Liansheng Wang
Generalizable Whole Slide Image Classification with Fine-Grained Visual-Semantic Interaction
Feb 29, 2024



Abstract:Whole Slide Image (WSI) classification is often formulated as a Multiple Instance Learning (MIL) problem. Recently, Vision-Language Models (VLMs) have demonstrated remarkable performance in WSI classification. However, existing methods leverage coarse-grained pathogenetic descriptions for visual representation supervision, which are insufficient to capture the complex visual appearance of pathogenetic images, hindering the generalizability of models on diverse downstream tasks. Additionally, processing high-resolution WSIs can be computationally expensive. In this paper, we propose a novel "Fine-grained Visual-Semantic Interaction" (FiVE) framework for WSI classification. It is designed to enhance the model's generalizability by leveraging the interplay between localized visual patterns and fine-grained pathological semantics. Specifically, with meticulously designed queries, we start by utilizing a large language model to extract fine-grained pathological descriptions from various non-standardized raw reports. The output descriptions are then reconstructed into fine-grained labels used for training. By introducing a Task-specific Fine-grained Semantics (TFS) module, we enable prompts to capture crucial visual information in WSIs, which enhances representation learning and augments generalization capabilities significantly. Furthermore, given that pathological visual patterns are redundantly distributed across tissue slices, we sample a subset of visual instances during training. Our method demonstrates robust generalizability and strong transferability, dominantly outperforming the counterparts on the TCGA Lung Cancer dataset with at least 9.19% higher accuracy in few-shot experiments.
Less Could Be Better: Parameter-efficient Fine-tuning Advances Medical Vision Foundation Models
Jan 22, 2024Abstract:Parameter-efficient fine-tuning (PEFT) that was initially developed for exploiting pre-trained large language models has recently emerged as an effective approach to perform transfer learning on computer vision tasks. However, the effectiveness of PEFT on medical vision foundation models is still unclear and remains to be explored. As a proof of concept, we conducted a detailed empirical study on applying PEFT to chest radiography foundation models. Specifically, we delved into LoRA, a representative PEFT method, and compared it against full-parameter fine-tuning (FFT) on two self-supervised radiography foundation models across three well-established chest radiograph datasets. Our results showed that LoRA outperformed FFT in 13 out of 18 transfer learning tasks by at most 2.9% using fewer than 1% tunable parameters. Combining LoRA with foundation models, we set up new state-of-the-art on a range of data-efficient learning tasks, such as an AUROC score of 80.6% using 1% labeled data on NIH ChestX-ray14. We hope this study can evoke more attention from the community in the use of PEFT for transfer learning on medical imaging tasks. Code and models are available at https://github.com/RL4M/MED-PEFT.
Simultaneous Alignment and Surface Regression Using Hybrid 2D-3D Networks for 3D Coherent Layer Segmentation of Retinal OCT Images with Full and Sparse Annotations
Dec 04, 2023



Abstract:Layer segmentation is important to quantitative analysis of retinal optical coherence tomography (OCT). Recently, deep learning based methods have been developed to automate this task and yield remarkable performance. However, due to the large spatial gap and potential mismatch between the B-scans of an OCT volume, all of them were based on 2D segmentation of individual B-scans, which may lose the continuity and diagnostic information of the retinal layers in 3D space. Besides, most of these methods required dense annotation of the OCT volumes, which is labor-intensive and expertise-demanding. This work presents a novel framework based on hybrid 2D-3D convolutional neural networks (CNNs) to obtain continuous 3D retinal layer surfaces from OCT volumes, which works well with both full and sparse annotations. The 2D features of individual B-scans are extracted by an encoder consisting of 2D convolutions. These 2D features are then used to produce the alignment displacement vectors and layer segmentation by two 3D decoders coupled via a spatial transformer module. Two losses are proposed to utilize the retinal layers' natural property of being smooth for B-scan alignment and layer segmentation, respectively, and are the key to the semi-supervised learning with sparse annotation. The entire framework is trained end-to-end. To the best of our knowledge, this is the first work that attempts 3D retinal layer segmentation in volumetric OCT images based on CNNs. Experiments on a synthetic dataset and three public clinical datasets show that our framework can effectively align the B-scans for potential motion correction, and achieves superior performance to state-of-the-art 2D deep learning methods in terms of both layer segmentation accuracy and cross-B-scan 3D continuity in both fully and semi-supervised settings, thus offering more clinical values than previous works.
Shifting More Attention to Breast Lesion Segmentation in Ultrasound Videos
Oct 03, 2023Abstract:Breast lesion segmentation in ultrasound (US) videos is essential for diagnosing and treating axillary lymph node metastasis. However, the lack of a well-established and large-scale ultrasound video dataset with high-quality annotations has posed a persistent challenge for the research community. To overcome this issue, we meticulously curated a US video breast lesion segmentation dataset comprising 572 videos and 34,300 annotated frames, covering a wide range of realistic clinical scenarios. Furthermore, we propose a novel frequency and localization feature aggregation network (FLA-Net) that learns temporal features from the frequency domain and predicts additional lesion location positions to assist with breast lesion segmentation. We also devise a localization-based contrastive loss to reduce the lesion location distance between neighboring video frames within the same video and enlarge the location distances between frames from different ultrasound videos. Our experiments on our annotated dataset and two public video polyp segmentation datasets demonstrate that our proposed FLA-Net achieves state-of-the-art performance in breast lesion segmentation in US videos and video polyp segmentation while significantly reducing time and space complexity. Our model and dataset are available at https://github.com/jhl-Det/FLA-Net.
You've Got Two Teachers: Co-evolutionary Image and Report Distillation for Semi-supervised Anatomical Abnormality Detection in Chest X-ray
Jul 18, 2023Abstract:Chest X-ray (CXR) anatomical abnormality detection aims at localizing and characterising cardiopulmonary radiological findings in the radiographs, which can expedite clinical workflow and reduce observational oversights. Most existing methods attempted this task in either fully supervised settings which demanded costly mass per-abnormality annotations, or weakly supervised settings which still lagged badly behind fully supervised methods in performance. In this work, we propose a co-evolutionary image and report distillation (CEIRD) framework, which approaches semi-supervised abnormality detection in CXR by grounding the visual detection results with text-classified abnormalities from paired radiology reports, and vice versa. Concretely, based on the classical teacher-student pseudo label distillation (TSD) paradigm, we additionally introduce an auxiliary report classification model, whose prediction is used for report-guided pseudo detection label refinement (RPDLR) in the primary vision detection task. Inversely, we also use the prediction of the vision detection model for abnormality-guided pseudo classification label refinement (APCLR) in the auxiliary report classification task, and propose a co-evolution strategy where the vision and report models mutually promote each other with RPDLR and APCLR performed alternatively. To this end, we effectively incorporate the weak supervision by reports into the semi-supervised TSD pipeline. Besides the cross-modal pseudo label refinement, we further propose an intra-image-modal self-adaptive non-maximum suppression, where the pseudo detection labels generated by the teacher vision model are dynamically rectified by high-confidence predictions by the student. Experimental results on the public MIMIC-CXR benchmark demonstrate CEIRD's superior performance to several up-to-date weakly and semi-supervised methods.
Why is the winner the best?
Mar 30, 2023



Abstract:International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
M3AE: Multimodal Representation Learning for Brain Tumor Segmentation with Missing Modalities
Mar 09, 2023



Abstract:Multimodal magnetic resonance imaging (MRI) provides complementary information for sub-region analysis of brain tumors. Plenty of methods have been proposed for automatic brain tumor segmentation using four common MRI modalities and achieved remarkable performance. In practice, however, it is common to have one or more modalities missing due to image corruption, artifacts, acquisition protocols, allergy to contrast agents, or simply cost. In this work, we propose a novel two-stage framework for brain tumor segmentation with missing modalities. In the first stage, a multimodal masked autoencoder (M3AE) is proposed, where both random modalities (i.e., modality dropout) and random patches of the remaining modalities are masked for a reconstruction task, for self-supervised learning of robust multimodal representations against missing modalities. To this end, we name our framework M3AE. Meanwhile, we employ model inversion to optimize a representative full-modal image at marginal extra cost, which will be used to substitute for the missing modalities and boost performance during inference. Then in the second stage, a memory-efficient self distillation is proposed to distill knowledge between heterogenous missing-modal situations while fine-tuning the model for supervised segmentation. Our M3AE belongs to the 'catch-all' genre where a single model can be applied to all possible subsets of modalities, thus is economic for both training and deployment. Extensive experiments on BraTS 2018 and 2020 datasets demonstrate its superior performance to existing state-of-the-art methods with missing modalities, as well as the efficacy of its components. Our code is available at: https://github.com/ccarliu/m3ae.
RECIST Weakly Supervised Lesion Segmentation via Label-Space Co-Training
Mar 01, 2023



Abstract:As an essential indicator for cancer progression and treatment response, tumor size is often measured following the response evaluation criteria in solid tumors (RECIST) guideline in CT slices. By marking each lesion with its longest axis and the longest perpendicular one, laborious pixel-wise manual annotation can be avoided. However, such a coarse substitute cannot provide a rich and accurate base to allow versatile quantitative analysis of lesions. To this end, we propose a novel weakly supervised framework to exploit the existing rich RECIST annotations for pixel-wise lesion segmentation. Specifically, a pair of under- and over-segmenting masks are constructed for each lesion based on its RECIST annotation and served as the label for co-training a pair of subnets, respectively, along with the proposed label-space perturbation induced consistency loss to bridge the gap between the two subnets and enable effective co-training. Extensive experiments are conducted on a public dataset to demonstrate the superiority of the proposed framework regarding the RECIST-based weakly supervised segmentation task and its universal applicability to various backbone networks.
Advancing Radiograph Representation Learning with Masked Record Modeling
Feb 15, 2023



Abstract:Modern studies in radiograph representation learning rely on either self-supervision to encode invariant semantics or associated radiology reports to incorporate medical expertise, while the complementarity between them is barely noticed. To explore this, we formulate the self- and report-completion as two complementary objectives and present a unified framework based on masked record modeling (MRM). In practice, MRM reconstructs masked image patches and masked report tokens following a multi-task scheme to learn knowledge-enhanced semantic representations. With MRM pre-training, we obtain pre-trained models that can be well transferred to various radiography tasks. Specifically, we find that MRM offers superior performance in label-efficient fine-tuning. For instance, MRM achieves 88.5% mean AUC on CheXpert using 1% labeled data, outperforming previous R$^2$L methods with 100% labels. On NIH ChestX-ray, MRM outperforms the best performing counterpart by about 3% under small labeling ratios. Besides, MRM surpasses self- and report-supervised pre-training in identifying the pneumonia type and the pneumothorax area, sometimes by large margins.
GraVIS: Grouping Augmented Views from Independent Sources for Dermatology Analysis
Jan 11, 2023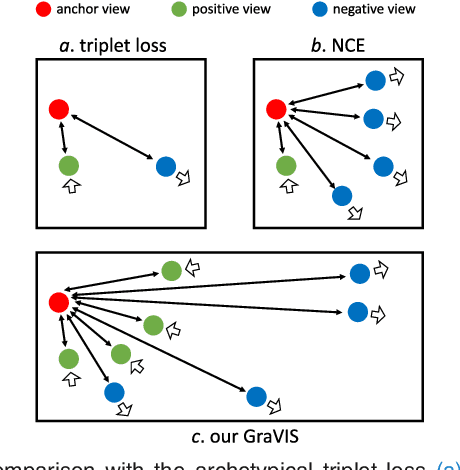
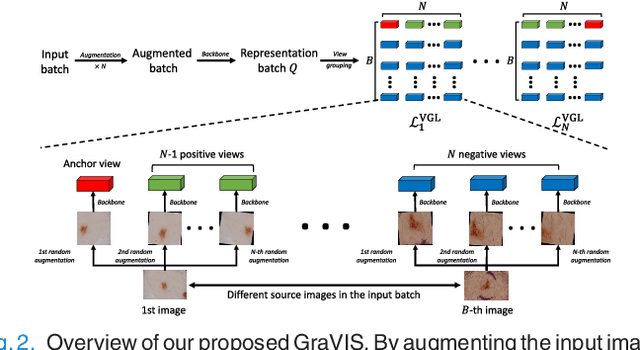
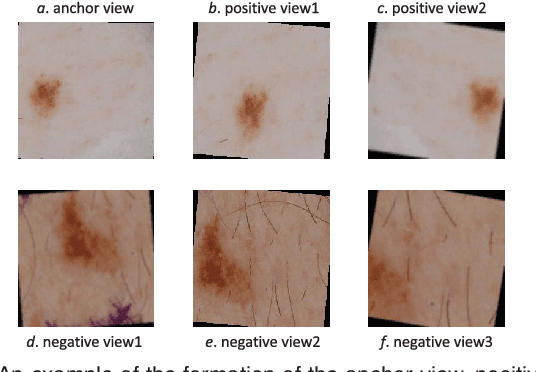
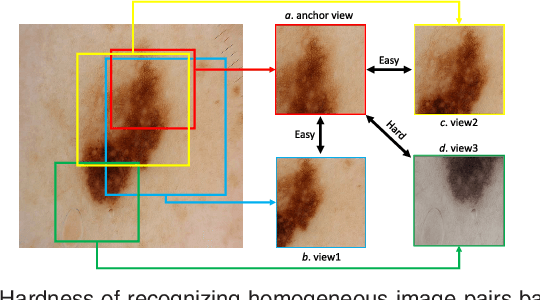
Abstract:Self-supervised representation learning has been extremely successful in medical image analysis, as it requires no human annotations to provide transferable representations for downstream tasks. Recent self-supervised learning methods are dominated by noise-contrastive estimation (NCE, also known as contrastive learning), which aims to learn invariant visual representations by contrasting one homogeneous image pair with a large number of heterogeneous image pairs in each training step. Nonetheless, NCE-based approaches still suffer from one major problem that is one homogeneous pair is not enough to extract robust and invariant semantic information. Inspired by the archetypical triplet loss, we propose GraVIS, which is specifically optimized for learning self-supervised features from dermatology images, to group homogeneous dermatology images while separating heterogeneous ones. In addition, a hardness-aware attention is introduced and incorporated to address the importance of homogeneous image views with similar appearance instead of those dissimilar homogeneous ones. GraVIS significantly outperforms its transfer learning and self-supervised learning counterparts in both lesion segmentation and disease classification tasks, sometimes by 5 percents under extremely limited supervision. More importantly, when equipped with the pre-trained weights provided by GraVIS, a single model could achieve better results than winners that heavily rely on ensemble strategies in the well-known ISIC 2017 challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge