Shufei Zhang
PolyReal: A Benchmark for Real-World Polymer Science Workflows
Apr 03, 2026Abstract:Multimodal Large Language Models (MLLMs) excel in general domains but struggle with complex, real-world science. We posit that polymer science, an interdisciplinary field spanning chemistry, physics, biology, and engineering, is an ideal high-stakes testbed due to its diverse multimodal data. Yet, existing benchmarks related to polymer science largely overlook real-world workflows, limiting their practical utility and failing to systematically evaluate MLLMs across the full, practice-grounded lifecycle of experimentation. We introduce PolyReal, a novel multimodal benchmark grounded in real-world scientific practices to evaluate MLLMs on the full lifecycle of polymer experimentation. It covers five critical capabilities: (1) foundational knowledge application; (2) lab safety analysis; (3) experiment mechanism reasoning; (4) raw data extraction; and (5) performance & application exploration. Our evaluation of leading MLLMs on PolyReal reveals a capability imbalance. While models perform well on knowledge-intensive reasoning (e.g., Experiment Mechanism Reasoning), they drop sharply on practice-based tasks (e.g., Lab Safety Analysis and Raw Data Extraction). This exposes a severe gap between abstract scientific knowledge and its practical, context-dependent application, showing that these real-world tasks remain challenging for MLLMs. Thus, PolyReal helps address this evaluation gap and provides a practical benchmark for assessing AI systems in real-world scientific workflows.
Equivariant Evidential Deep Learning for Interatomic Potentials
Feb 11, 2026Abstract:Uncertainty quantification (UQ) is critical for assessing the reliability of machine learning interatomic potentials (MLIPs) in molecular dynamics (MD) simulations, identifying extrapolation regimes and enabling uncertainty-aware workflows such as active learning for training dataset construction. Existing UQ approaches for MLIPs are often limited by high computational cost or suboptimal performance. Evidential deep learning (EDL) provides a theoretically grounded single-model alternative that determines both aleatoric and epistemic uncertainty in a single forward pass. However, extending evidential formulations from scalar targets to vector-valued quantities such as atomic forces introduces substantial challenges, particularly in maintaining statistical self-consistency under rotational transformations. To address this, we propose \textit{Equivariant Evidential Deep Learning for Interatomic Potentials} ($\text{e}^2$IP), a backbone-agnostic framework that models atomic forces and their uncertainty jointly by representing uncertainty as a full $3\times3$ symmetric positive definite covariance tensor that transforms equivariantly under rotations. Experiments on diverse molecular benchmarks show that $\text{e}^2$IP provides a stronger accuracy-efficiency-reliability balance than the non-equivariant evidential baseline and the widely used ensemble method. It also achieves better data efficiency through the fully equivariant architecture while retaining single-model inference efficiency.
InternAgent-1.5: A Unified Agentic Framework for Long-Horizon Autonomous Scientific Discovery
Feb 09, 2026Abstract:We introduce InternAgent-1.5, a unified system designed for end-to-end scientific discovery across computational and empirical domains. The system is built on a structured architecture composed of three coordinated subsystems for generation, verification, and evolution. These subsystems are supported by foundational capabilities for deep research, solution optimization, and long horizon memory. The architecture allows InternAgent-1.5 to operate continuously across extended discovery cycles while maintaining coherent and improving behavior. It also enables the system to coordinate computational modeling and laboratory experimentation within a single unified system. We evaluate InternAgent-1.5 on scientific reasoning benchmarks such as GAIA, HLE, GPQA, and FrontierScience, and the system achieves leading performance that demonstrates strong foundational capabilities. Beyond these benchmarks, we further assess two categories of discovery tasks. In algorithm discovery tasks, InternAgent-1.5 autonomously designs competitive methods for core machine learning problems. In empirical discovery tasks, it executes complete computational or wet lab experiments and produces scientific findings in earth, life, biological, and physical domains. Overall, these results show that InternAgent-1.5 provides a general and scalable framework for autonomous scientific discovery.
SciEvalKit: An Open-source Evaluation Toolkit for Scientific General Intelligence
Dec 30, 2025Abstract:We introduce SciEvalKit, a unified benchmarking toolkit designed to evaluate AI models for science across a broad range of scientific disciplines and task capabilities. Unlike general-purpose evaluation platforms, SciEvalKit focuses on the core competencies of scientific intelligence, including Scientific Multimodal Perception, Scientific Multimodal Reasoning, Scientific Multimodal Understanding, Scientific Symbolic Reasoning, Scientific Code Generation, Science Hypothesis Generation and Scientific Knowledge Understanding. It supports six major scientific domains, spanning from physics and chemistry to astronomy and materials science. SciEvalKit builds a foundation of expert-grade scientific benchmarks, curated from real-world, domain-specific datasets, ensuring that tasks reflect authentic scientific challenges. The toolkit features a flexible, extensible evaluation pipeline that enables batch evaluation across models and datasets, supports custom model and dataset integration, and provides transparent, reproducible, and comparable results. By bridging capability-based evaluation and disciplinary diversity, SciEvalKit offers a standardized yet customizable infrastructure to benchmark the next generation of scientific foundation models and intelligent agents. The toolkit is open-sourced and actively maintained to foster community-driven development and progress in AI4Science.
An Agentic Framework for Autonomous Materials Computation
Dec 22, 2025



Abstract:Large Language Models (LLMs) have emerged as powerful tools for accelerating scientific discovery, yet their static knowledge and hallucination issues hinder autonomous research applications. Recent advances integrate LLMs into agentic frameworks, enabling retrieval, reasoning, and tool use for complex scientific workflows. Here, we present a domain-specialized agent designed for reliable automation of first-principles materials computations. By embedding domain expertise, the agent ensures physically coherent multi-step workflows and consistently selects convergent, well-posed parameters, thereby enabling reliable end-to-end computational execution. A new benchmark of diverse computational tasks demonstrates that our system significantly outperforms standalone LLMs in both accuracy and robustness. This work establishes a verifiable foundation for autonomous computational experimentation and represents a key step toward fully automated scientific discovery.
Probing Scientific General Intelligence of LLMs with Scientist-Aligned Workflows
Dec 18, 2025Abstract:Despite advances in scientific AI, a coherent framework for Scientific General Intelligence (SGI)-the ability to autonomously conceive, investigate, and reason across scientific domains-remains lacking. We present an operational SGI definition grounded in the Practical Inquiry Model (PIM: Deliberation, Conception, Action, Perception) and operationalize it via four scientist-aligned tasks: deep research, idea generation, dry/wet experiments, and experimental reasoning. SGI-Bench comprises over 1,000 expert-curated, cross-disciplinary samples inspired by Science's 125 Big Questions, enabling systematic evaluation of state-of-the-art LLMs. Results reveal gaps: low exact match (10--20%) in deep research despite step-level alignment; ideas lacking feasibility and detail; high code executability but low execution result accuracy in dry experiments; low sequence fidelity in wet protocols; and persistent multimodal comparative-reasoning challenges. We further introduce Test-Time Reinforcement Learning (TTRL), which optimizes retrieval-augmented novelty rewards at inference, enhancing hypothesis novelty without reference answer. Together, our PIM-grounded definition, workflow-centric benchmark, and empirical insights establish a foundation for AI systems that genuinely participate in scientific discovery.
Plan Then Action:High-Level Planning Guidance Reinforcement Learning for LLM Reasoning
Oct 02, 2025Abstract:Large language models (LLMs) have demonstrated remarkable reasoning abilities in complex tasks, often relying on Chain-of-Thought (CoT) reasoning. However, due to their autoregressive token-level generation, the reasoning process is largely constrained to local decision-making and lacks global planning. This limitation frequently results in redundant, incoherent, or inaccurate reasoning, which significantly degrades overall performance. Existing approaches, such as tree-based algorithms and reinforcement learning (RL), attempt to address this issue but suffer from high computational costs and often fail to produce optimal reasoning trajectories. To tackle this challenge, we propose Plan-Then-Action Enhanced Reasoning with Group Relative Policy Optimization PTA-GRPO, a two-stage framework designed to improve both high-level planning and fine-grained CoT reasoning. In the first stage, we leverage advanced LLMs to distill CoT into compact high-level guidance, which is then used for supervised fine-tuning (SFT). In the second stage, we introduce a guidance-aware RL method that jointly optimizes the final output and the quality of high-level guidance, thereby enhancing reasoning effectiveness. We conduct extensive experiments on multiple mathematical reasoning benchmarks, including MATH, AIME2024, AIME2025, and AMC, across diverse base models such as Qwen2.5-7B-Instruct, Qwen3-8B, Qwen3-14B, and LLaMA3.2-3B. Experimental results demonstrate that PTA-GRPO consistently achieves stable and significant improvements across different models and tasks, validating its effectiveness and generalization.
ChemBOMAS: Accelerated BO in Chemistry with LLM-Enhanced Multi-Agent System
Sep 10, 2025Abstract:The efficiency of Bayesian optimization (BO) in chemistry is often hindered by sparse experimental data and complex reaction mechanisms. To overcome these limitations, we introduce ChemBOMAS, a new framework named LLM-Enhanced Multi-Agent System for accelerating BO in chemistry. ChemBOMAS's optimization process is enhanced by LLMs and synergistically employs two strategies: knowledge-driven coarse-grained optimization and data-driven fine-grained optimization. First, in the knowledge-driven coarse-grained optimization stage, LLMs intelligently decompose the vast search space by reasoning over existing chemical knowledge to identify promising candidate regions. Subsequently, in the data-driven fine-grained optimization stage, LLMs enhance the BO process within these candidate regions by generating pseudo-data points, thereby improving data utilization efficiency and accelerating convergence. Benchmark evaluations** further confirm that ChemBOMAS significantly enhances optimization effectiveness and efficiency compared to various BO algorithms. Importantly, the practical utility of ChemBOMAS was validated through wet-lab experiments conducted under pharmaceutical industry protocols, targeting conditional optimization for a previously unreported and challenging chemical reaction. In the wet experiment, ChemBOMAS achieved an optimal objective value of 96%. This was substantially higher than the 15% achieved by domain experts. This real-world success, together with strong performance on benchmark evaluations, highlights ChemBOMAS as a powerful tool to accelerate chemical discovery.
CMPhysBench: A Benchmark for Evaluating Large Language Models in Condensed Matter Physics
Aug 25, 2025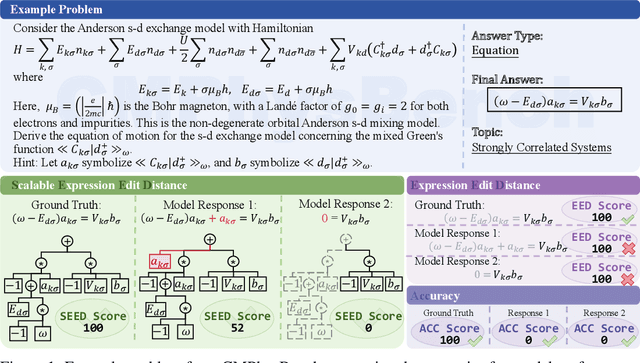
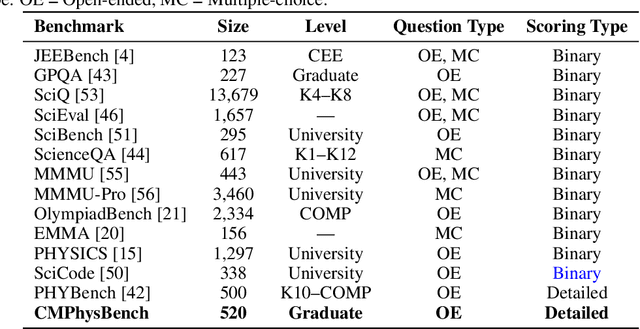
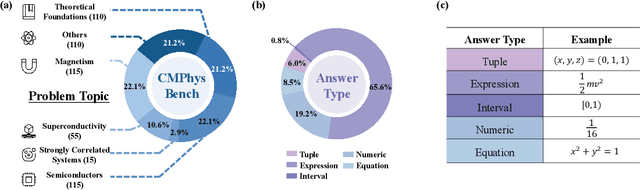
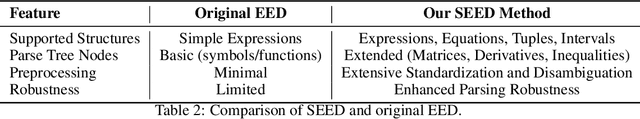
Abstract:We introduce CMPhysBench, designed to assess the proficiency of Large Language Models (LLMs) in Condensed Matter Physics, as a novel Benchmark. CMPhysBench is composed of more than 520 graduate-level meticulously curated questions covering both representative subfields and foundational theoretical frameworks of condensed matter physics, such as magnetism, superconductivity, strongly correlated systems, etc. To ensure a deep understanding of the problem-solving process,we focus exclusively on calculation problems, requiring LLMs to independently generate comprehensive solutions. Meanwhile, leveraging tree-based representations of expressions, we introduce the Scalable Expression Edit Distance (SEED) score, which provides fine-grained (non-binary) partial credit and yields a more accurate assessment of similarity between prediction and ground-truth. Our results show that even the best models, Grok-4, reach only 36 average SEED score and 28% accuracy on CMPhysBench, underscoring a significant capability gap, especially for this practical and frontier domain relative to traditional physics. The code anddataset are publicly available at https://github.com/CMPhysBench/CMPhysBench.
Mol-R1: Towards Explicit Long-CoT Reasoning in Molecule Discovery
Aug 11, 2025Abstract:Large language models (LLMs), especially Explicit Long Chain-of-Thought (CoT) reasoning models like DeepSeek-R1 and QWQ, have demonstrated powerful reasoning capabilities, achieving impressive performance in commonsense reasoning and mathematical inference. Despite their effectiveness, Long-CoT reasoning models are often criticized for their limited ability and low efficiency in knowledge-intensive domains such as molecule discovery. Success in this field requires a precise understanding of domain knowledge, including molecular structures and chemical principles, which is challenging due to the inherent complexity of molecular data and the scarcity of high-quality expert annotations. To bridge this gap, we introduce Mol-R1, a novel framework designed to improve explainability and reasoning performance of R1-like Explicit Long-CoT reasoning LLMs in text-based molecule generation. Our approach begins with a high-quality reasoning dataset curated through Prior Regulation via In-context Distillation (PRID), a dedicated distillation strategy to effectively generate paired reasoning traces guided by prior regulations. Building upon this, we introduce MoIA, Molecular Iterative Adaptation, a sophisticated training strategy that iteratively combines Supervised Fine-tuning (SFT) with Reinforced Policy Optimization (RPO), tailored to boost the reasoning performance of R1-like reasoning models for molecule discovery. Finally, we examine the performance of Mol-R1 in the text-based molecule reasoning generation task, showing superior performance against existing baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge