Qi Dou
for the ALFA study
Robust Medical Image Classification from Noisy Labeled Data with Global and Local Representation Guided Co-training
May 10, 2022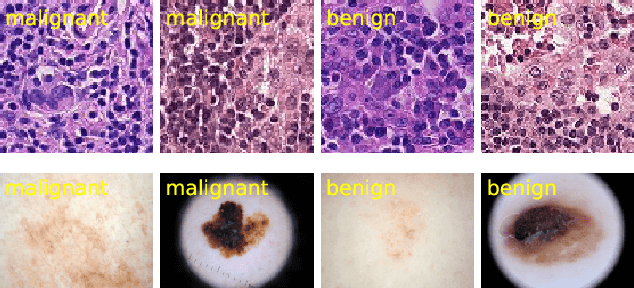
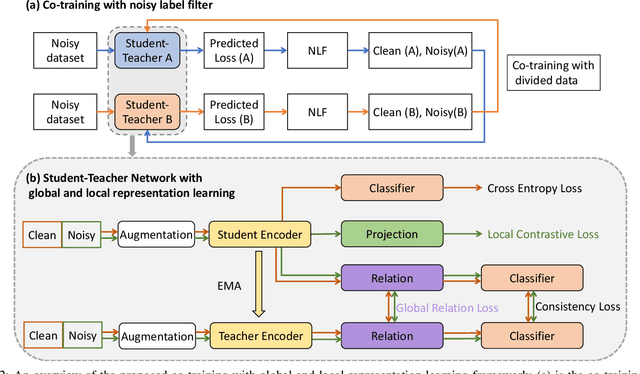
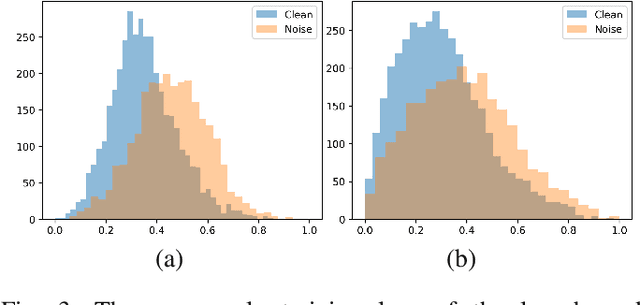
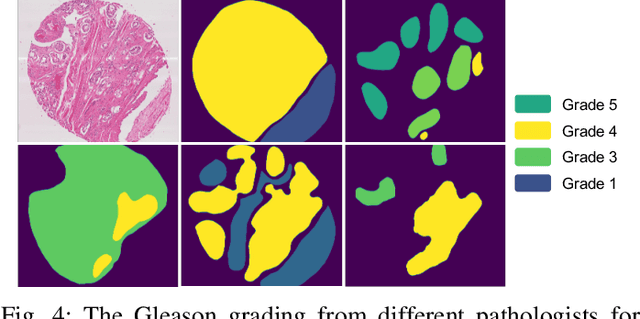
Abstract:Deep neural networks have achieved remarkable success in a wide variety of natural image and medical image computing tasks. However, these achievements indispensably rely on accurately annotated training data. If encountering some noisy-labeled images, the network training procedure would suffer from difficulties, leading to a sub-optimal classifier. This problem is even more severe in the medical image analysis field, as the annotation quality of medical images heavily relies on the expertise and experience of annotators. In this paper, we propose a novel collaborative training paradigm with global and local representation learning for robust medical image classification from noisy-labeled data to combat the lack of high quality annotated medical data. Specifically, we employ the self-ensemble model with a noisy label filter to efficiently select the clean and noisy samples. Then, the clean samples are trained by a collaborative training strategy to eliminate the disturbance from imperfect labeled samples. Notably, we further design a novel global and local representation learning scheme to implicitly regularize the networks to utilize noisy samples in a self-supervised manner. We evaluated our proposed robust learning strategy on four public medical image classification datasets with three types of label noise,ie,random noise, computer-generated label noise, and inter-observer variability noise. Our method outperforms other learning from noisy label methods and we also conducted extensive experiments to analyze each component of our method.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
IOP-FL: Inside-Outside Personalization for Federated Medical Image Segmentation
Apr 16, 2022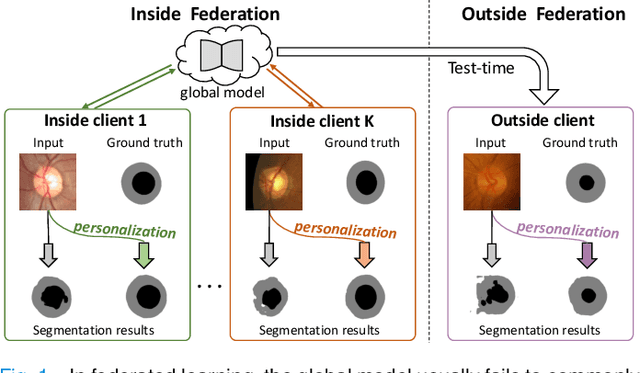
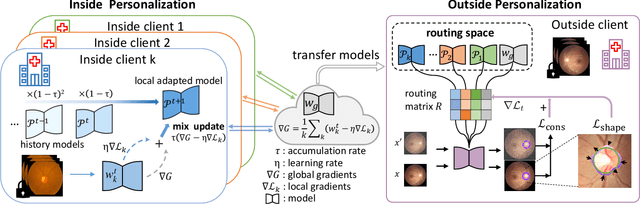
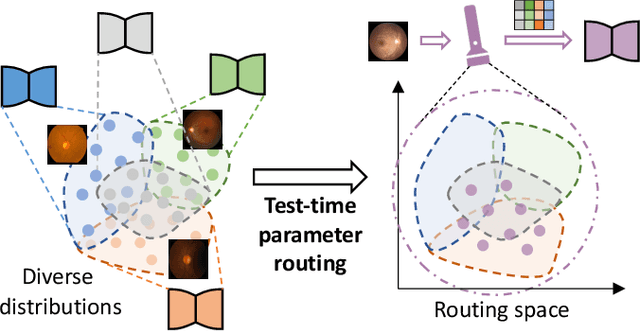
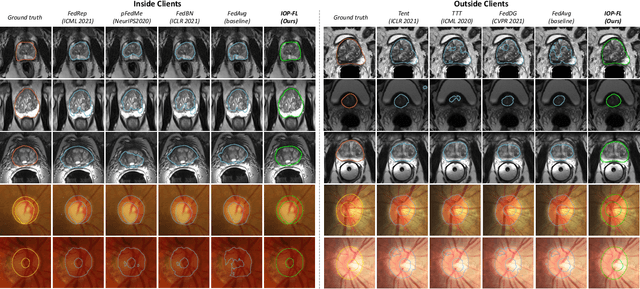
Abstract:Federated learning (FL) allows multiple medical institutions to collaboratively learn a global model without centralizing all clients data. It is difficult, if possible at all, for such a global model to commonly achieve optimal performance for each individual client, due to the heterogeneity of medical data from various scanners and patient demographics. This problem becomes even more significant when deploying the global model to unseen clients outside the FL with new distributions not presented during federated training. To optimize the prediction accuracy of each individual client for critical medical tasks, we propose a novel unified framework for both Inside and Outside model Personalization in FL (IOP-FL). Our inside personalization is achieved by a lightweight gradient-based approach that exploits the local adapted model for each client, by accumulating both the global gradients for common knowledge and local gradients for client-specific optimization. Moreover, and importantly, the obtained local personalized models and the global model can form a diverse and informative routing space to personalize a new model for outside FL clients. Hence, we design a new test-time routing scheme inspired by the consistency loss with a shape constraint to dynamically incorporate the models, given the distribution information conveyed by the test data. Our extensive experimental results on two medical image segmentation tasks present significant improvements over SOTA methods on both inside and outside personalization, demonstrating the great potential of our IOP-FL scheme for clinical practice. Code will be released at https://github.com/med-air/IOP-FL.
Sim-to-Real 6D Object Pose Estimation via Iterative Self-training for Robotic Bin-picking
Apr 14, 2022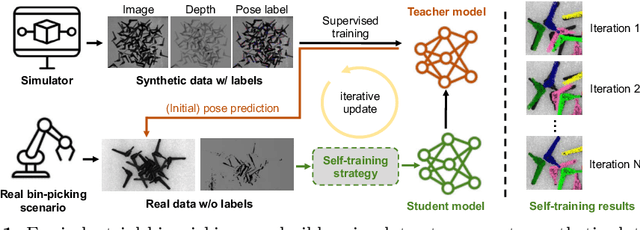
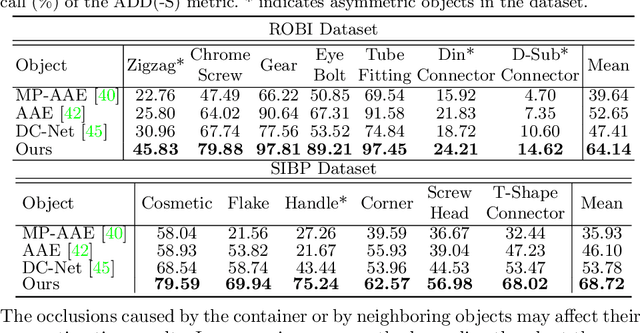
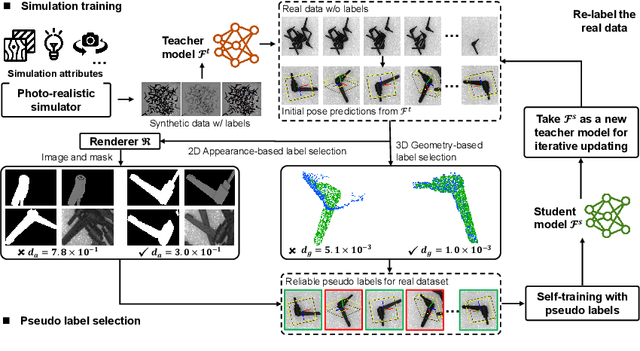
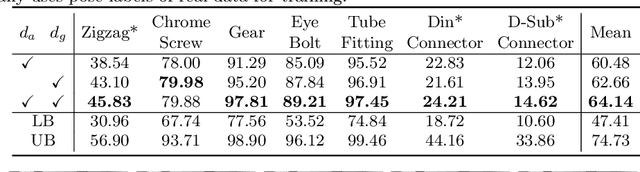
Abstract:In this paper, we propose an iterative self-training framework for sim-to-real 6D object pose estimation to facilitate cost-effective robotic grasping. Given a bin-picking scenario, we establish a photo-realistic simulator to synthesize abundant virtual data, and use this to train an initial pose estimation network. This network then takes the role of a teacher model, which generates pose predictions for unlabeled real data. With these predictions, we further design a comprehensive adaptive selection scheme to distinguish reliable results, and leverage them as pseudo labels to update a student model for pose estimation on real data. To continuously improve the quality of pseudo labels, we iterate the above steps by taking the trained student model as a new teacher and re-label real data using the refined teacher model. We evaluate our method on a public benchmark and our newly-released dataset, achieving an ADD(-S) improvement of 11.49% and 22.62% respectively. Our method is also able to improve robotic bin-picking success by 19.54%, demonstrating the potential of iterative sim-to-real solutions for robotic applications.
Rapid model transfer for medical image segmentation via iterative human-in-the-loop update: from labelled public to unlabelled clinical datasets for multi-organ segmentation in CT
Apr 13, 2022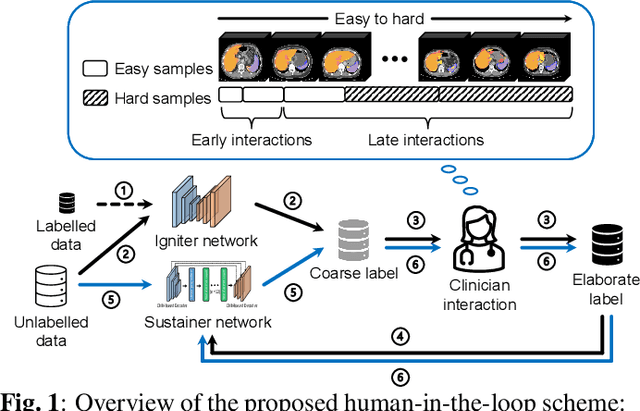
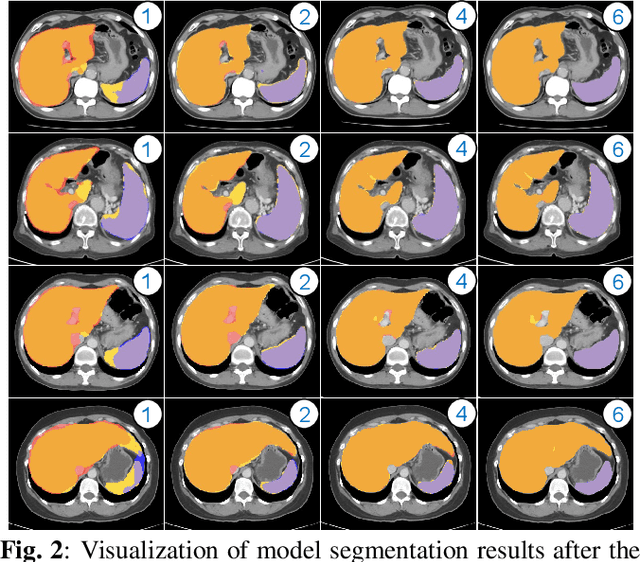
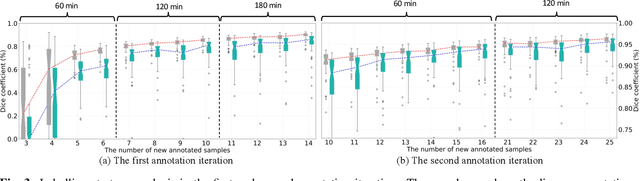
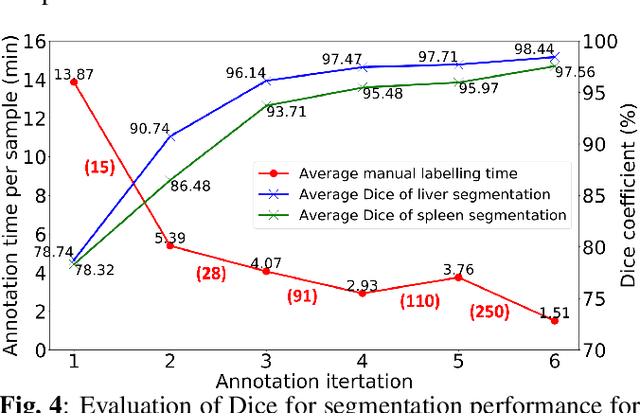
Abstract:Despite the remarkable success on medical image analysis with deep learning, it is still under exploration regarding how to rapidly transfer AI models from one dataset to another for clinical applications. This paper presents a novel and generic human-in-the-loop scheme for efficiently transferring a segmentation model from a small-scale labelled dataset to a larger-scale unlabelled dataset for multi-organ segmentation in CT. To achieve this, we propose to use an igniter network which can learn from a small-scale labelled dataset and generate coarse annotations to start the process of human-machine interaction. Then, we use a sustainer network for our larger-scale dataset, and iteratively updated it on the new annotated data. Moreover, we propose a flexible labelling strategy for the annotator to reduce the initial annotation workload. The model performance and the time cost of annotation in each subject evaluated on our private dataset are reported and analysed. The results show that our scheme can not only improve the performance by 19.7% on Dice, but also expedite the cost time of manual labelling from 13.87 min to 1.51 min per CT volume during the model transfer, demonstrating the clinical usefulness with promising potentials.
Robotic Surgery Remote Mentoring via AR with 3D Scene Streaming and Hand Interaction
Apr 09, 2022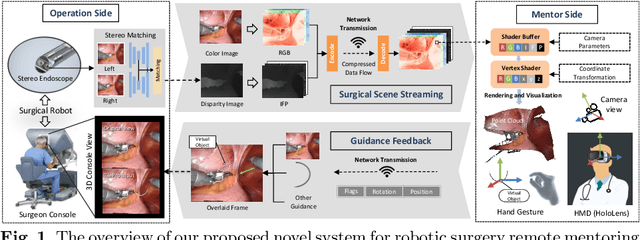
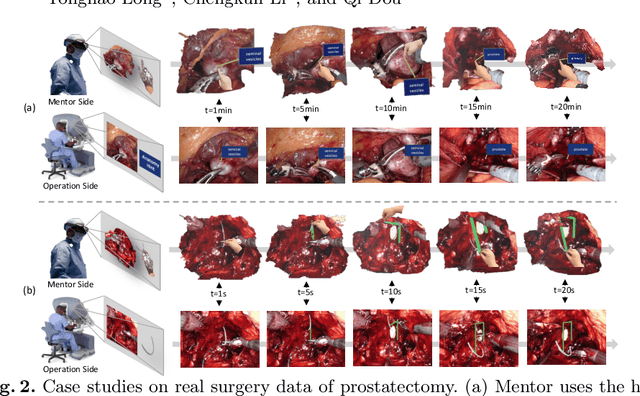

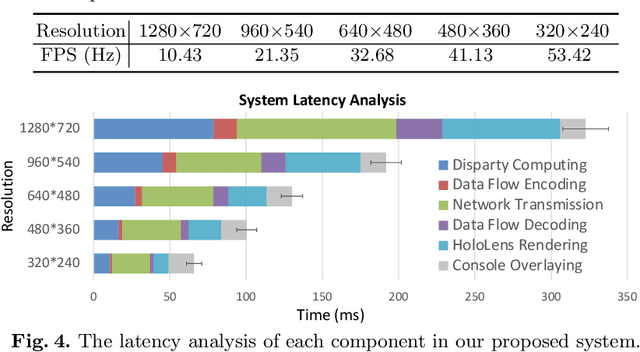
Abstract:With the growing popularity of robotic surgery, education becomes increasingly important and urgently needed for the sake of patient safety. However, experienced surgeons have limited accessibility due to their busy clinical schedule or working in a distant city, thus can hardly provide sufficient education resources for novices. Remote mentoring, as an effective way, can help solve this problem, but traditional methods are limited to plain text, audio, or 2D video, which are not intuitive nor vivid. Augmented reality (AR), a thriving technique being widely used for various education scenarios, is promising to offer new possibilities of visual experience and interactive teaching. In this paper, we propose a novel AR-based robotic surgery remote mentoring system with efficient 3D scene visualization and natural 3D hand interaction. Using a head-mounted display (i.e., HoloLens), the mentor can remotely monitor the procedure streamed from the trainee's operation side. The mentor can also provide feedback directly with hand gestures, which is in-turn transmitted to the trainee and viewed in surgical console as guidance. We comprehensively validate the system on both real surgery stereo videos and ex-vivo scenarios of common robotic training tasks (i.e., peg-transfer and suturing). Promising results are demonstrated regarding the fidelity of streamed scene visualization, the accuracy of feedback with hand interaction, and the low-latency of each component in the entire remote mentoring system. This work showcases the feasibility of leveraging AR technology for reliable, flexible and low-cost solutions to robotic surgical education, and holds great potential for clinical applications.
Federated Learning from Only Unlabeled Data with Class-Conditional-Sharing Clients
Apr 07, 2022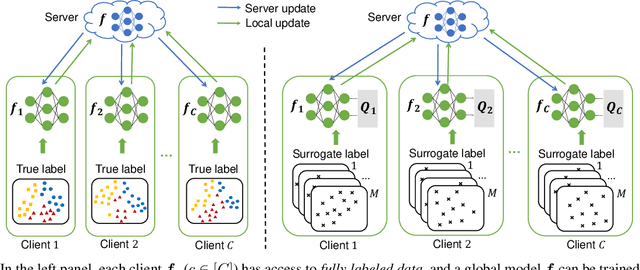
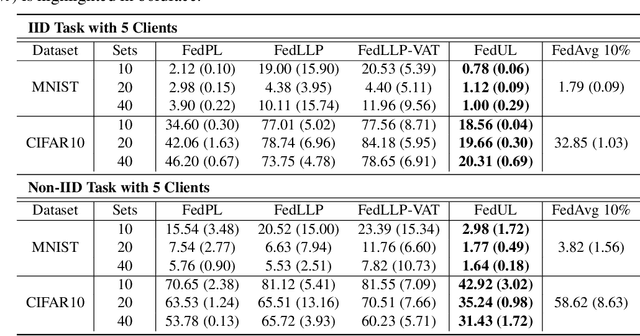
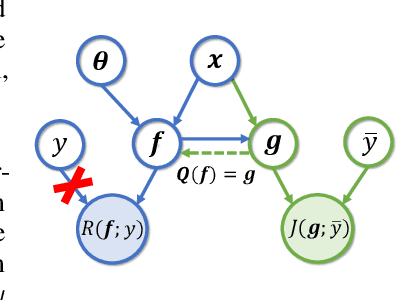
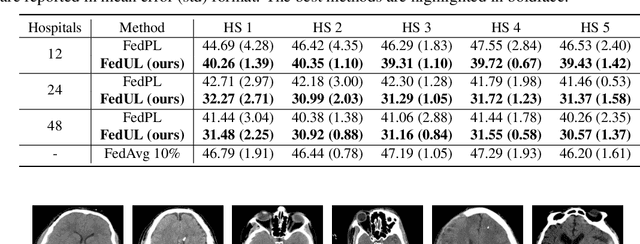
Abstract:Supervised federated learning (FL) enables multiple clients to share the trained model without sharing their labeled data. However, potential clients might even be reluctant to label their own data, which could limit the applicability of FL in practice. In this paper, we show the possibility of unsupervised FL whose model is still a classifier for predicting class labels, if the class-prior probabilities are shifted while the class-conditional distributions are shared among the unlabeled data owned by the clients. We propose federation of unsupervised learning (FedUL), where the unlabeled data are transformed into surrogate labeled data for each of the clients, a modified model is trained by supervised FL, and the wanted model is recovered from the modified model. FedUL is a very general solution to unsupervised FL: it is compatible with many supervised FL methods, and the recovery of the wanted model can be theoretically guaranteed as if the data have been labeled. Experiments on benchmark and real-world datasets demonstrate the effectiveness of FedUL. Code is available at https://github.com/lunanbit/FedUL.
3D Perception based Imitation Learning under Limited Demonstration for Laparoscope Control in Robotic Surgery
Apr 07, 2022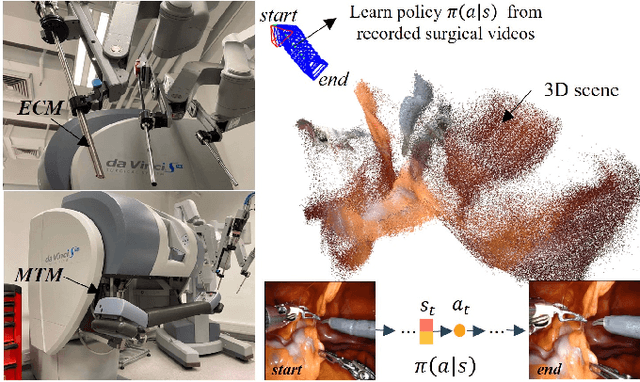
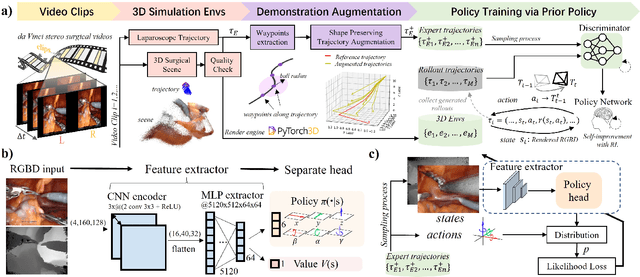
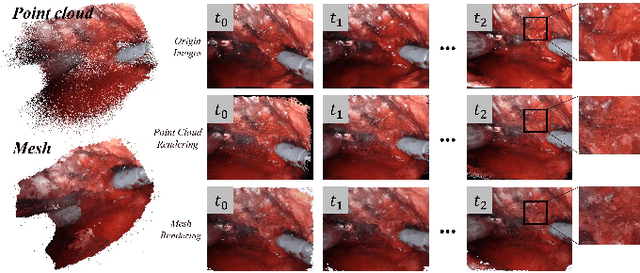
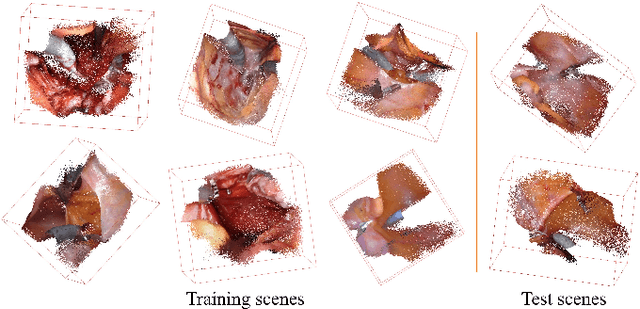
Abstract:Automatic laparoscope motion control is fundamentally important for surgeons to efficiently perform operations. However, its traditional control methods based on tool tracking without considering information hidden in surgical scenes are not intelligent enough, while the latest supervised imitation learning (IL)-based methods require expensive sensor data and suffer from distribution mismatch issues caused by limited demonstrations. In this paper, we propose a novel Imitation Learning framework for Laparoscope Control (ILLC) with reinforcement learning (RL), which can efficiently learn the control policy from limited surgical video clips. Specially, we first extract surgical laparoscope trajectories from unlabeled videos as the demonstrations and reconstruct the corresponding surgical scenes. To fully learn from limited motion trajectory demonstrations, we propose Shape Preserving Trajectory Augmentation (SPTA) to augment these data, and build a simulation environment that supports parallel RGB-D rendering to reinforce the RL policy for interacting with the environment efficiently. With adversarial training for IL, we obtain the laparoscope control policy based on the generated rollouts and surgical demonstrations. Extensive experiments are conducted in unseen reconstructed surgical scenes, and our method outperforms the previous IL methods, which proves the feasibility of our unified learning-based framework for laparoscope control.
Transformer-empowered Multi-scale Contextual Matching and Aggregation for Multi-contrast MRI Super-resolution
Mar 26, 2022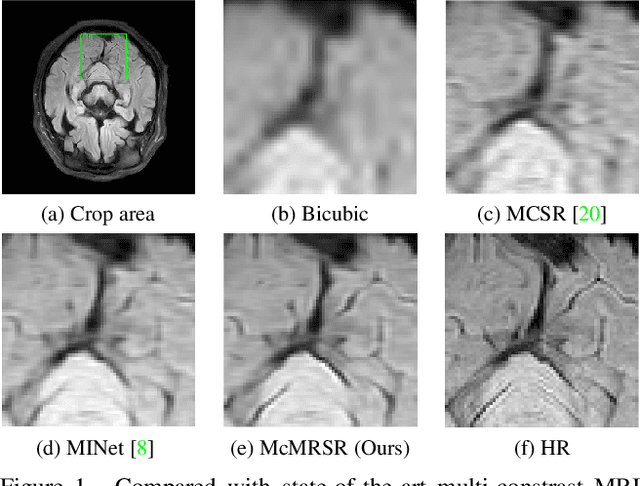
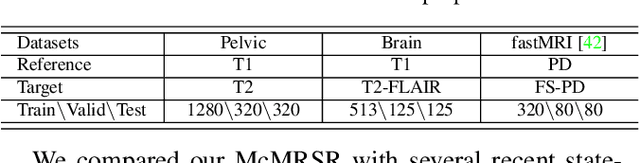
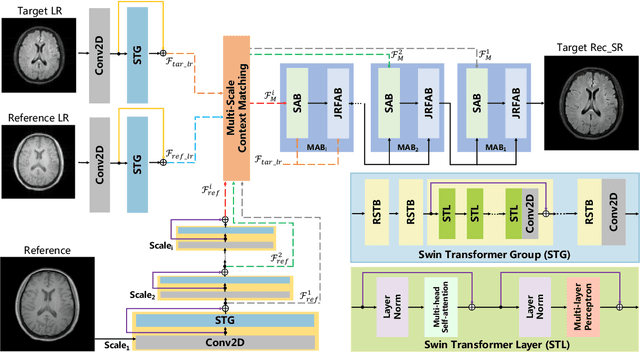
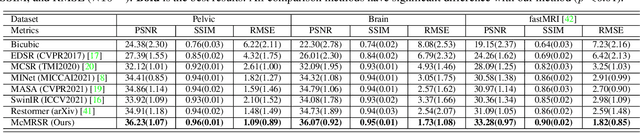
Abstract:Magnetic resonance imaging (MRI) can present multi-contrast images of the same anatomical structures, enabling multi-contrast super-resolution (SR) techniques. Compared with SR reconstruction using a single-contrast, multi-contrast SR reconstruction is promising to yield SR images with higher quality by leveraging diverse yet complementary information embedded in different imaging modalities. However, existing methods still have two shortcomings: (1) they neglect that the multi-contrast features at different scales contain different anatomical details and hence lack effective mechanisms to match and fuse these features for better reconstruction; and (2) they are still deficient in capturing long-range dependencies, which are essential for the regions with complicated anatomical structures. We propose a novel network to comprehensively address these problems by developing a set of innovative Transformer-empowered multi-scale contextual matching and aggregation techniques; we call it McMRSR. Firstly, we tame transformers to model long-range dependencies in both reference and target images. Then, a new multi-scale contextual matching method is proposed to capture corresponding contexts from reference features at different scales. Furthermore, we introduce a multi-scale aggregation mechanism to gradually and interactively aggregate multi-scale matched features for reconstructing the target SR MR image. Extensive experiments demonstrate that our network outperforms state-of-the-art approaches and has great potential to be applied in clinical practice. Codes are available at https://github.com/XAIMI-Lab/McMRSR.
Towards Robust Part-aware Instance Segmentation for Industrial Bin Picking
Mar 05, 2022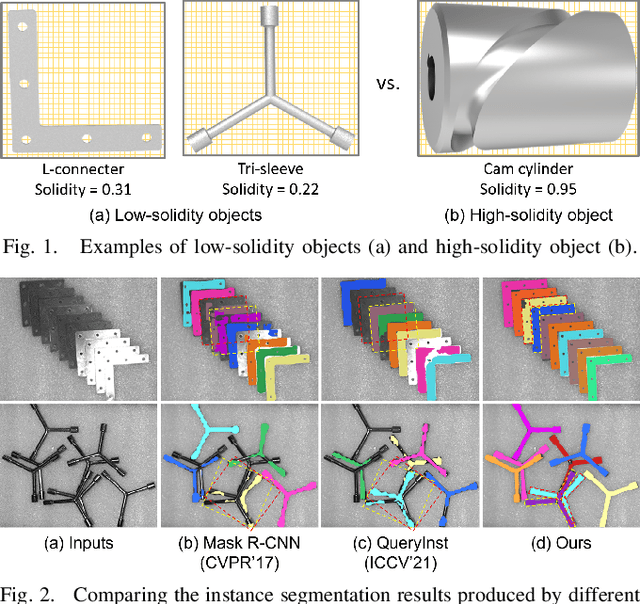
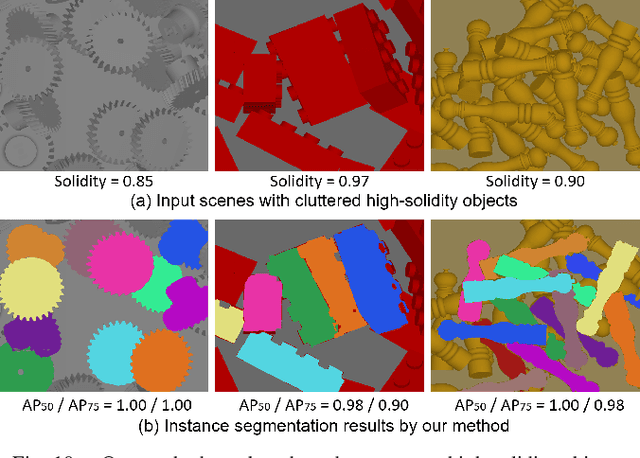

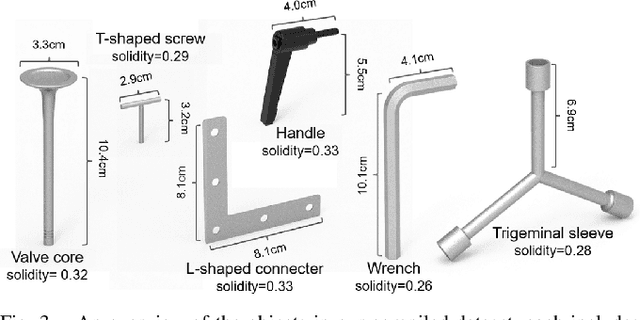
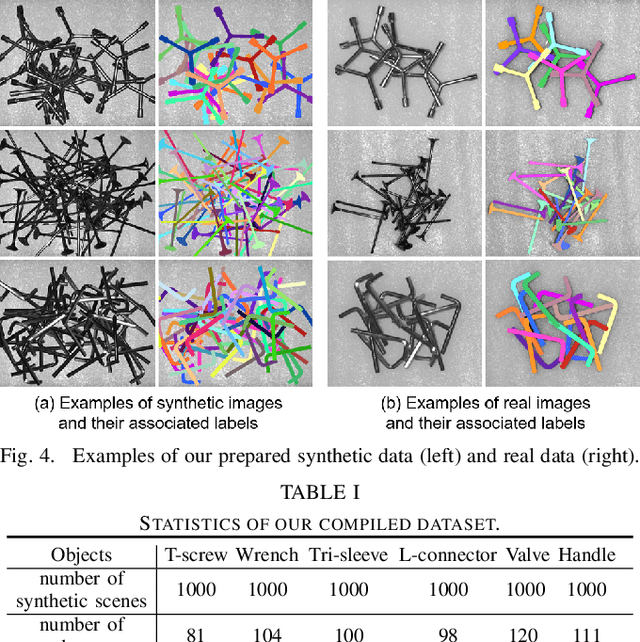
Abstract:Industrial bin picking is a challenging task that requires accurate and robust segmentation of individual object instances. Particularly, industrial objects can have irregular shapes, that is, thin and concave, whereas in bin-picking scenarios, objects are often closely packed with strong occlusion. To address these challenges, we formulate a novel part-aware instance segmentation pipeline. The key idea is to decompose industrial objects into correlated approximate convex parts and enhance the object-level segmentation with part-level segmentation. We design a part-aware network to predict part masks and part-to-part offsets, followed by a part aggregation module to assemble the recognized parts into instances. To guide the network learning, we also propose an automatic label decoupling scheme to generate ground-truth part-level labels from instance-level labels. Finally, we contribute the first instance segmentation dataset, which contains a variety of industrial objects that are thin and have non-trivial shapes. Extensive experimental results on various industrial objects demonstrate that our method can achieve the best segmentation results compared with the state-of-the-art approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge