Zhen Guo
Department of Electrical Engineering and Computer Science, Massachusetts Institute of Technology, Cambridge, Massachusetts, 02139, USA
RLinf-USER: A Unified and Extensible System for Real-World Online Policy Learning in Embodied AI
Feb 08, 2026Abstract:Online policy learning directly in the physical world is a promising yet challenging direction for embodied intelligence. Unlike simulation, real-world systems cannot be arbitrarily accelerated, cheaply reset, or massively replicated, which makes scalable data collection, heterogeneous deployment, and long-horizon effective training difficult. These challenges suggest that real-world policy learning is not only an algorithmic issue but fundamentally a systems problem. We present USER, a Unified and extensible SystEm for Real-world online policy learning. USER treats physical robots as first-class hardware resources alongside GPUs through a unified hardware abstraction layer, enabling automatic discovery, management, and scheduling of heterogeneous robots. To address cloud-edge communication, USER introduces an adaptive communication plane with tunneling-based networking, distributed data channels for traffic localization, and streaming-multiprocessor-aware weight synchronization to regulate GPU-side overhead. On top of this infrastructure, USER organizes learning as a fully asynchronous framework with a persistent, cache-aware buffer, enabling efficient long-horizon experiments with robust crash recovery and reuse of historical data. In addition, USER provides extensible abstractions for rewards, algorithms, and policies, supporting online imitation or reinforcement learning of CNN/MLP, generative policies, and large vision-language-action (VLA) models within a unified pipeline. Results in both simulation and the real world show that USER enables multi-robot coordination, heterogeneous manipulators, edge-cloud collaboration with large models, and long-running asynchronous training, offering a unified and extensible systems foundation for real-world online policy learning.
CL-bench: A Benchmark for Context Learning
Feb 03, 2026Abstract:Current language models (LMs) excel at reasoning over prompts using pre-trained knowledge. However, real-world tasks are far more complex and context-dependent: models must learn from task-specific context and leverage new knowledge beyond what is learned during pre-training to reason and resolve tasks. We term this capability context learning, a crucial ability that humans naturally possess but has been largely overlooked. To this end, we introduce CL-bench, a real-world benchmark consisting of 500 complex contexts, 1,899 tasks, and 31,607 verification rubrics, all crafted by experienced domain experts. Each task is designed such that the new content required to resolve it is contained within the corresponding context. Resolving tasks in CL-bench requires models to learn from the context, ranging from new domain-specific knowledge, rule systems, and complex procedures to laws derived from empirical data, all of which are absent from pre-training. This goes far beyond long-context tasks that primarily test retrieval or reading comprehension, and in-context learning tasks, where models learn simple task patterns via instructions and demonstrations. Our evaluations of ten frontier LMs find that models solve only 17.2% of tasks on average. Even the best-performing model, GPT-5.1, solves only 23.7%, revealing that LMs have yet to achieve effective context learning, which poses a critical bottleneck for tackling real-world, complex context-dependent tasks. CL-bench represents a step towards building LMs with this fundamental capability, making them more intelligent and advancing their deployment in real-world scenarios.
MICo-150K: A Comprehensive Dataset Advancing Multi-Image Composition
Dec 08, 2025



Abstract:In controllable image generation, synthesizing coherent and consistent images from multiple reference inputs, i.e., Multi-Image Composition (MICo), remains a challenging problem, partly hindered by the lack of high-quality training data. To bridge this gap, we conduct a systematic study of MICo, categorizing it into 7 representative tasks and curate a large-scale collection of high-quality source images and construct diverse MICo prompts. Leveraging powerful proprietary models, we synthesize a rich amount of balanced composite images, followed by human-in-the-loop filtering and refinement, resulting in MICo-150K, a comprehensive dataset for MICo with identity consistency. We further build a Decomposition-and-Recomposition (De&Re) subset, where 11K real-world complex images are decomposed into components and recomposed, enabling both real and synthetic compositions. To enable comprehensive evaluation, we construct MICo-Bench with 100 cases per task and 300 challenging De&Re cases, and further introduce a new metric, Weighted-Ref-VIEScore, specifically tailored for MICo evaluation. Finally, we fine-tune multiple models on MICo-150K and evaluate them on MICo-Bench. The results show that MICo-150K effectively equips models without MICo capability and further enhances those with existing skills. Notably, our baseline model, Qwen-MICo, fine-tuned from Qwen-Image-Edit, matches Qwen-Image-2509 in 3-image composition while supporting arbitrary multi-image inputs beyond the latter's limitation. Our dataset, benchmark, and baseline collectively offer valuable resources for further research on Multi-Image Composition.
$π_\texttt{RL}$: Online RL Fine-tuning for Flow-based Vision-Language-Action Models
Oct 29, 2025


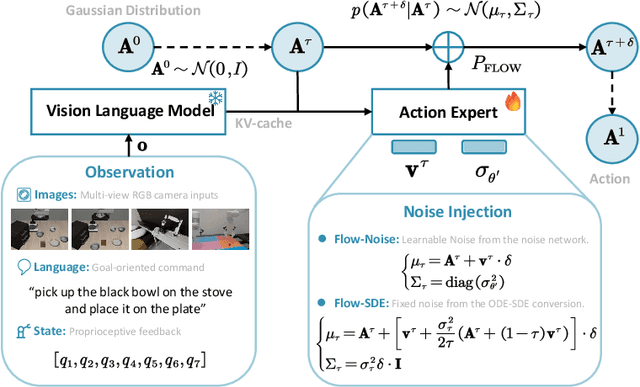
Abstract:Vision-Language-Action (VLA) models enable robots to understand and perform complex tasks from multimodal input. Although recent work explores using reinforcement learning (RL) to automate the laborious data collection process in scaling supervised fine-tuning (SFT), applying large-scale RL to flow-based VLAs (e.g., $\pi_0$, $\pi_{0.5}$) remains challenging due to intractable action log-likelihoods from iterative denoising. We address this challenge with $\pi_{\text{RL}}$, an open-source framework for training flow-based VLAs in parallel simulation. $\pi_{\text{RL}}$ implements two RL algorithms: (1) {Flow-Noise} models the denoising process as a discrete-time MDP with a learnable noise network for exact log-likelihood computation. (2) {Flow-SDE} integrates denoising with agent-environment interaction, formulating a two-layer MDP that employs ODE-to-SDE conversion for efficient RL exploration. We evaluate $\pi_{\text{RL}}$ on LIBERO and ManiSkill benchmarks. On LIBERO, $\pi_{\text{RL}}$ boosts few-shot SFT models $\pi_0$ and $\pi_{0.5}$ from 57.6% to 97.6% and from 77.1% to 98.3%, respectively. In ManiSkill, we train $\pi_{\text{RL}}$ in 320 parallel environments, improving $\pi_0$ from 41.6% to 85.7% and $\pi_{0.5}$ from 40.0% to 84.8% across 4352 pick-and-place tasks, demonstrating scalable multitask RL under heterogeneous simulation. Overall, $\pi_{\text{RL}}$ achieves significant performance gains and stronger generalization over SFT-models, validating the effectiveness of online RL for flow-based VLAs.
RLinf: Flexible and Efficient Large-scale Reinforcement Learning via Macro-to-Micro Flow Transformation
Sep 19, 2025
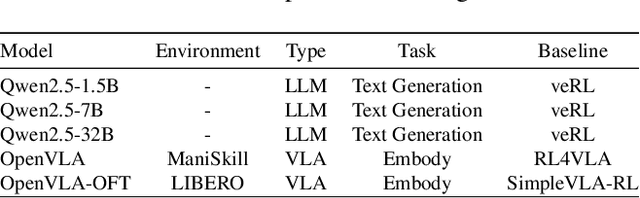

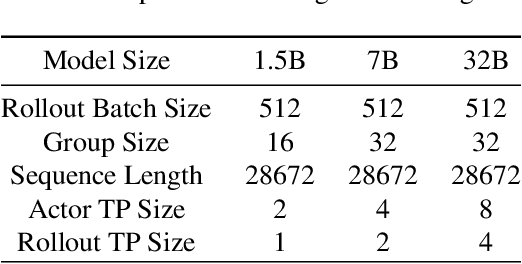
Abstract:Reinforcement learning (RL) has demonstrated immense potential in advancing artificial general intelligence, agentic intelligence, and embodied intelligence. However, the inherent heterogeneity and dynamicity of RL workflows often lead to low hardware utilization and slow training on existing systems. In this paper, we present RLinf, a high-performance RL training system based on our key observation that the major roadblock to efficient RL training lies in system flexibility. To maximize flexibility and efficiency, RLinf is built atop a novel RL system design paradigm called macro-to-micro flow transformation (M2Flow), which automatically breaks down high-level, easy-to-compose RL workflows at both the temporal and spatial dimensions, and recomposes them into optimized execution flows. Supported by RLinf worker's adaptive communication capability, we devise context switching and elastic pipelining to realize M2Flow transformation, and a profiling-guided scheduling policy to generate optimal execution plans. Extensive evaluations on both reasoning RL and embodied RL tasks demonstrate that RLinf consistently outperforms state-of-the-art systems, achieving 1.1x-2.13x speedup in end-to-end training throughput.
Persistent Backdoor Attacks in Continual Learning
Sep 20, 2024



Abstract:Backdoor attacks pose a significant threat to neural networks, enabling adversaries to manipulate model outputs on specific inputs, often with devastating consequences, especially in critical applications. While backdoor attacks have been studied in various contexts, little attention has been given to their practicality and persistence in continual learning, particularly in understanding how the continual updates to model parameters, as new data distributions are learned and integrated, impact the effectiveness of these attacks over time. To address this gap, we introduce two persistent backdoor attacks-Blind Task Backdoor and Latent Task Backdoor-each leveraging minimal adversarial influence. Our blind task backdoor subtly alters the loss computation without direct control over the training process, while the latent task backdoor influences only a single task's training, with all other tasks trained benignly. We evaluate these attacks under various configurations, demonstrating their efficacy with static, dynamic, physical, and semantic triggers. Our results show that both attacks consistently achieve high success rates across different continual learning algorithms, while effectively evading state-of-the-art defenses, such as SentiNet and I-BAU.
Scaling Law Hypothesis for Multimodal Model
Sep 10, 2024Abstract:We propose a scaling law hypothesis for multimodal models processing text, audio, images, and video within a shared token and embedding space. Our framework predicts model performance based on modality-specific compression and tokenization efficiency, extending established scaling laws from text-based decoder models to mixed-modality systems. We explore whether leveraging more training data in multiple modalities can reduce the size of the multimodal model, enabling efficient deployment on resource-constrained devices.
Unveiling the Unseen: Exploring Whitebox Membership Inference through the Lens of Explainability
Jul 01, 2024



Abstract:The increasing prominence of deep learning applications and reliance on personalized data underscore the urgent need to address privacy vulnerabilities, particularly Membership Inference Attacks (MIAs). Despite numerous MIA studies, significant knowledge gaps persist, particularly regarding the impact of hidden features (in isolation) on attack efficacy and insufficient justification for the root causes of attacks based on raw data features. In this paper, we aim to address these knowledge gaps by first exploring statistical approaches to identify the most informative neurons and quantifying the significance of the hidden activations from the selected neurons on attack accuracy, in isolation and combination. Additionally, we propose an attack-driven explainable framework by integrating the target and attack models to identify the most influential features of raw data that lead to successful membership inference attacks. Our proposed MIA shows an improvement of up to 26% on state-of-the-art MIA.
Octo-planner: On-device Language Model for Planner-Action Agents
Jun 26, 2024



Abstract:AI agents have become increasingly significant in various domains, enabling autonomous decision-making and problem-solving. To function effectively, these agents require a planning process that determines the best course of action and then executes the planned actions. In this paper, we present an efficient on-device Planner-Action framework that separates planning and action execution into two distinct components: a planner agent based on Phi-3 Mini, a 3.8 billion parameter LLM optimized for edge devices, and an action agent using the Octopus model for function execution. The planner agent first responds to user queries by decomposing tasks into a sequence of sub-steps, which are then executed by the action agent. To optimize performance on resource-constrained devices, we employ model fine-tuning instead of in-context learning, reducing computational costs and energy consumption while improving response times. Our approach involves using GPT-4 to generate diverse planning queries and responses based on available functions, with subsequent validations to ensure data quality. We fine-tune the Phi-3 Mini model on this curated dataset, achieving a 97\% success rate in our in-domain test environment. To address multi-domain planning challenges, we developed a multi-LoRA training method that merges weights from LoRAs trained on distinct function subsets. This approach enables flexible handling of complex, multi-domain queries while maintaining computational efficiency on resource-constrained devices. To support further research, we have open-sourced our model weights at \url{https://huggingface.co/NexaAIDev/octopus-planning}. For the demo, please refer to \url{https://www.nexa4ai.com/octo-planner}.
More Compute Is What You Need
May 02, 2024
Abstract:Large language model pre-training has become increasingly expensive, with most practitioners relying on scaling laws to allocate compute budgets for model size and training tokens, commonly referred to as Compute-Optimal or Chinchilla Optimal. In this paper, we hypothesize a new scaling law that suggests model performance depends mostly on the amount of compute spent for transformer-based models, independent of the specific allocation to model size and dataset size. Using this unified scaling law, we predict that (a) for inference efficiency, training should prioritize smaller model sizes and larger training datasets, and (b) assuming the exhaustion of available web datasets, scaling the model size might be the only way to further improve model performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge