Yuqiang Li
PolyReal: A Benchmark for Real-World Polymer Science Workflows
Apr 03, 2026Abstract:Multimodal Large Language Models (MLLMs) excel in general domains but struggle with complex, real-world science. We posit that polymer science, an interdisciplinary field spanning chemistry, physics, biology, and engineering, is an ideal high-stakes testbed due to its diverse multimodal data. Yet, existing benchmarks related to polymer science largely overlook real-world workflows, limiting their practical utility and failing to systematically evaluate MLLMs across the full, practice-grounded lifecycle of experimentation. We introduce PolyReal, a novel multimodal benchmark grounded in real-world scientific practices to evaluate MLLMs on the full lifecycle of polymer experimentation. It covers five critical capabilities: (1) foundational knowledge application; (2) lab safety analysis; (3) experiment mechanism reasoning; (4) raw data extraction; and (5) performance & application exploration. Our evaluation of leading MLLMs on PolyReal reveals a capability imbalance. While models perform well on knowledge-intensive reasoning (e.g., Experiment Mechanism Reasoning), they drop sharply on practice-based tasks (e.g., Lab Safety Analysis and Raw Data Extraction). This exposes a severe gap between abstract scientific knowledge and its practical, context-dependent application, showing that these real-world tasks remain challenging for MLLMs. Thus, PolyReal helps address this evaluation gap and provides a practical benchmark for assessing AI systems in real-world scientific workflows.
Project Imaging-X: A Survey of 1000+ Open-Access Medical Imaging Datasets for Foundation Model Development
Mar 29, 2026Abstract:Foundation models have demonstrated remarkable success across diverse domains and tasks, primarily due to the thrive of large-scale, diverse, and high-quality datasets. However, in the field of medical imaging, the curation and assembling of such medical datasets are highly challenging due to the reliance on clinical expertise and strict ethical and privacy constraints, resulting in a scarcity of large-scale unified medical datasets and hindering the development of powerful medical foundation models. In this work, we present the largest survey to date of medical image datasets, covering over 1,000 open-access datasets with a systematic catalog of their modalities, tasks, anatomies, annotations, limitations, and potential for integration. Our analysis exposes a landscape that is modest in scale, fragmented across narrowly scoped tasks, and unevenly distributed across organs and modalities, which in turn limits the utility of existing medical image datasets for developing versatile and robust medical foundation models. To turn fragmentation into scale, we propose a metadata-driven fusion paradigm (MDFP) that integrates public datasets with shared modalities or tasks, thereby transforming multiple small data silos into larger, more coherent resources. Building on MDFP, we release an interactive discovery portal that enables end-to-end, automated medical image dataset integration, and compile all surveyed datasets into a unified, structured table that clearly summarizes their key characteristics and provides reference links, offering the community an accessible and comprehensive repository. By charting the current terrain and offering a principled path to dataset consolidation, our survey provides a practical roadmap for scaling medical imaging corpora, supporting faster data discovery, more principled dataset creation, and more capable medical foundation models.
WIST: Web-Grounded Iterative Self-Play Tree for Domain-Targeted Reasoning Improvement
Mar 22, 2026Abstract:Recent progress in reinforcement learning with verifiable rewards (RLVR) offers a practical path to self-improvement of language models, but existing methods face a key trade-off: endogenous self-play can drift over iterations, while corpus-grounded approaches rely on curated data environments. We present \textbf{WIST}, a \textbf{W}eb-grounded \textbf{I}terative \textbf{S}elf-play \textbf{T}ree framework for domain-targeted reasoning improvement that learns directly from the open web without requiring any pre-arranged domain corpus. WIST incrementally expands a domain tree for exploration, and retrieves and cleans path-consistent web corpus to construct a controllable training environment. It then performs Challenger--Solver self-play with verifiable rewards, and feeds learnability signals back to update node posteriors and guide subsequent exploration through an adaptive curriculum. Across four backbones, WIST consistently improves over the base models and typically outperforms both purely endogenous self-evolution and corpus-grounded self-play baselines, with the Overall gains reaching \textbf{+9.8} (\textit{Qwen3-4B-Base}) and \textbf{+9.7} (\textit{OctoThinker-8B}). WIST is also domain-steerable, improving \textit{Qwen3-8B-Base} by \textbf{+14.79} in medicine and \textit{Qwen3-4B-Base} by \textbf{+5.28} on PhyBench. Ablations further confirm the importance of WIST's key components for stable open-web learning. Our Code is available at https://github.com/lfy-123/WIST.
Does the Question Really Matter? Training-Free Data Selection for Vision-Language SFT
Mar 10, 2026Abstract:Visual instruction tuning is crucial for improving vision-language large models (VLLMs). However, many samples can be solved via linguistic patterns or common-sense shortcuts, without genuine cross-modal reasoning, limiting the effectiveness of multimodal learning. Prior data selection methods often rely on costly proxy model training and focus on difficulty or diversity, failing to capture a sample's true contribution to vision-language joint reasoning. In this paper, we propose CVS, a training-free data selection method based on the insight that, for high-quality multimodal samples, introducing the question should substantially alter the model's assessment of answer validity given an image. CVS leverages a frozen VLLM as an evaluator and measures the discrepancy in answer validity with and without conditioning on the question, enabling the identification of samples that require vision-language joint reasoning while filtering semantic-conflict noise. Experiments on Vision-Flan and The Cauldron show that CVS achieves solid performance across datasets. On Vision-Flan, CVS outperforms full-data training by 3.5% and 4.8% using only 10% and 15% of the data, respectively, and remains robust on the highly heterogeneous Cauldron dataset. Moreover, CVS reduces computational cost by 17.3% and 44.4% compared to COINCIDE and XMAS.
NMRTrans: Structure Elucidation from Experimental NMR Spectra via Set Transformers
Feb 10, 2026Abstract:Nuclear Magnetic Resonance (NMR) spectroscopy is fundamental for molecular structure elucidation, yet interpreting spectra at scale remains time-consuming and highly expertise-dependent. While recent spectrum-as-language modeling and retrieval-based methods have shown promise, they rely heavily on large corpora of computed spectra and exhibit notable performance drops when applied to experimental measurements. To address these issues, we build NMRSpec, a large-scale corpus of experimental $^1$H and $^{13}$C spectra mined from chemical literature, and propose NMRTrans, which models spectra as unordered peak sets and aligns the model's inductive bias with the physical nature of NMR. To our best knowledge, NMRTrans is the first NMR Transformer trained solely on large-scale experimental spectra and achieves state-of-the-art performance on experimental benchmarks, improving Top-10 Accuracy over the strongest baseline by +17.82 points (61.15% vs. 43.33%), and underscoring the importance of experimental data and structure-aware architectures for reliable NMR structure elucidation.
MARTI-MARS$^2$: Scaling Multi-Agent Self-Search via Reinforcement Learning for Code Generation
Feb 08, 2026Abstract:While the complex reasoning capability of Large Language Models (LLMs) has attracted significant attention, single-agent systems often encounter inherent performance ceilings in complex tasks such as code generation. Multi-agent collaboration offers a promising avenue to transcend these boundaries. However, existing frameworks typically rely on prompt-based test-time interactions or multi-role configurations trained with homogeneous parameters, limiting error correction capabilities and strategic diversity. In this paper, we propose a Multi-Agent Reinforced Training and Inference Framework with Self-Search Scaling (MARTI-MARS2), which integrates policy learning with multi-agent tree search by formulating the multi-agent collaborative exploration process as a dynamic and learnable environment. By allowing agents to iteratively explore and refine within the environment, the framework facilitates evolution from parameter-sharing homogeneous multi-role training to heterogeneous multi-agent training, breaking through single-agent capability limits. We also introduce an efficient inference strategy MARTI-MARS2-T+ to fully exploit the scaling potential of multi-agent collaboration at test time. We conduct extensive experiments across varied model scales (8B, 14B, and 32B) on challenging code generation benchmarks. Utilizing two collaborating 32B models, MARTI-MARS2 achieves 77.7%, outperforming strong baselines like GPT-5.1. Furthermore, MARTI-MARS2 reveals a novel scaling law: shifting from single-agent to homogeneous multi-role and ultimately to heterogeneous multi-agent paradigms progressively yields higher RL performance ceilings, robust TTS capabilities, and greater policy diversity, suggesting that policy diversity is critical for scaling intelligence via multi-agent reinforcement learning.
MolAct: An Agentic RL Framework for Molecular Editing and Property Optimization
Dec 24, 2025Abstract:Molecular editing and optimization are multi-step problems that require iteratively improving properties while keeping molecules chemically valid and structurally similar. We frame both tasks as sequential, tool-guided decisions and introduce MolAct, an agentic reinforcement learning framework that employs a two-stage training paradigm: first building editing capability, then optimizing properties while reusing the learned editing behaviors. To the best of our knowledge, this is the first work to formalize molecular design as an Agentic Reinforcement Learning problem, where an LLM agent learns to interleave reasoning, tool-use, and molecular optimization. The framework enables agents to interact in multiple turns, invoking chemical tools for validity checking, property assessment, and similarity control, and leverages their feedback to refine subsequent edits. We instantiate the MolAct framework to train two model families: MolEditAgent for molecular editing tasks and MolOptAgent for molecular optimization tasks. In molecular editing, MolEditAgent-7B delivers 100, 95, and 98 valid add, delete, and substitute edits, outperforming strong closed "thinking" baselines such as DeepSeek-R1; MolEditAgent-3B approaches the performance of much larger open "thinking" models like Qwen3-32B-think. In molecular optimization, MolOptAgent-7B (trained on MolEditAgent-7B) surpasses the best closed "thinking" baseline (e.g., Claude 3.7) on LogP and remains competitive on solubility, while maintaining balanced performance across other objectives. These results highlight that treating molecular design as a multi-step, tool-augmented process is key to reliable and interpretable improvements.
An Agentic Framework for Autonomous Materials Computation
Dec 22, 2025



Abstract:Large Language Models (LLMs) have emerged as powerful tools for accelerating scientific discovery, yet their static knowledge and hallucination issues hinder autonomous research applications. Recent advances integrate LLMs into agentic frameworks, enabling retrieval, reasoning, and tool use for complex scientific workflows. Here, we present a domain-specialized agent designed for reliable automation of first-principles materials computations. By embedding domain expertise, the agent ensures physically coherent multi-step workflows and consistently selects convergent, well-posed parameters, thereby enabling reliable end-to-end computational execution. A new benchmark of diverse computational tasks demonstrates that our system significantly outperforms standalone LLMs in both accuracy and robustness. This work establishes a verifiable foundation for autonomous computational experimentation and represents a key step toward fully automated scientific discovery.
Revisiting the Broken Symmetry Phase of Solid Hydrogen: A Neural Network Variational Monte Carlo Study
Dec 19, 2025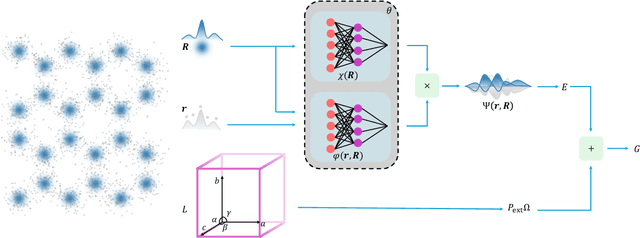
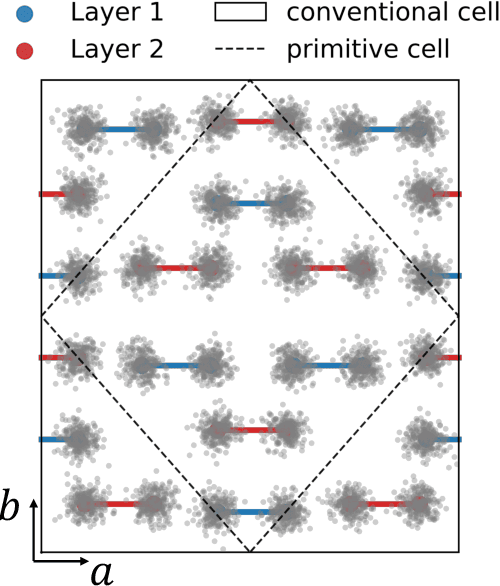
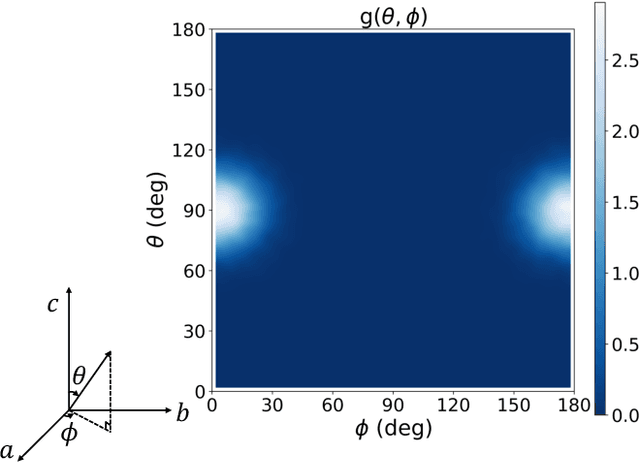
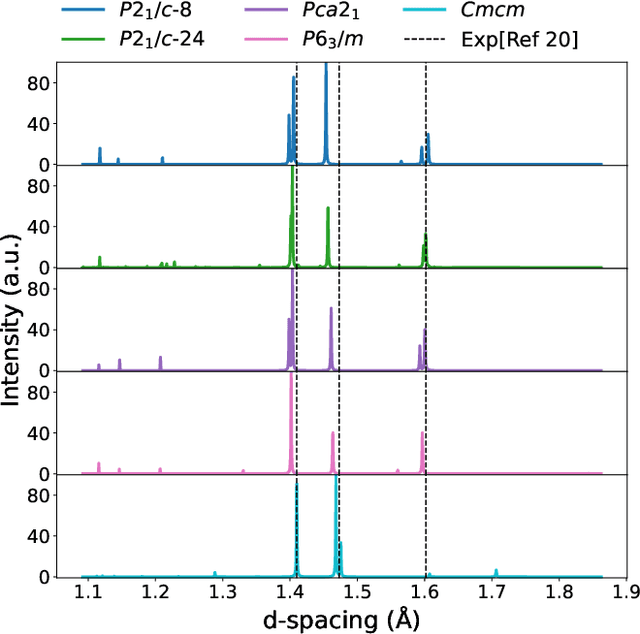
Abstract:The crystal structure of high-pressure solid hydrogen remains a fundamental open problem. Although the research frontier has mostly shifted toward ultra-high pressure phases above 400 GPa, we show that even the broken symmetry phase observed around 130~GPa requires revisiting due to its intricate coupling of electronic and nuclear degrees of freedom. Here, we develop a first principle quantum Monte Carlo framework based on a deep neural network wave function that treats both electrons and nuclei quantum mechanically within the constant pressure ensemble. Our calculations reveal an unreported ground-state structure candidate for the broken symmetry phase with $Cmcm$ space group symmetry, and we test its stability up to 96 atoms. The predicted structure quantitatively matches the experimental equation of state and X-ray diffraction patterns. Furthermore, our group-theoretical analysis shows that the $Cmcm$ structure is compatible with existing Raman and infrared spectroscopic data. Crucially, static density functional theory calculation reveals the $Cmcm$ structure as a dynamically unstable saddle point on the Born-Oppenheimer potential energy surface, demonstrating that a full quantum many-body treatment of the problem is necessary. These results shed new light on the phase diagram of high-pressure hydrogen and call for further experimental verifications.
Probing Scientific General Intelligence of LLMs with Scientist-Aligned Workflows
Dec 18, 2025Abstract:Despite advances in scientific AI, a coherent framework for Scientific General Intelligence (SGI)-the ability to autonomously conceive, investigate, and reason across scientific domains-remains lacking. We present an operational SGI definition grounded in the Practical Inquiry Model (PIM: Deliberation, Conception, Action, Perception) and operationalize it via four scientist-aligned tasks: deep research, idea generation, dry/wet experiments, and experimental reasoning. SGI-Bench comprises over 1,000 expert-curated, cross-disciplinary samples inspired by Science's 125 Big Questions, enabling systematic evaluation of state-of-the-art LLMs. Results reveal gaps: low exact match (10--20%) in deep research despite step-level alignment; ideas lacking feasibility and detail; high code executability but low execution result accuracy in dry experiments; low sequence fidelity in wet protocols; and persistent multimodal comparative-reasoning challenges. We further introduce Test-Time Reinforcement Learning (TTRL), which optimizes retrieval-augmented novelty rewards at inference, enhancing hypothesis novelty without reference answer. Together, our PIM-grounded definition, workflow-centric benchmark, and empirical insights establish a foundation for AI systems that genuinely participate in scientific discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge