Pranav Rajpurkar
Harvard University
Imitation Learning for Robot Assistance in Open Surgery: A Multi-Policy Evaluation on Suture Following
May 27, 2026Abstract:This study presents the first evaluation of general-purpose imitation learning for surgeon-robot collaborative assistance in open surgery, targeting suture following: the grab-pull-release motion an assistant performs at every stitch. We collect 160 teleoperated demonstrations (32,374 frames) on an open-source robot arm, benchmark four architecturally diverse imitation learning policies (ACT, Diffusion Policy, SmolVLA, $π_0$) across 28 trained models evaluated in 32 configurations along three clinically motivated dimensions: dataset size, camera viewpoint, and background variation. Our results demonstrate that under ideal conditions, the four policies achieve $50$-$75\%$ task success, with depth error as the dominant failure mode across all architectures. Among all policies, $π_0$ achieves the strongest results with a pretrained vision-language backbone, demonstrating superior data efficiency, greater robustness to background variation, and smoother trajectories compatible with surgical workflow. When deployed in a surgeon-robot suturing trial, $π_0$ yields a $92\%$ stitch completion rate. These findings establish collaborative robotic assistance in open surgery as a feasible target for imitation learning and highlight depth perception and end-effector design as key priorities for clinical translation.
ReXSonoVQA: A Video QA Benchmark for Procedure-Centric Ultrasound Understanding
Apr 14, 2026Abstract:Ultrasound acquisition requires skilled probe manipulation and real-time adjustments. Vision-language models (VLMs) could enable autonomous ultrasound systems, but existing benchmarks evaluate only static images, not dynamic procedural understanding. We introduce ReXSonoVQA, a video QA benchmark with 514 video clips and 514 questions (249 MCQ, 265 free-response) targeting three competencies: Action-Goal Reasoning, Artifact Resolution & Optimization, and Procedure Context & Planning. Zero-shot evaluation of Gemini 3 Pro, Qwen3.5-397B, LLaVA-Video-72B, and Seed 2.0 Pro shows VLMs can extract some procedural information, but troubleshooting questions remain challenging with minimal gains over text-only baselines, exposing limitations in causal reasoning. ReXSonoVQA enables developing perception systems for ultrasound training, guidance, and robotic automation.
ReXInTheWild: A Unified Benchmark for Medical Photograph Understanding
Mar 19, 2026Abstract:Everyday photographs taken with ordinary cameras are already widely used in telemedicine and other online health conversations, yet no comprehensive benchmark evaluates whether vision-language models can interpret their medical content. Analyzing these images requires both fine-grained natural image understanding and domain-specific medical reasoning, a combination that challenges both general-purpose and specialized models. We introduce ReXInTheWild, a benchmark of 955 clinician-verified multiple-choice questions spanning seven clinical topics across 484 photographs sourced from the biomedical literature. When evaluated on ReXInTheWild, leading multimodal large language models show substantial performance variation: Gemini-3 achieves 78% accuracy, followed by Claude Opus 4.5 (72%) and GPT-5 (68%), while the medical specialist model MedGemma reaches only 37%. A systematic error analysis also reveals four categories of common errors, ranging from low-level geometric errors to high-level reasoning failures and requiring different mitigation strategies. ReXInTheWild provides a challenging, clinically grounded benchmark at the intersection of natural image understanding and medical reasoning. The dataset is available on HuggingFace.
Evaluating Contextual Intelligence in Recyclability: A Comprehensive Study of Image-Based Reasoning Systems
Dec 31, 2025Abstract:While the importance of efficient recycling is widely acknowledged, accurately determining the recyclability of items and their proper disposal remains a complex task for the general public. In this study, we explore the application of cutting-edge vision-language models (GPT-4o, GPT-4o-mini, and Claude 3.5) for predicting the recyclability of commonly disposed items. Utilizing a curated dataset of images, we evaluated the models' ability to match objects to appropriate recycling bins, including assessing whether the items could physically fit into the available bins. Additionally, we investigated the models' performance across several challenging scenarios: (i) adjusting predictions based on location-specific recycling guidelines; (ii) accounting for contamination or structural damage; and (iii) handling objects composed of multiple materials. Our findings highlight the significant advancements in contextual understanding offered by these models compared to previous iterations, while also identifying areas where they still fall short. The continued refinement of context-aware models is crucial for enhancing public recycling practices and advancing environmental sustainability.
ReX-MLE: The Autonomous Agent Benchmark for Medical Imaging Challenges
Dec 19, 2025



Abstract:Autonomous coding agents built on large language models (LLMs) can now solve many general software and machine learning tasks, but they remain ineffective on complex, domain-specific scientific problems. Medical imaging is a particularly demanding domain, requiring long training cycles, high-dimensional data handling, and specialized preprocessing and validation pipelines, capabilities not fully measured in existing agent benchmarks. To address this gap, we introduce ReX-MLE, a benchmark of 20 challenges derived from high-impact medical imaging competitions spanning diverse modalities and task types. Unlike prior ML-agent benchmarks, ReX-MLE evaluates full end-to-end workflows, requiring agents to independently manage data preprocessing, model training, and submission under realistic compute and time constraints. Evaluating state-of-the-art agents (AIDE, ML-Master, R&D-Agent) with different LLM backends (GPT-5, Gemini, Claude), we observe a severe performance gap: most submissions rank in the 0th percentile compared to human experts. Failures stem from domain-knowledge and engineering limitations. ReX-MLE exposes these bottlenecks and provides a foundation for developing domain-aware autonomous AI systems.
RadGame: An AI-Powered Platform for Radiology Education
Sep 16, 2025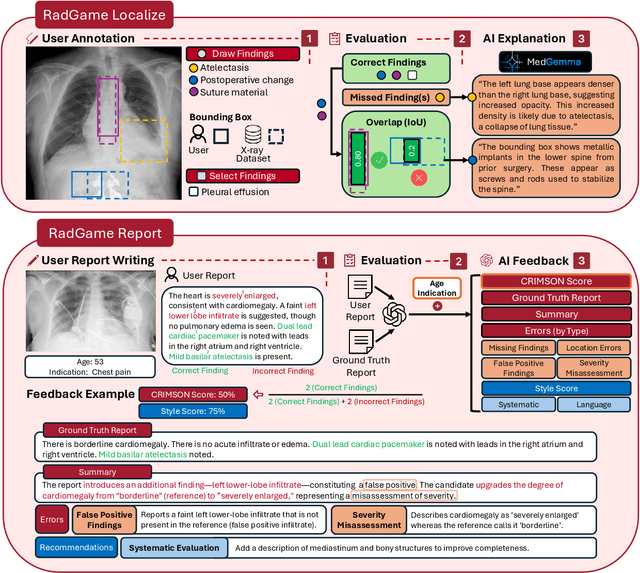
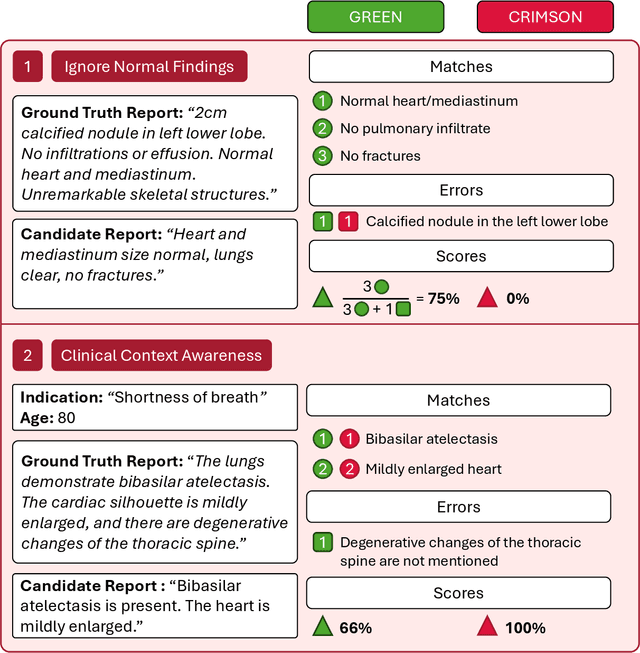
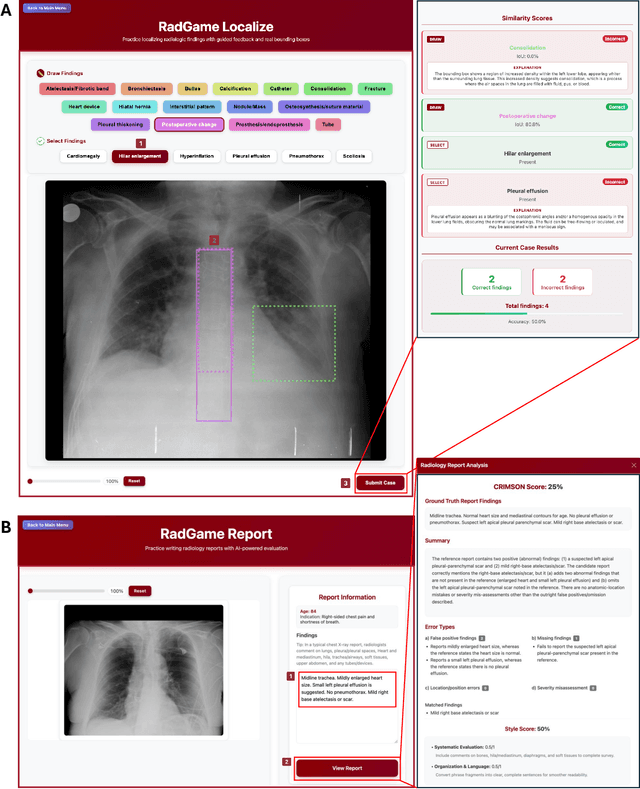
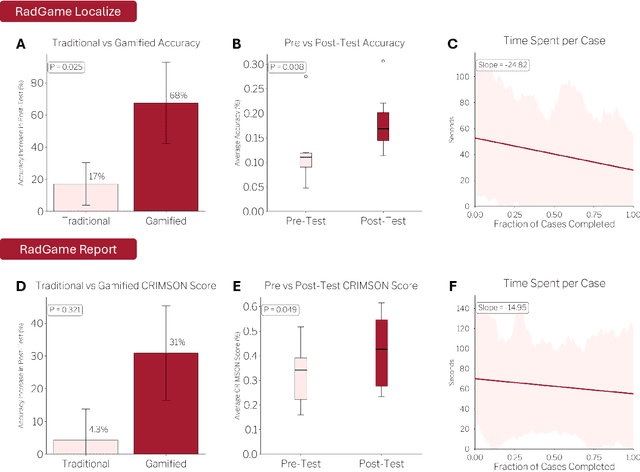

Abstract:We introduce RadGame, an AI-powered gamified platform for radiology education that targets two core skills: localizing findings and generating reports. Traditional radiology training is based on passive exposure to cases or active practice with real-time input from supervising radiologists, limiting opportunities for immediate and scalable feedback. RadGame addresses this gap by combining gamification with large-scale public datasets and automated, AI-driven feedback that provides clear, structured guidance to human learners. In RadGame Localize, players draw bounding boxes around abnormalities, which are automatically compared to radiologist-drawn annotations from public datasets, and visual explanations are generated by vision-language models for user missed findings. In RadGame Report, players compose findings given a chest X-ray, patient age and indication, and receive structured AI feedback based on radiology report generation metrics, highlighting errors and omissions compared to a radiologist's written ground truth report from public datasets, producing a final performance and style score. In a prospective evaluation, participants using RadGame achieved a 68% improvement in localization accuracy compared to 17% with traditional passive methods and a 31% improvement in report-writing accuracy compared to 4% with traditional methods after seeing the same cases. RadGame highlights the potential of AI-driven gamification to deliver scalable, feedback-rich radiology training and reimagines the application of medical AI resources in education.
ReXGroundingCT: A 3D Chest CT Dataset for Segmentation of Findings from Free-Text Reports
Jul 29, 2025Abstract:We present ReXGroundingCT, the first publicly available dataset to link free-text radiology findings with pixel-level segmentations in 3D chest CT scans that is manually annotated. While prior datasets have relied on structured labels or predefined categories, ReXGroundingCT captures the full expressiveness of clinical language represented in free text and grounds it to spatially localized 3D segmentation annotations in volumetric imaging. This addresses a critical gap in medical AI: the ability to connect complex, descriptive text, such as "3 mm nodule in the left lower lobe", to its precise anatomical location in three-dimensional space, a capability essential for grounded radiology report generation systems. The dataset comprises 3,142 non-contrast chest CT scans paired with standardized radiology reports from the CT-RATE dataset. Using a systematic three-stage pipeline, GPT-4 was used to extract positive lung and pleural findings, which were then manually segmented by expert annotators. A total of 8,028 findings across 16,301 entities were annotated, with quality control performed by board-certified radiologists. Approximately 79% of findings are focal abnormalities, while 21% are non-focal. The training set includes up to three representative segmentations per finding, while the validation and test sets contain exhaustive labels for each finding entity. ReXGroundingCT establishes a new benchmark for developing and evaluating sentence-level grounding and free-text medical segmentation models in chest CT. The dataset can be accessed at https://huggingface.co/datasets/rajpurkarlab/ReXGroundingCT.
ReXVQA: A Large-scale Visual Question Answering Benchmark for Generalist Chest X-ray Understanding
Jun 04, 2025



Abstract:We present ReXVQA, the largest and most comprehensive benchmark for visual question answering (VQA) in chest radiology, comprising approximately 696,000 questions paired with 160,000 chest X-rays studies across training, validation, and test sets. Unlike prior efforts that rely heavily on template based queries, ReXVQA introduces a diverse and clinically authentic task suite reflecting five core radiological reasoning skills: presence assessment, location analysis, negation detection, differential diagnosis, and geometric reasoning. We evaluate eight state-of-the-art multimodal large language models, including MedGemma-4B-it, Qwen2.5-VL, Janus-Pro-7B, and Eagle2-9B. The best-performing model (MedGemma) achieves 83.24% overall accuracy. To bridge the gap between AI performance and clinical expertise, we conducted a comprehensive human reader study involving 3 radiology residents on 200 randomly sampled cases. Our evaluation demonstrates that MedGemma achieved superior performance (83.84% accuracy) compared to human readers (best radiology resident: 77.27%), representing a significant milestone where AI performance exceeds expert human evaluation on chest X-ray interpretation. The reader study reveals distinct performance patterns between AI models and human experts, with strong inter-reader agreement among radiologists while showing more variable agreement patterns between human readers and AI models. ReXVQA establishes a new standard for evaluating generalist radiological AI systems, offering public leaderboards, fine-grained evaluation splits, structured explanations, and category-level breakdowns. This benchmark lays the foundation for next-generation AI systems capable of mimicking expert-level clinical reasoning beyond narrow pathology classification. Our dataset will be open-sourced at https://huggingface.co/datasets/rajpurkarlab/ReXVQA
ReXGradient-160K: A Large-Scale Publicly Available Dataset of Chest Radiographs with Free-text Reports
May 01, 2025

Abstract:We present ReXGradient-160K, representing the largest publicly available chest X-ray dataset to date in terms of the number of patients. This dataset contains 160,000 chest X-ray studies with paired radiological reports from 109,487 unique patients across 3 U.S. health systems (79 medical sites). This comprehensive dataset includes multiple images per study and detailed radiology reports, making it particularly valuable for the development and evaluation of AI systems for medical imaging and automated report generation models. The dataset is divided into training (140,000 studies), validation (10,000 studies), and public test (10,000 studies) sets, with an additional private test set (10,000 studies) reserved for model evaluation on the ReXrank benchmark. By providing this extensive dataset, we aim to accelerate research in medical imaging AI and advance the state-of-the-art in automated radiological analysis. Our dataset will be open-sourced at https://huggingface.co/datasets/rajpurkarlab/ReXGradient-160K.
FreeTumor: Large-Scale Generative Tumor Synthesis in Computed Tomography Images for Improving Tumor Recognition
Feb 23, 2025Abstract:Tumor is a leading cause of death worldwide, with an estimated 10 million deaths attributed to tumor-related diseases every year. AI-driven tumor recognition unlocks new possibilities for more precise and intelligent tumor screening and diagnosis. However, the progress is heavily hampered by the scarcity of annotated datasets, which demands extensive annotation efforts by radiologists. To tackle this challenge, we introduce FreeTumor, an innovative Generative AI (GAI) framework to enable large-scale tumor synthesis for mitigating data scarcity. Specifically, FreeTumor effectively leverages a combination of limited labeled data and large-scale unlabeled data for tumor synthesis training. Unleashing the power of large-scale data, FreeTumor is capable of synthesizing a large number of realistic tumors on images for augmenting training datasets. To this end, we create the largest training dataset for tumor synthesis and recognition by curating 161,310 publicly available Computed Tomography (CT) volumes from 33 sources, with only 2.3% containing annotated tumors. To validate the fidelity of synthetic tumors, we engaged 13 board-certified radiologists in a Visual Turing Test to discern between synthetic and real tumors. Rigorous clinician evaluation validates the high quality of our synthetic tumors, as they achieved only 51.1% sensitivity and 60.8% accuracy in distinguishing our synthetic tumors from real ones. Through high-quality tumor synthesis, FreeTumor scales up the recognition training datasets by over 40 times, showcasing a notable superiority over state-of-the-art AI methods including various synthesis methods and foundation models. These findings indicate promising prospects of FreeTumor in clinical applications, potentially advancing tumor treatments and improving the survival rates of patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge