Fengze Liu
InfoLaw: Information Scaling Laws for Large Language Models with Quality-Weighted Mixture Data and Repetition
May 04, 2026Abstract:Upweighting high-quality data in LLM pretraining often improves performance, but in datalimited regimes, especially under overtraining, stronger upweighting increases repetition and can degrade performance. However, standard scaling laws do not reliably extrapolate across mixture recipes or under repetitions, making the selection for optimal data recipes at scaling underdetermined. To solve this, we introduce InfoLaw (Information Scaling Laws), a data-aware scaling framework that predicts loss from consumed tokens, model size, data mixture weights, and repetition. The key idea is to model pretraining as information accumulation, where quality controls information density and repetition induces scaledependent diminishing returns. We first collect the model performance after training on datasets that vary in scale, quality distribution, and repetition level. Then we build up the modeling for information so that information accurately predicts those model performance. InfoLaw predicts performance on unseen data recipes and larger scale runs (up to 7B, 425B tokens) with 0.15% mean and 0.96% max absolute error in loss, and it extrapolates reliably across overtraining levels, enabling efficient data-recipe selection under varying compute budgets.
Your Agent, Their Asset: A Real-World Safety Analysis of OpenClaw
Apr 06, 2026Abstract:OpenClaw, the most widely deployed personal AI agent in early 2026, operates with full local system access and integrates with sensitive services such as Gmail, Stripe, and the filesystem. While these broad privileges enable high levels of automation and powerful personalization, they also expose a substantial attack surface that existing sandboxed evaluations fail to capture. To address this gap, we present the first real-world safety evaluation of OpenClaw and introduce the CIK taxonomy, which unifies an agent's persistent state into three dimensions, i.e., Capability, Identity, and Knowledge, for safety analysis. Our evaluations cover 12 attack scenarios on a live OpenClaw instance across four backbone models (Claude Sonnet 4.5, Opus 4.6, Gemini 3.1 Pro, and GPT-5.4). The results show that poisoning any single CIK dimension increases the average attack success rate from 24.6% to 64-74%, with even the most robust model exhibiting more than a threefold increase over its baseline vulnerability. We further assess three CIK-aligned defense strategies alongside a file-protection mechanism; however, the strongest defense still yields a 63.8% success rate under Capability-targeted attacks, while file protection blocks 97% of malicious injections but also prevents legitimate updates. Taken together, these findings show that the vulnerabilities are inherent to the agent architecture, necessitating more systematic safeguards to secure personal AI agents. Our project page is https://ucsc-vlaa.github.io/CIK-Bench.
Exploring Polyglot Harmony: On Multilingual Data Allocation for Large Language Models Pretraining
Sep 19, 2025Abstract:Large language models (LLMs) have become integral to a wide range of applications worldwide, driving an unprecedented global demand for effective multilingual capabilities. Central to achieving robust multilingual performance is the strategic allocation of language proportions within training corpora. However, determining optimal language ratios is highly challenging due to intricate cross-lingual interactions and sensitivity to dataset scale. This paper introduces Climb (Cross-Lingual Interaction-aware Multilingual Balancing), a novel framework designed to systematically optimize multilingual data allocation. At its core, Climb introduces a cross-lingual interaction-aware language ratio, explicitly quantifying each language's effective allocation by capturing inter-language dependencies. Leveraging this ratio, Climb proposes a principled two-step optimization procedure--first equalizing marginal benefits across languages, then maximizing the magnitude of the resulting language allocation vectors--significantly simplifying the inherently complex multilingual optimization problem. Extensive experiments confirm that Climb can accurately measure cross-lingual interactions across various multilingual settings. LLMs trained with Climb-derived proportions consistently achieve state-of-the-art multilingual performance, even achieving competitive performance with open-sourced LLMs trained with more tokens.
TiKMiX: Take Data Influence into Dynamic Mixture for Language Model Pre-training
Aug 25, 2025Abstract:The data mixture used in the pre-training of a language model is a cornerstone of its final performance. However, a static mixing strategy is suboptimal, as the model's learning preferences for various data domains shift dynamically throughout training. Crucially, observing these evolving preferences in a computationally efficient manner remains a significant challenge. To address this, we propose TiKMiX, a method that dynamically adjusts the data mixture according to the model's evolving preferences. TiKMiX introduces Group Influence, an efficient metric for evaluating the impact of data domains on the model. This metric enables the formulation of the data mixing problem as a search for an optimal, influence-maximizing distribution. We solve this via two approaches: TiKMiX-D for direct optimization, and TiKMiX-M, which uses a regression model to predict a superior mixture. We trained models with different numbers of parameters, on up to 1 trillion tokens. TiKMiX-D exceeds the performance of state-of-the-art methods like REGMIX while using just 20% of the computational resources. TiKMiX-M leads to an average performance gain of 2% across 9 downstream benchmarks. Our experiments reveal that a model's data preferences evolve with training progress and scale, and we demonstrate that dynamically adjusting the data mixture based on Group Influence, a direct measure of these preferences, significantly improves performance by mitigating the underdigestion of data seen with static ratios.
MuRating: A High Quality Data Selecting Approach to Multilingual Large Language Model Pretraining
Jul 02, 2025Abstract:Data quality is a critical driver of large language model performance, yet existing model-based selection methods focus almost exclusively on English. We introduce MuRating, a scalable framework that transfers high-quality English data-quality signals into a single rater for 17 target languages. MuRating aggregates multiple English "raters" via pairwise comparisons to learn unified document-quality scores,then projects these judgments through translation to train a multilingual evaluator on monolingual, cross-lingual, and parallel text pairs. Applied to web data, MuRating selects balanced subsets of English and multilingual content to pretrain a 1.2 B-parameter LLaMA model. Compared to strong baselines, including QuRater, AskLLM, DCLM and so on, our approach boosts average accuracy on both English benchmarks and multilingual evaluations, with especially large gains on knowledge-intensive tasks. We further analyze translation fidelity, selection biases, and underrepresentation of narrative material, outlining directions for future work.
AUTOCT: Automating Interpretable Clinical Trial Prediction with LLM Agents
Jun 04, 2025



Abstract:Clinical trials are critical for advancing medical treatments but remain prohibitively expensive and time-consuming. Accurate prediction of clinical trial outcomes can significantly reduce research and development costs and accelerate drug discovery. While recent deep learning models have shown promise by leveraging unstructured data, their black-box nature, lack of interpretability, and vulnerability to label leakage limit their practical use in high-stakes biomedical contexts. In this work, we propose AutoCT, a novel framework that combines the reasoning capabilities of large language models with the explainability of classical machine learning. AutoCT autonomously generates, evaluates, and refines tabular features based on public information without human input. Our method uses Monte Carlo Tree Search to iteratively optimize predictive performance. Experimental results show that AutoCT performs on par with or better than SOTA methods on clinical trial prediction tasks within only a limited number of self-refinement iterations, establishing a new paradigm for scalable, interpretable, and cost-efficient clinical trial prediction.
QuaDMix: Quality-Diversity Balanced Data Selection for Efficient LLM Pretraining
Apr 23, 2025



Abstract:Quality and diversity are two critical metrics for the training data of large language models (LLMs), positively impacting performance. Existing studies often optimize these metrics separately, typically by first applying quality filtering and then adjusting data proportions. However, these approaches overlook the inherent trade-off between quality and diversity, necessitating their joint consideration. Given a fixed training quota, it is essential to evaluate both the quality of each data point and its complementary effect on the overall dataset. In this paper, we introduce a unified data selection framework called QuaDMix, which automatically optimizes the data distribution for LLM pretraining while balancing both quality and diversity. Specifically, we first propose multiple criteria to measure data quality and employ domain classification to distinguish data points, thereby measuring overall diversity. QuaDMix then employs a unified parameterized data sampling function that determines the sampling probability of each data point based on these quality and diversity related labels. To accelerate the search for the optimal parameters involved in the QuaDMix framework, we conduct simulated experiments on smaller models and use LightGBM for parameters searching, inspired by the RegMix method. Our experiments across diverse models and datasets demonstrate that QuaDMix achieves an average performance improvement of 7.2% across multiple benchmarks. These results outperform the independent strategies for quality and diversity, highlighting the necessity and ability to balance data quality and diversity.
SAME++: A Self-supervised Anatomical eMbeddings Enhanced medical image registration framework using stable sampling and regularized transformation
Nov 25, 2023Abstract:Image registration is a fundamental medical image analysis task. Ideally, registration should focus on aligning semantically corresponding voxels, i.e., the same anatomical locations. However, existing methods often optimize similarity measures computed directly on intensities or on hand-crafted features, which lack anatomical semantic information. These similarity measures may lead to sub-optimal solutions where large deformations, complex anatomical differences, or cross-modality imagery exist. In this work, we introduce a fast and accurate method for unsupervised 3D medical image registration building on top of a Self-supervised Anatomical eMbedding (SAM) algorithm, which is capable of computing dense anatomical correspondences between two images at the voxel level. We name our approach SAM-Enhanced registration (SAME++), which decomposes image registration into four steps: affine transformation, coarse deformation, deep non-parametric transformation, and instance optimization. Using SAM embeddings, we enhance these steps by finding more coherent correspondence and providing features with better semantic guidance. We extensively evaluated SAME++ using more than 50 labeled organs on three challenging inter-subject registration tasks of different body parts. As a complete registration framework, SAME++ markedly outperforms leading methods by $4.2\%$ - $8.2\%$ in terms of Dice score while being orders of magnitude faster than numerical optimization-based methods. Code is available at \url{https://github.com/alibaba-damo-academy/same}.
Unsupervised Domain Adaptation through Shape Modeling for Medical Image Segmentation
Jul 06, 2022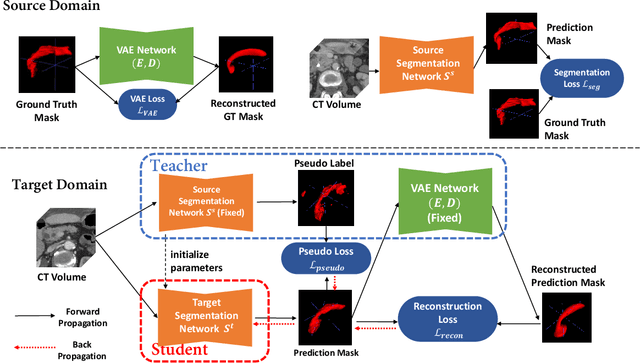
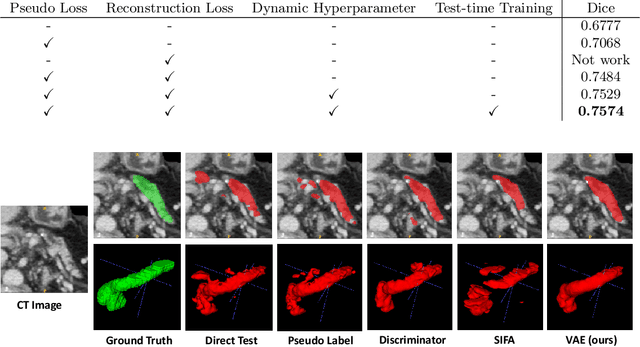
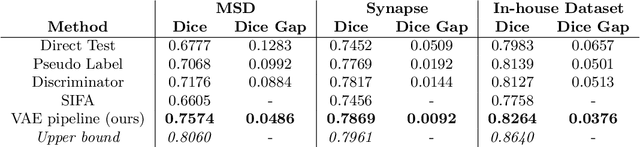
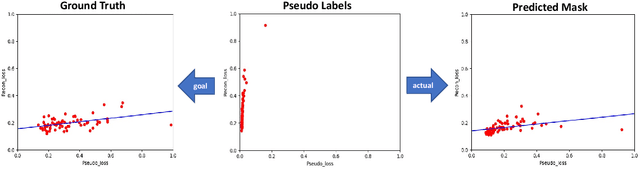
Abstract:Shape information is a strong and valuable prior in segmenting organs in medical images. However, most current deep learning based segmentation algorithms have not taken shape information into consideration, which can lead to bias towards texture. We aim at modeling shape explicitly and using it to help medical image segmentation. Previous methods proposed Variational Autoencoder (VAE) based models to learn the distribution of shape for a particular organ and used it to automatically evaluate the quality of a segmentation prediction by fitting it into the learned shape distribution. Based on which we aim at incorporating VAE into current segmentation pipelines. Specifically, we propose a new unsupervised domain adaptation pipeline based on a pseudo loss and a VAE reconstruction loss under a teacher-student learning paradigm. Both losses are optimized simultaneously and, in return, boost the segmentation task performance. Extensive experiments on three public Pancreas segmentation datasets as well as two in-house Pancreas segmentation datasets show consistent improvements with at least 2.8 points gain in the Dice score, demonstrating the effectiveness of our method in challenging unsupervised domain adaptation scenarios for medical image segmentation. We hope this work will advance shape analysis and geometric learning in medical imaging.
Learning to Bootstrap for Combating Label Noise
Feb 09, 2022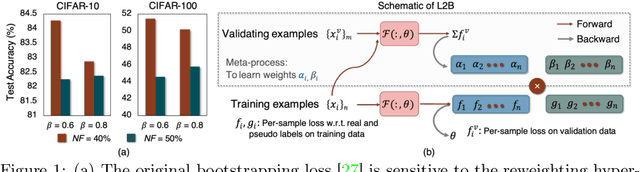
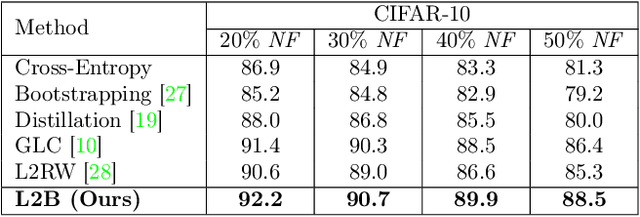
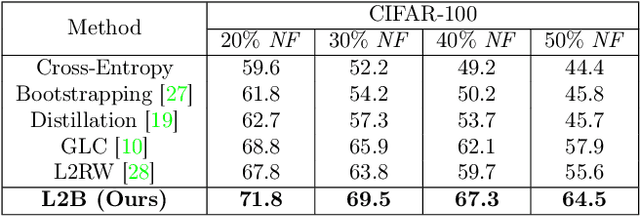
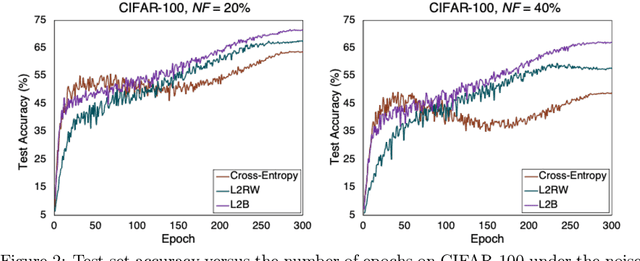
Abstract:Deep neural networks are powerful tools for representation learning, but can easily overfit to noisy labels which are prevalent in many real-world scenarios. Generally, noisy supervision could stem from variation among labelers, label corruption by adversaries, etc. To combat such label noises, one popular line of approach is to apply customized weights to the training instances, so that the corrupted examples contribute less to the model learning. However, such learning mechanisms potentially erase important information about the data distribution and therefore yield suboptimal results. To leverage useful information from the corrupted instances, an alternative is the bootstrapping loss, which reconstructs new training targets on-the-fly by incorporating the network's own predictions (i.e., pseudo-labels). In this paper, we propose a more generic learnable loss objective which enables a joint reweighting of instances and labels at once. Specifically, our method dynamically adjusts the per-sample importance weight between the real observed labels and pseudo-labels, where the weights are efficiently determined in a meta process. Compared to the previous instance reweighting methods, our approach concurrently conducts implicit relabeling, and thereby yield substantial improvements with almost no extra cost. Extensive experimental results demonstrated the strengths of our approach over existing methods on multiple natural and medical image benchmark datasets, including CIFAR-10, CIFAR-100, ISIC2019 and Clothing 1M. The code is publicly available at https://github.com/yuyinzhou/L2B.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge