can partially solve the problem of insufficient labeled data by generating pseudo labels for iterative training. However, noise generated due to the model's uncertainty during training directly affects the segmentation results. Therefore, we added sample-level and pixel-level uncertainty to stabilize the training process based on the self-training framework. Specifically, we saved several moments of the model during pre-training, and used the difference between their predictions on unlabeled samples as the sample-level uncertainty estimate for that sample. Then, we gradually add unlabeled samples from easy to hard during training. At the same time, we added a decoder with different upsampling methods to the segmentation network and used the difference between the outputs of the two decoders as pixel-level uncertainty. In short, we selectively retrained unlabeled samples and assigned pixel-level uncertainty to pseudo labels to optimize the self-training process. We compared the segmentation results of our model with five semi-supervised approaches on the public 2017 ACDC dataset and 2018 Prostate dataset. Our proposed method achieves better segmentation performance on both datasets under the same settings, demonstrating its effectiveness, robustness, and potential transferability to other medical image segmentation tasks. Keywords: Medical image segmentation, semi-supervised learning, self-training, uncertainty estimation
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge