"Image": models, code, and papers
Long-Tailed Continual Learning For Visual Food Recognition
Jul 01, 2023
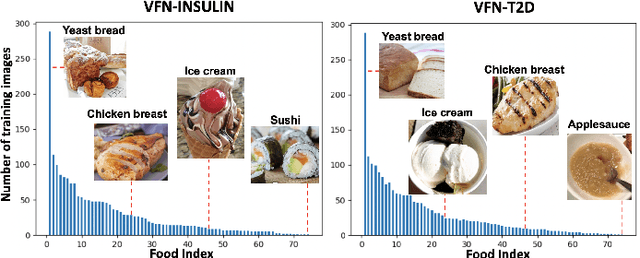
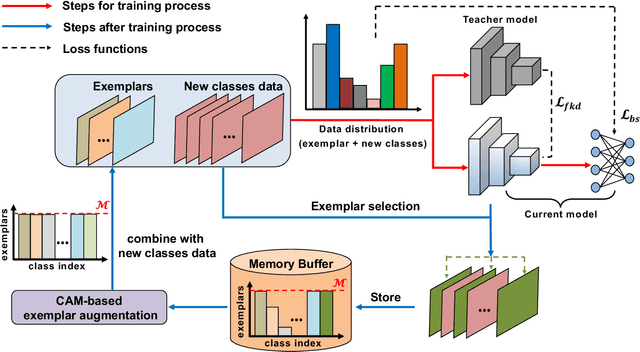
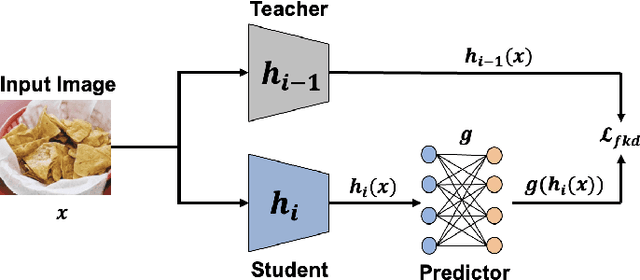
Deep learning based food recognition has achieved remarkable progress in predicting food types given an eating occasion image. However, there are two major obstacles that hinder deployment in real world scenario. First, as new foods appear sequentially overtime, a trained model needs to learn the new classes continuously without causing catastrophic forgetting for already learned knowledge of existing food types. Second, the distribution of food images in real life is usually long-tailed as a small number of popular food types are consumed more frequently than others, which can vary in different populations. This requires the food recognition method to learn from class-imbalanced data by improving the generalization ability on instance-rare food classes. In this work, we focus on long-tailed continual learning and aim to address both aforementioned challenges. As existing long-tailed food image datasets only consider healthy people population, we introduce two new benchmark food image datasets, VFN-INSULIN and VFN-T2D, which exhibits on the real world food consumption for insulin takers and individuals with type 2 diabetes without taking insulin, respectively. We propose a novel end-to-end framework for long-tailed continual learning, which effectively addresses the catastrophic forgetting by applying an additional predictor for knowledge distillation to avoid misalignment of representation during continual learning. We also introduce a novel data augmentation technique by integrating class-activation-map (CAM) and CutMix, which significantly improves the generalization ability for instance-rare food classes to address the class-imbalance issue. The proposed method show promising performance with large margin improvements compared with existing methods.
Spatio-Temporal Classification of Lung Ventilation Patterns using 3D EIT Images: A General Approach for Individualized Lung Function Evaluation
Jul 01, 2023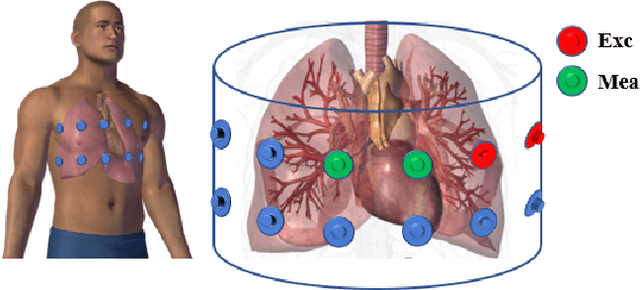
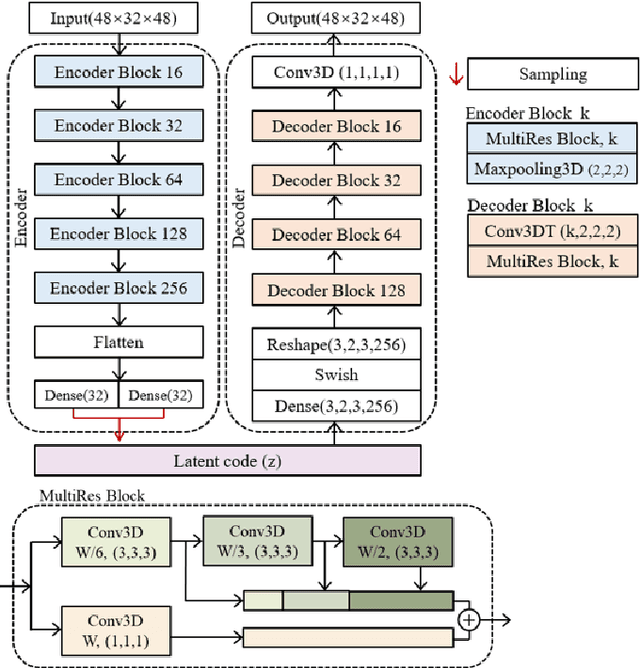
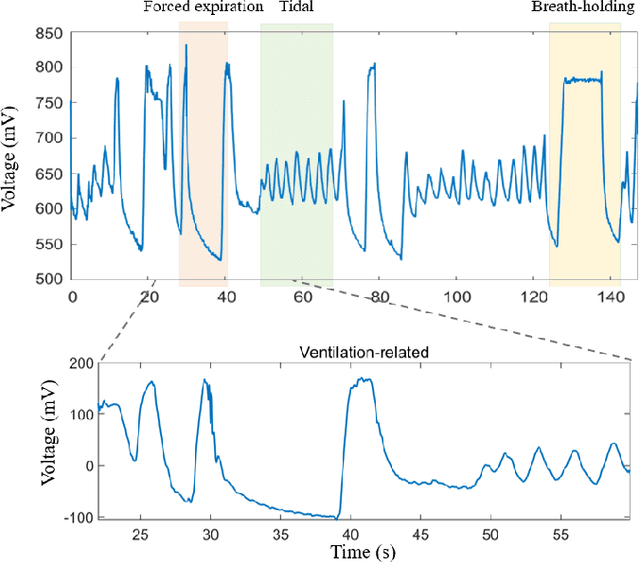
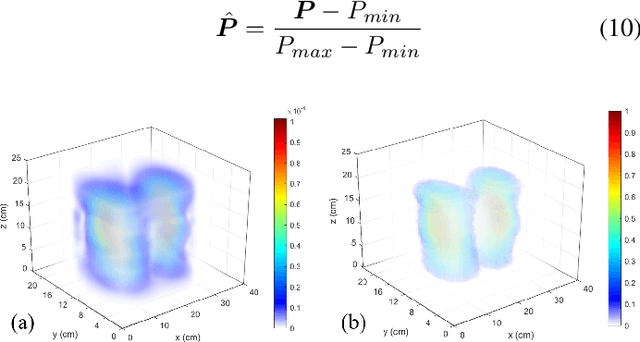
The Pulmonary Function Test (PFT) is an widely utilized and rigorous classification test for lung function evaluation, serving as a comprehensive tool for lung diagnosis. Meanwhile, Electrical Impedance Tomography (EIT) is a rapidly advancing clinical technique that visualizes conductivity distribution induced by ventilation. EIT provides additional spatial and temporal information on lung ventilation beyond traditional PFT. However, relying solely on conventional isolated interpretations of PFT results and EIT images overlooks the continuous dynamic aspects of lung ventilation. This study aims to classify lung ventilation patterns by extracting spatial and temporal features from the 3D EIT image series. The study uses a Variational Autoencoder network with a MultiRes block to compress the spatial distribution in a 3D image into a one-dimensional vector. These vectors are then concatenated to create a feature map for the exhibition of temporal features. A simple convolutional neural network is used for classification. Data collected from 137 subjects were finally used for training. The model is validated by ten-fold and leave-one-out cross-validation first. The accuracy and sensitivity of normal ventilation mode are 0.95 and 1.00, and the f1-score is 0.94. Furthermore, we check the reliability and feasibility of the proposed pipeline by testing it on newly recruited nine subjects. Our results show that the pipeline correctly predicts the ventilation mode of 8 out of 9 subjects. The study demonstrates the potential of using image series for lung ventilation mode classification, providing a feasible method for patient prescreening and presenting an alternative form of PFT.
Augment Features Beyond Color for Domain Generalized Segmentation
Jul 04, 2023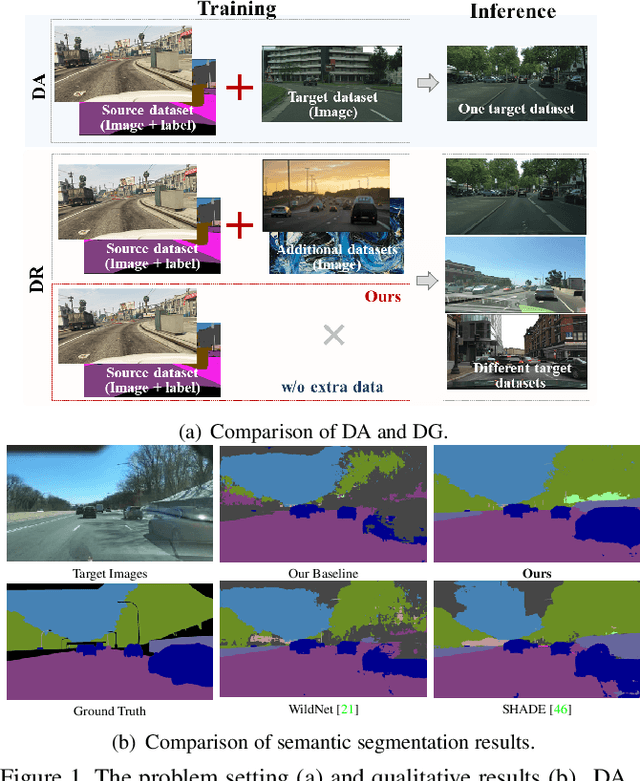
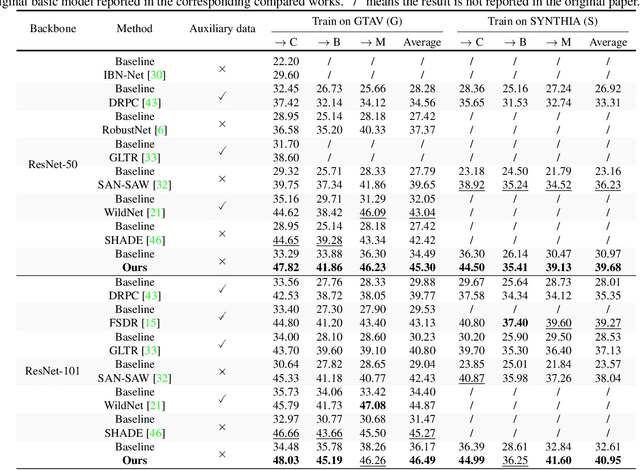
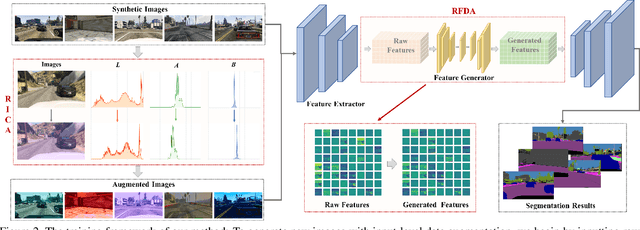
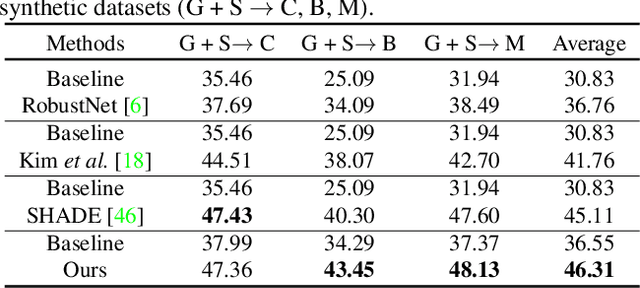
Domain generalized semantic segmentation (DGSS) is an essential but highly challenging task, in which the model is trained only on source data and any target data is not available. Previous DGSS methods can be partitioned into augmentation-based and normalization-based ones. The former either introduces extra biased data or only conducts channel-wise adjustments for data augmentation, and the latter may discard beneficial visual information, both of which lead to limited performance in DGSS. Contrarily, our method performs inter-channel transformation and meanwhile evades domain-specific biases, thus diversifying data and enhancing model generalization performance. Specifically, our method consists of two modules: random image color augmentation (RICA) and random feature distribution augmentation (RFDA). RICA converts images from RGB to the CIELAB color model and randomizes color maps in a perception-based way for image enhancement purposes. We further this augmentation by extending it beyond color to feature space using a CycleGAN-based generative network, which complements RICA and further boosts generalization capability. We conduct extensive experiments, and the generalization results from the synthetic GTAV and SYNTHIA to the real Cityscapes, BDDS, and Mapillary datasets show that our method achieves state-of-the-art performance in DGSS.
Realistic Data Enrichment for Robust Image Segmentation in Histopathology
Apr 19, 2023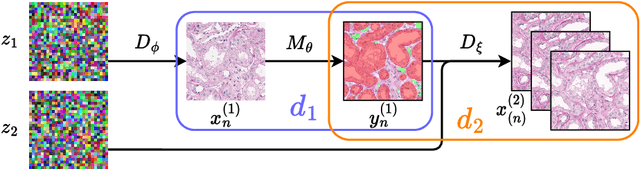
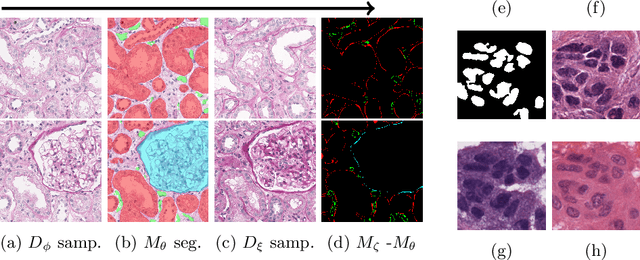
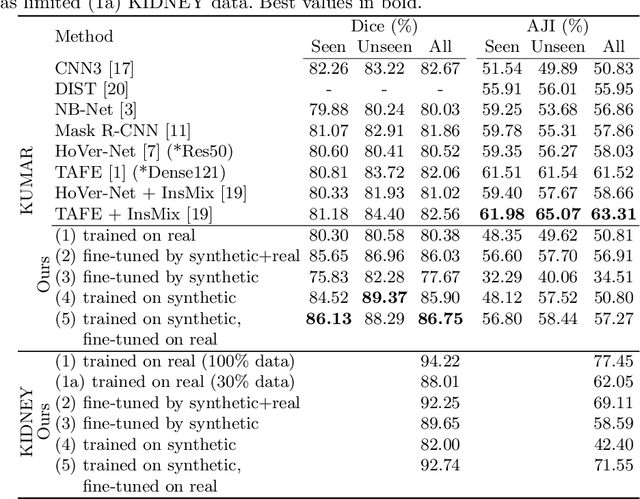

Poor performance of quantitative analysis in histopathological Whole Slide Images (WSI) has been a significant obstacle in clinical practice. Annotating large-scale WSIs manually is a demanding and time-consuming task, unlikely to yield the expected results when used for fully supervised learning systems. Rarely observed disease patterns and large differences in object scales are difficult to model through conventional patient intake. Prior methods either fall back to direct disease classification, which only requires learning a few factors per image, or report on average image segmentation performance, which is highly biased towards majority observations. Geometric image augmentation is commonly used to improve robustness for average case predictions and to enrich limited datasets. So far no method provided sampling of a realistic posterior distribution to improve stability, e.g. for the segmentation of imbalanced objects within images. Therefore, we propose a new approach, based on diffusion models, which can enrich an imbalanced dataset with plausible examples from underrepresented groups by conditioning on segmentation maps. Our method can simply expand limited clinical datasets making them suitable to train machine learning pipelines, and provides an interpretable and human-controllable way of generating histopathology images that are indistinguishable from real ones to human experts. We validate our findings on two datasets, one from the public domain and one from a Kidney Transplant study.
VVC+M: Plug and Play Scalable Image Coding for Humans and Machines
May 17, 2023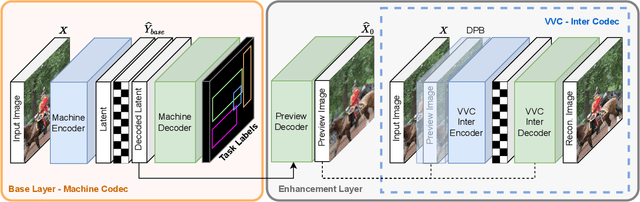
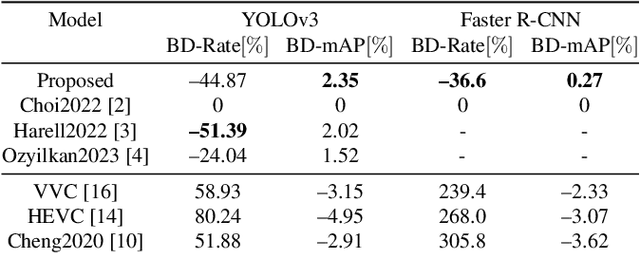
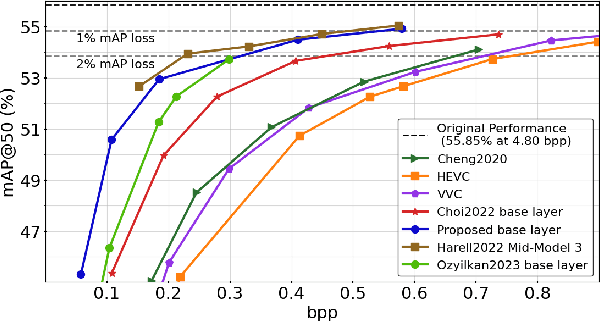
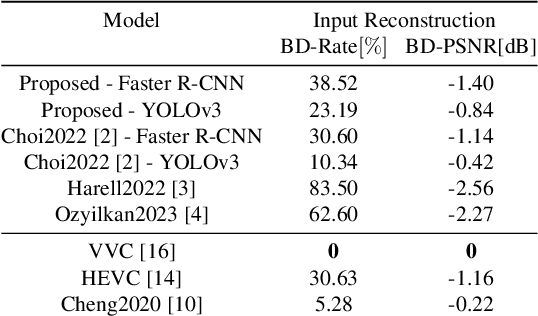
Compression for machines is an emerging field, where inputs are encoded while optimizing the performance of downstream automated analysis. In scalable coding for humans and machines, the compressed representation used for machines is further utilized to enable input reconstruction. Often performed by jointly optimizing the compression scheme for both machine task and human perception, this results in sub-optimal rate-distortion (RD) performance for the machine side. We focus on the case of images, proposing to utilize the pre-existing residual coding capabilities of video codecs such as VVC to create a scalable codec from any image compression for machines (ICM) scheme. Using our approach we improve an existing scalable codec to achieve superior RD performance on the machine task, while remaining competitive for human perception. Moreover, our approach can be trained post-hoc for any given ICM scheme, and without creating a coupling between the quality of the machine analysis and human vision.
Goal Representations for Instruction Following: A Semi-Supervised Language Interface to Control
Jun 30, 2023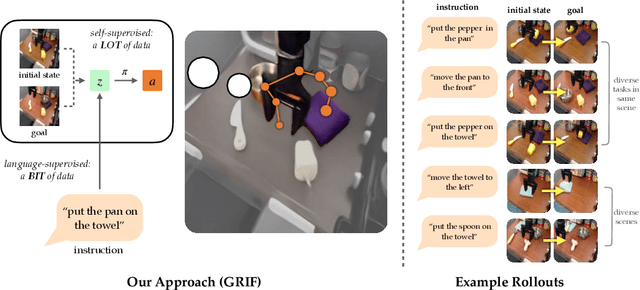

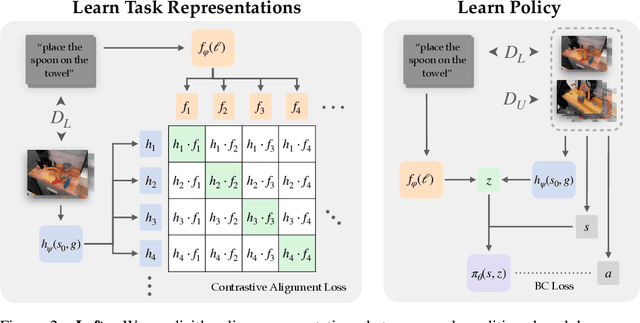
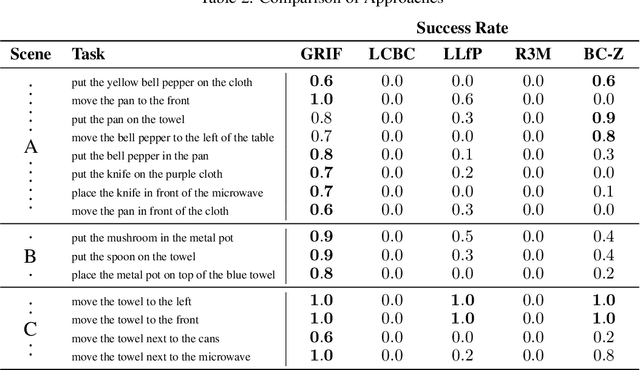
Our goal is for robots to follow natural language instructions like "put the towel next to the microwave." But getting large amounts of labeled data, i.e. data that contains demonstrations of tasks labeled with the language instruction, is prohibitive. In contrast, obtaining policies that respond to image goals is much easier, because any autonomous trial or demonstration can be labeled in hindsight with its final state as the goal. In this work, we contribute a method that taps into joint image- and goal- conditioned policies with language using only a small amount of language data. Prior work has made progress on this using vision-language models or by jointly training language-goal-conditioned policies, but so far neither method has scaled effectively to real-world robot tasks without significant human annotation. Our method achieves robust performance in the real world by learning an embedding from the labeled data that aligns language not to the goal image, but rather to the desired change between the start and goal images that the instruction corresponds to. We then train a policy on this embedding: the policy benefits from all the unlabeled data, but the aligned embedding provides an interface for language to steer the policy. We show instruction following across a variety of manipulation tasks in different scenes, with generalization to language instructions outside of the labeled data. Videos and code for our approach can be found on our website: http://tiny.cc/grif .
Reconstructing Three-decade Global Fine-Grained Nighttime Light Observations by a New Super-Resolution Framework
Jul 14, 2023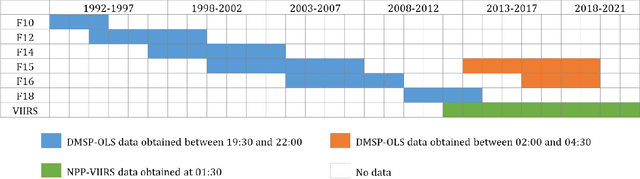
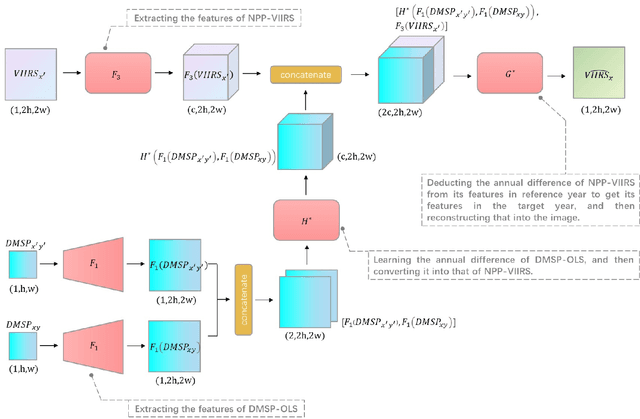
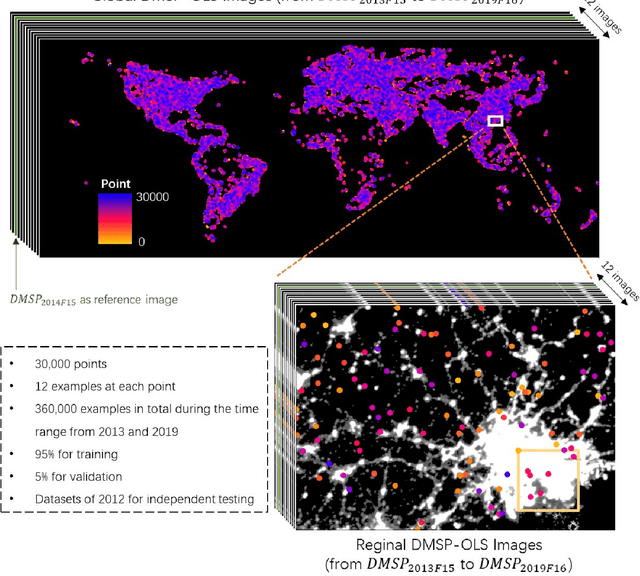
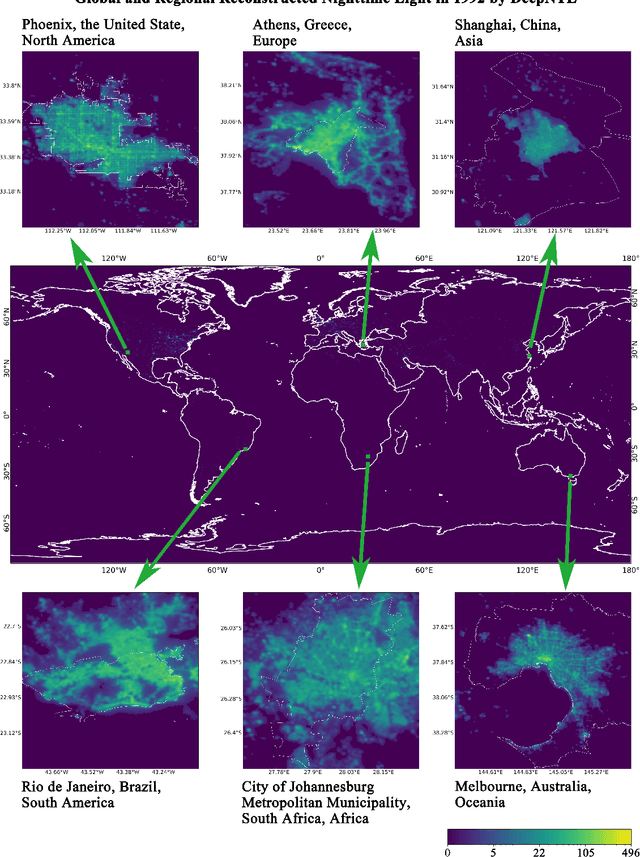
Satellite-collected nighttime light provides a unique perspective on human activities, including urbanization, population growth, and epidemics. Yet, long-term and fine-grained nighttime light observations are lacking, leaving the analysis and applications of decades of light changes in urban facilities undeveloped. To fill this gap, we developed an innovative framework and used it to design a new super-resolution model that reconstructs low-resolution nighttime light data into high resolution. The validation of one billion data points shows that the correlation coefficient of our model at the global scale reaches 0.873, which is significantly higher than that of other existing models (maximum = 0.713). Our model also outperforms existing models at the national and urban scales. Furthermore, through an inspection of airports and roads, only our model's image details can reveal the historical development of these facilities. We provide the long-term and fine-grained nighttime light observations to promote research on human activities. The dataset is available at \url{https://doi.org/10.5281/zenodo.7859205}.
AdaptiveClick: Clicks-aware Transformer with Adaptive Focal Loss for Interactive Image Segmentation
May 07, 2023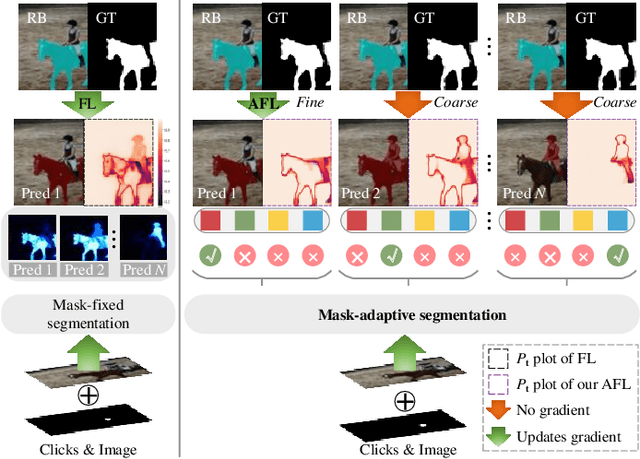
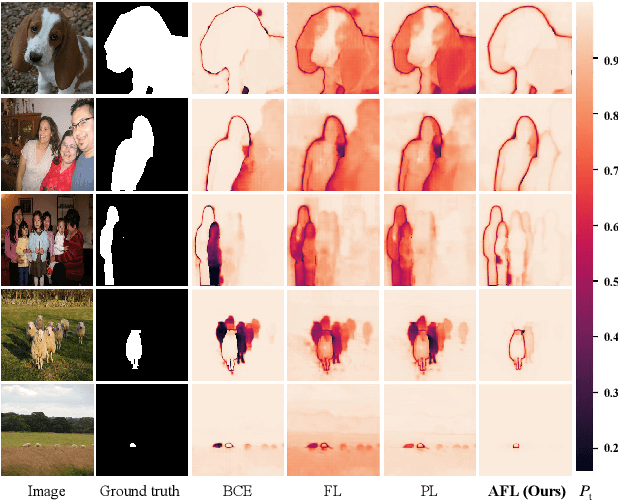
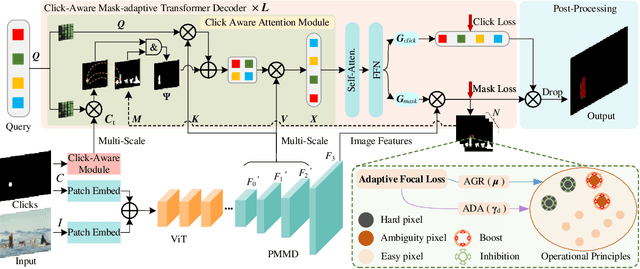

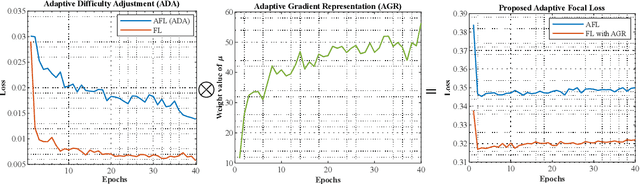
Interactive Image Segmentation (IIS) has emerged as a promising technique for decreasing annotation time. Substantial progress has been made in pre- and post-processing for IIS, but the critical issue of interaction ambiguity notably hindering segmentation quality, has been under-researched. To address this, we introduce AdaptiveClick -- a clicks-aware transformer incorporating an adaptive focal loss, which tackles annotation inconsistencies with tools for mask- and pixel-level ambiguity resolution. To the best of our knowledge, AdaptiveClick is the first transformer-based, mask-adaptive segmentation framework for IIS. The key ingredient of our method is the Clicks-aware Mask-adaptive Transformer Decoder (CAMD), which enhances the interaction between clicks and image features. Additionally, AdaptiveClick enables pixel-adaptive differentiation of hard and easy samples in the decision space, independent of their varying distributions. This is primarily achieved by optimizing a generalized Adaptive Focal Loss (AFL) with a theoretical guarantee, where two adaptive coefficients control the ratio of gradient values for hard and easy pixels. Our analysis reveals that the commonly used Focal and BCE losses can be considered special cases of the proposed AFL loss. With a plain ViT backbone, extensive experimental results on nine datasets demonstrate the superiority of AdaptiveClick compared to state-of-the-art methods. Code will be publicly available at https://github.com/lab206/AdaptiveClick.
Multi-Level Global Context Cross Consistency Model for Semi-Supervised Ultrasound Image Segmentation with Diffusion Model
May 17, 2023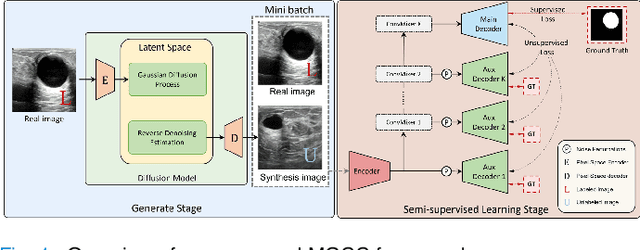
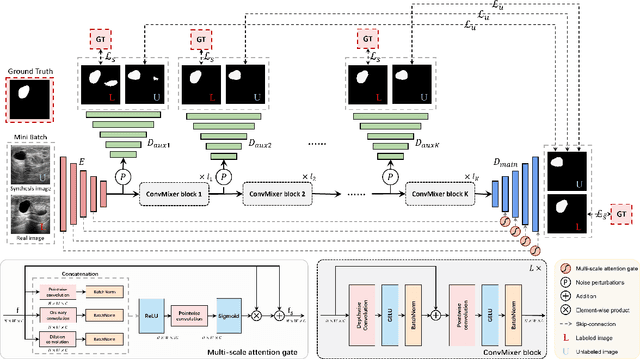
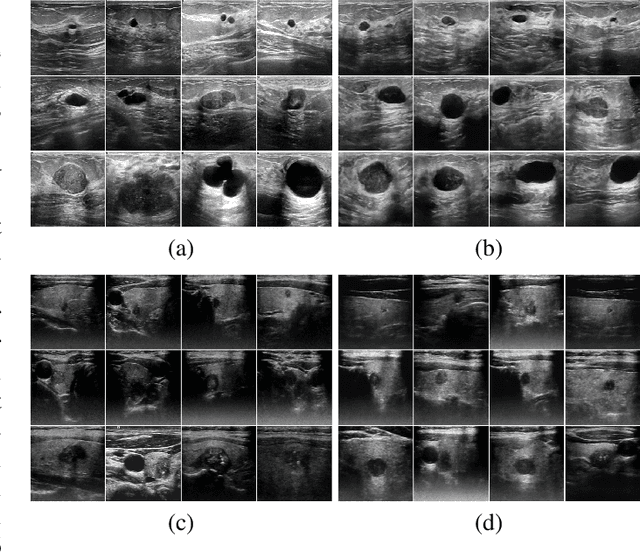
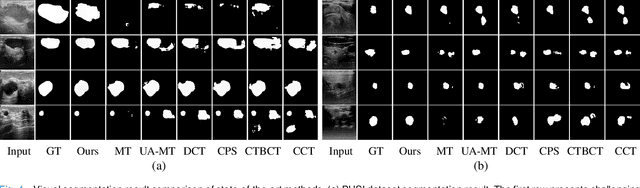
Medical image segmentation is a critical step in computer-aided diagnosis, and convolutional neural networks are popular segmentation networks nowadays. However, the inherent local operation characteristics make it difficult to focus on the global contextual information of lesions with different positions, shapes, and sizes. Semi-supervised learning can be used to learn from both labeled and unlabeled samples, alleviating the burden of manual labeling. However, obtaining a large number of unlabeled images in medical scenarios remains challenging. To address these issues, we propose a Multi-level Global Context Cross-consistency (MGCC) framework that uses images generated by a Latent Diffusion Model (LDM) as unlabeled images for semi-supervised learning. The framework involves of two stages. In the first stage, a LDM is used to generate synthetic medical images, which reduces the workload of data annotation and addresses privacy concerns associated with collecting medical data. In the second stage, varying levels of global context noise perturbation are added to the input of the auxiliary decoder, and output consistency is maintained between decoders to improve the representation ability. Experiments conducted on open-source breast ultrasound and private thyroid ultrasound datasets demonstrate the effectiveness of our framework in bridging the probability distribution and the semantic representation of the medical image. Our approach enables the effective transfer of probability distribution knowledge to the segmentation network, resulting in improved segmentation accuracy. The code is available at https://github.com/FengheTan9/Multi-Level-Global-Context-Cross-Consistency.
Prompting Diffusion Representations for Cross-Domain Semantic Segmentation
Jul 05, 2023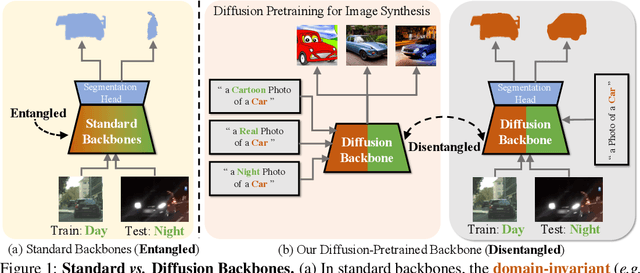
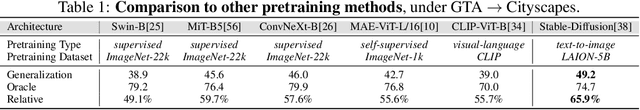
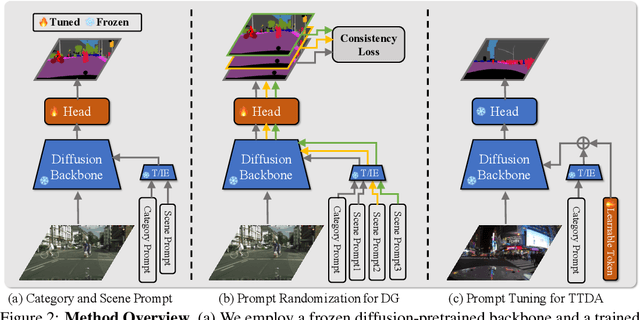
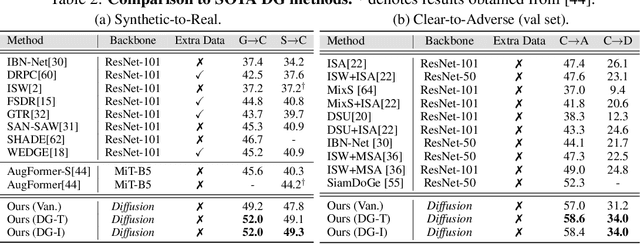
While originally designed for image generation, diffusion models have recently shown to provide excellent pretrained feature representations for semantic segmentation. Intrigued by this result, we set out to explore how well diffusion-pretrained representations generalize to new domains, a crucial ability for any representation. We find that diffusion-pretraining achieves extraordinary domain generalization results for semantic segmentation, outperforming both supervised and self-supervised backbone networks. Motivated by this, we investigate how to utilize the model's unique ability of taking an input prompt, in order to further enhance its cross-domain performance. We introduce a scene prompt and a prompt randomization strategy to help further disentangle the domain-invariant information when training the segmentation head. Moreover, we propose a simple but highly effective approach for test-time domain adaptation, based on learning a scene prompt on the target domain in an unsupervised manner. Extensive experiments conducted on four synthetic-to-real and clear-to-adverse weather benchmarks demonstrate the effectiveness of our approaches. Without resorting to any complex techniques, such as image translation, augmentation, or rare-class sampling, we set a new state-of-the-art on all benchmarks. Our implementation will be publicly available at \url{https://github.com/ETHRuiGong/PTDiffSeg}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge