Graph-in-Graph Network for Automatic Gene Ontology Description Generation
Jun 10, 2022Fenglin Liu, Bang Yang, Chenyu You, Xian Wu, Shen Ge, Adelaide Woicik, Sheng Wang
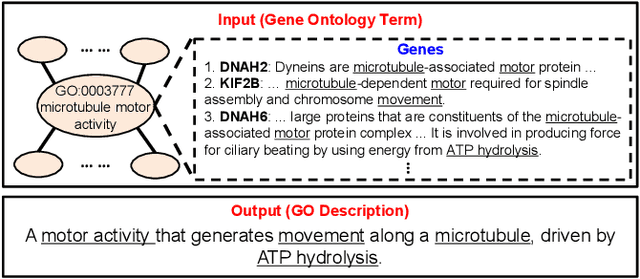
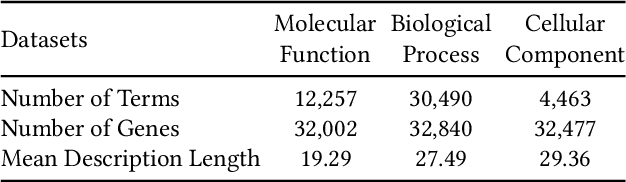
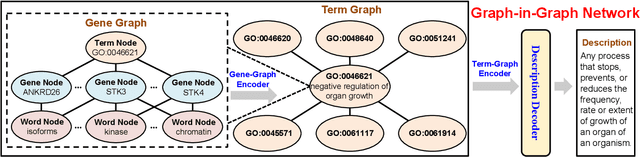
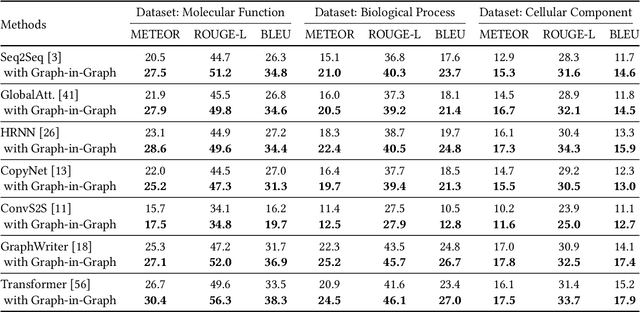
Gene Ontology (GO) is the primary gene function knowledge base that enables computational tasks in biomedicine. The basic element of GO is a term, which includes a set of genes with the same function. Existing research efforts of GO mainly focus on predicting gene term associations. Other tasks, such as generating descriptions of new terms, are rarely pursued. In this paper, we propose a novel task: GO term description generation. This task aims to automatically generate a sentence that describes the function of a GO term belonging to one of the three categories, i.e., molecular function, biological process, and cellular component. To address this task, we propose a Graph-in-Graph network that can efficiently leverage the structural information of GO. The proposed network introduces a two-layer graph: the first layer is a graph of GO terms where each node is also a graph (gene graph). Such a Graph-in-Graph network can derive the biological functions of GO terms and generate proper descriptions. To validate the effectiveness of the proposed network, we build three large-scale benchmark datasets. By incorporating the proposed Graph-in-Graph network, the performances of seven different sequence-to-sequence models can be substantially boosted across all evaluation metrics, with up to 34.7%, 14.5%, and 39.1% relative improvements in BLEU, ROUGE-L, and METEOR, respectively.
Hedging option books using neural-SDE market models
May 31, 2022Samuel N. Cohen, Christoph Reisinger, Sheng Wang
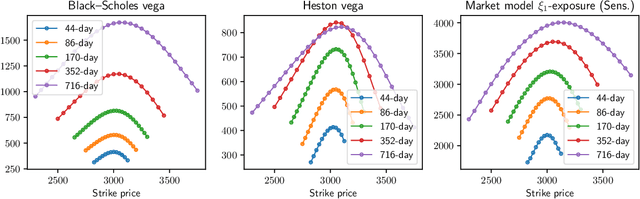

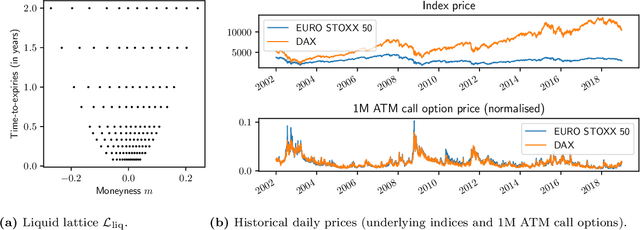

We study the capability of arbitrage-free neural-SDE market models to yield effective strategies for hedging options. In particular, we derive sensitivity-based and minimum-variance-based hedging strategies using these models and examine their performance when applied to various option portfolios using real-world data. Through backtesting analysis over typical and stressed market periods, we show that neural-SDE market models achieve lower hedging errors than Black--Scholes delta and delta-vega hedging consistently over time, and are less sensitive to the tenor choice of hedging instruments. In addition, hedging using market models leads to similar performance to hedging using Heston models, while the former tends to be more robust during stressed market periods.
Eye-gaze-guided Vision Transformer for Rectifying Shortcut Learning
May 25, 2022Chong Ma, Lin Zhao, Yuzhong Chen, Lu Zhang, Zhenxiang Xiao, Haixing Dai, David Liu, Zihao Wu, Zhengliang Liu, Sheng Wang, Jiaxing Gao, Changhe Li, Xi Jiang, Tuo Zhang, Qian Wang, Dinggang Shen, Dajiang Zhu, Tianming Liu
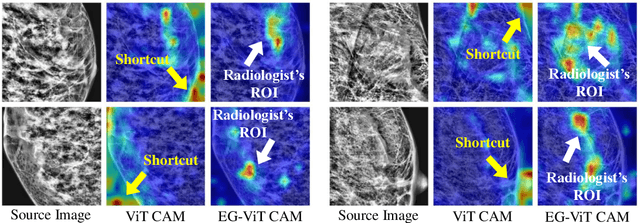
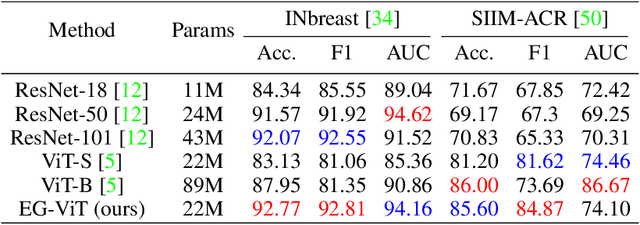
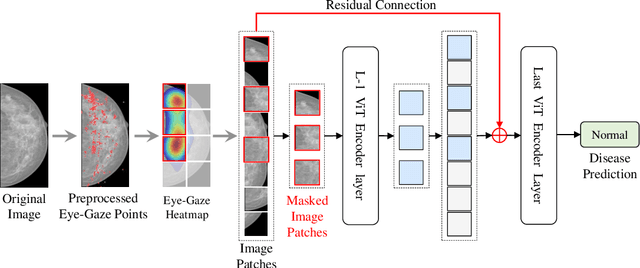
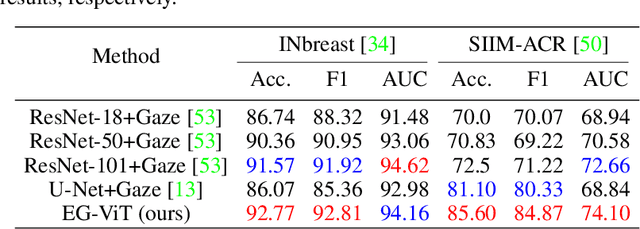
Learning harmful shortcuts such as spurious correlations and biases prevents deep neural networks from learning the meaningful and useful representations, thus jeopardizing the generalizability and interpretability of the learned representation. The situation becomes even more serious in medical imaging, where the clinical data (e.g., MR images with pathology) are limited and scarce while the reliability, generalizability and transparency of the learned model are highly required. To address this problem, we propose to infuse human experts' intelligence and domain knowledge into the training of deep neural networks. The core idea is that we infuse the visual attention information from expert radiologists to proactively guide the deep model to focus on regions with potential pathology and avoid being trapped in learning harmful shortcuts. To do so, we propose a novel eye-gaze-guided vision transformer (EG-ViT) for diagnosis with limited medical image data. We mask the input image patches that are out of the radiologists' interest and add an additional residual connection in the last encoder layer of EG-ViT to maintain the correlations of all patches. The experiments on two public datasets of INbreast and SIIM-ACR demonstrate our EG-ViT model can effectively learn/transfer experts' domain knowledge and achieve much better performance than baselines. Meanwhile, it successfully rectifies the harmful shortcut learning and significantly improves the EG-ViT model's interpretability. In general, EG-ViT takes the advantages of both human expert's prior knowledge and the power of deep neural networks. This work opens new avenues for advancing current artificial intelligence paradigms by infusing human intelligence.
Arbitrary Reduction of MRI Slice Spacing Based on Local-Aware Implicit Representation
May 23, 2022Xin Wang, Kai Xuan, Sheng Wang, Honglin Xiong, Lichi Zhang, Qian Wang
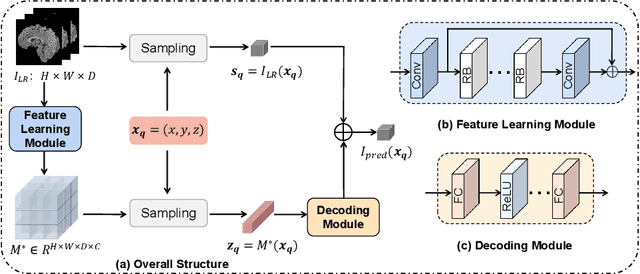
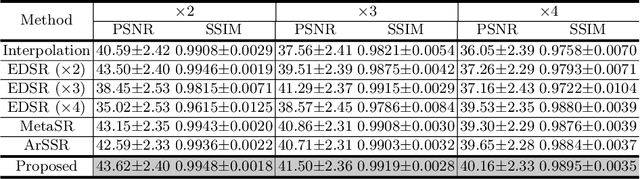
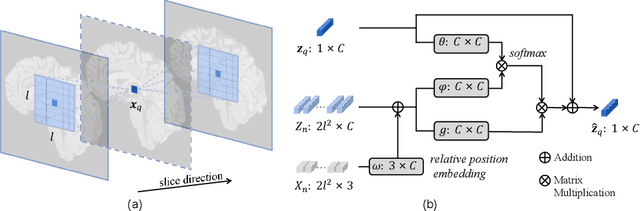
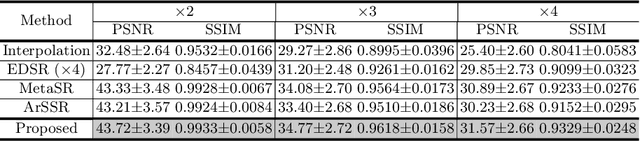
Magnetic resonance (MR) images are often acquired in 2D settings for real clinical applications. The 3D volumes reconstructed by stacking multiple 2D slices have large inter-slice spacing, resulting in lower inter-slice resolution than intra-slice resolution. Super-resolution is a powerful tool to reduce the inter-slice spacing of 3D images to facilitate subsequent visualization and computation tasks. However, most existing works train the super-resolution network at a fixed ratio, which is inconvenient in clinical scenes due to the heterogeneous parameters in MR scanning. In this paper, we propose a single super-resolution network to reduce the inter-slice spacing of MR images at an arbitrarily adjustable ratio. Specifically, we view the input image as a continuous implicit function of coordinates. The intermediate slices of different spacing ratios could be constructed according to the implicit representation up-sampled in the continuous domain. We particularly propose a novel local-aware spatial attention mechanism and long-range residual learning to boost the quality of the output image. The experimental results demonstrate the superiority of our proposed method, even compared to the models trained at a fixed ratio.
Seed-Guided Topic Discovery with Out-of-Vocabulary Seeds
May 04, 2022Yu Zhang, Yu Meng, Xuan Wang, Sheng Wang, Jiawei Han
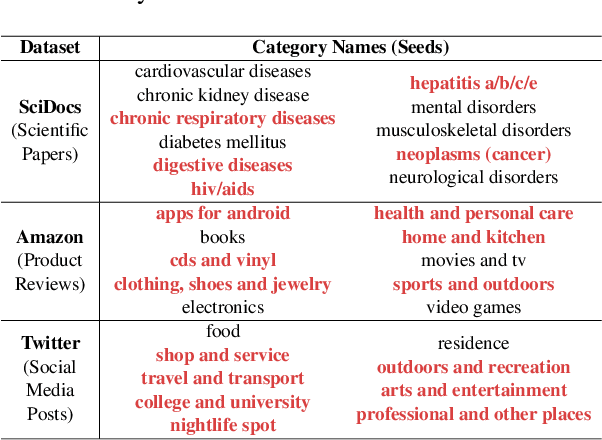
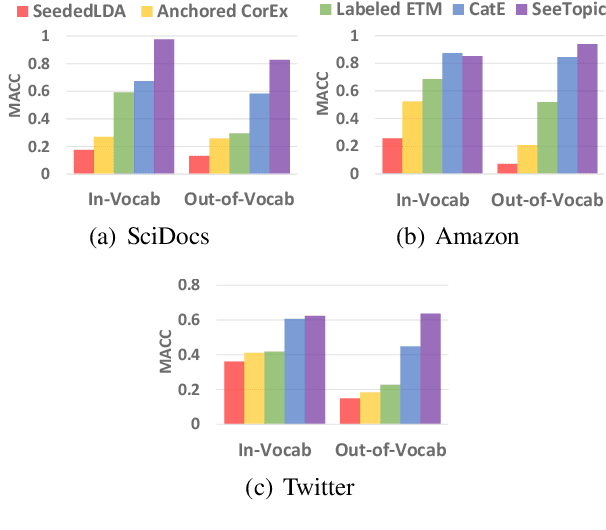
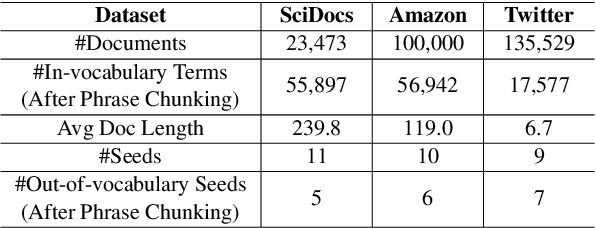
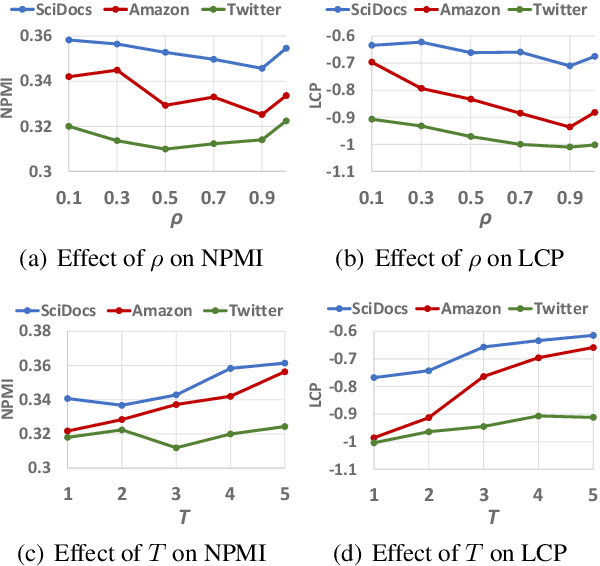
Discovering latent topics from text corpora has been studied for decades. Many existing topic models adopt a fully unsupervised setting, and their discovered topics may not cater to users' particular interests due to their inability of leveraging user guidance. Although there exist seed-guided topic discovery approaches that leverage user-provided seeds to discover topic-representative terms, they are less concerned with two factors: (1) the existence of out-of-vocabulary seeds and (2) the power of pre-trained language models (PLMs). In this paper, we generalize the task of seed-guided topic discovery to allow out-of-vocabulary seeds. We propose a novel framework, named SeeTopic, wherein the general knowledge of PLMs and the local semantics learned from the input corpus can mutually benefit each other. Experiments on three real datasets from different domains demonstrate the effectiveness of SeeTopic in terms of topic coherence, accuracy, and diversity.
Follow My Eye: Using Gaze to Supervise Computer-Aided Diagnosis
Apr 06, 2022Sheng Wang, Xi Ouyang, Tianming Liu, Qian Wang, Dinggang Shen
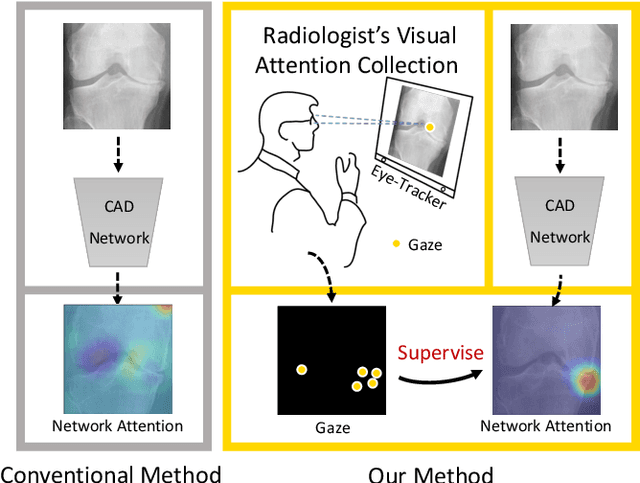
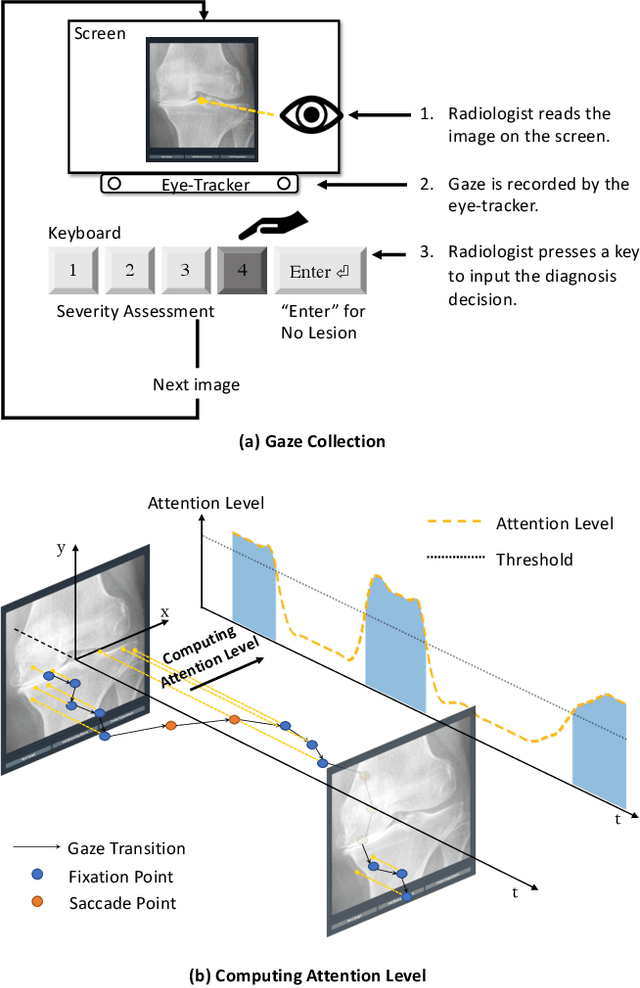
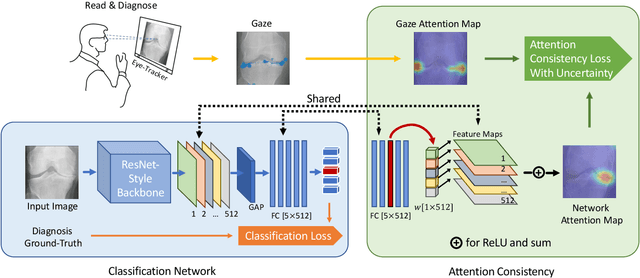
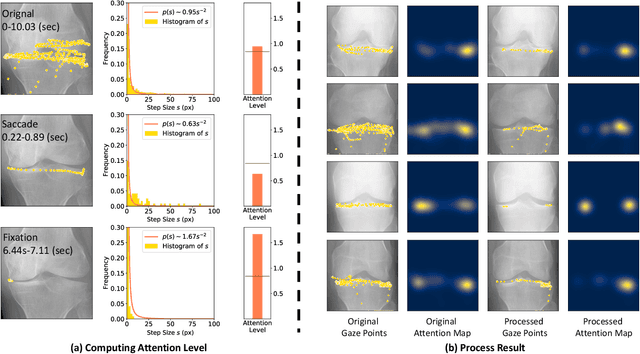
When deep neural network (DNN) was first introduced to the medical image analysis community, researchers were impressed by its performance. However, it is evident now that a large number of manually labeled data is often a must to train a properly functioning DNN. This demand for supervision data and labels is a major bottleneck in current medical image analysis, since collecting a large number of annotations from experienced experts can be time-consuming and expensive. In this paper, we demonstrate that the eye movement of radiologists reading medical images can be a new form of supervision to train the DNN-based computer-aided diagnosis (CAD) system. Particularly, we record the tracks of the radiologists' gaze when they are reading images. The gaze information is processed and then used to supervise the DNN's attention via an Attention Consistency module. To the best of our knowledge, the above pipeline is among the earliest efforts to leverage expert eye movement for deep-learning-based CAD. We have conducted extensive experiments on knee X-ray images for osteoarthritis assessment. The results show that our method can achieve considerable improvement in diagnosis performance, with the help of gaze supervision.
Estimating risks of option books using neural-SDE market models
Feb 15, 2022Samuel N. Cohen, Christoph Reisinger, Sheng Wang
In this paper, we examine the capacity of an arbitrage-free neural-SDE market model to produce realistic scenarios for the joint dynamics of multiple European options on a single underlying. We subsequently demonstrate its use as a risk simulation engine for option portfolios. Through backtesting analysis, we show that our models are more computationally efficient and accurate for evaluating the Value-at-Risk (VaR) of option portfolios, with better coverage performance and less procyclicality than standard filtered historical simulation approaches.
Adaptive Transfer Learning for Plant Phenotyping
Jan 14, 2022Jun Wu, Elizabeth A. Ainsworth, Sheng Wang, Kaiyu Guan, Jingrui He
Plant phenotyping (Guo et al. 2021; Pieruschka et al. 2019) focuses on studying the diverse traits of plants related to the plants' growth. To be more specific, by accurately measuring the plant's anatomical, ontogenetical, physiological and biochemical properties, it allows identifying the crucial factors of plants' growth in different environments. One commonly used approach is to predict the plant's traits using hyperspectral reflectance (Yendrek et al. 2017; Wang et al. 2021). However, the data distributions of the hyperspectral reflectance data in plant phenotyping might vary in different environments for different plants. That is, it would be computationally expansive to learn the machine learning models separately for one plant in different environments. To solve this problem, we focus on studying the knowledge transferability of modern machine learning models in plant phenotyping. More specifically, this work aims to answer the following questions. (1) How is the performance of conventional machine learning models, e.g., partial least squares regression (PLSR), Gaussian process regression (GPR) and multi-layer perceptron (MLP), affected by the number of annotated samples for plant phenotyping? (2) Whether could the neural network based transfer learning models improve the performance of plant phenotyping? (3) Could the neural network based transfer learning be improved by using infinite-width hidden layers for plant phenotyping?
Knee Cartilage Defect Assessment by Graph Representation and Surface Convolution
Jan 12, 2022Zixu Zhuang, Liping Si, Sheng Wang, Kai Xuan, Xi Ouyang, Yiqiang Zhan, Zhong Xue, Lichi Zhang, Dinggang Shen, Weiwu Yao, Qian Wang
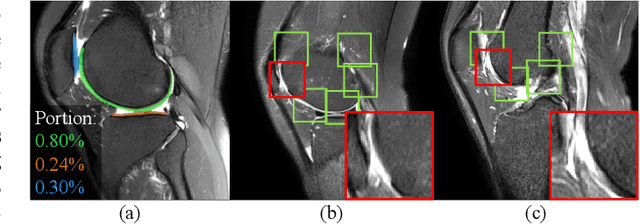
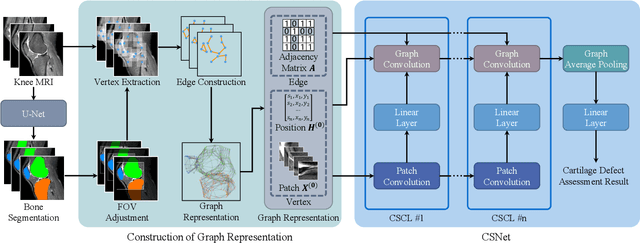
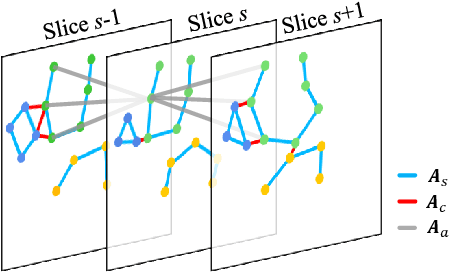
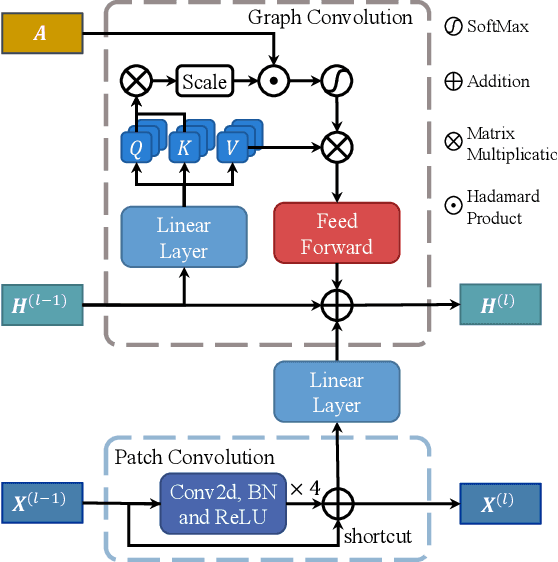
Knee osteoarthritis (OA) is the most common osteoarthritis and a leading cause of disability. Cartilage defects are regarded as major manifestations of knee OA, which are visible by magnetic resonance imaging (MRI). Thus early detection and assessment for knee cartilage defects are important for protecting patients from knee OA. In this way, many attempts have been made on knee cartilage defect assessment by applying convolutional neural networks (CNNs) to knee MRI. However, the physiologic characteristics of the cartilage may hinder such efforts: the cartilage is a thin curved layer, implying that only a small portion of voxels in knee MRI can contribute to the cartilage defect assessment; heterogeneous scanning protocols further challenge the feasibility of the CNNs in clinical practice; the CNN-based knee cartilage evaluation results lack interpretability. To address these challenges, we model the cartilages structure and appearance from knee MRI into a graph representation, which is capable of handling highly diverse clinical data. Then, guided by the cartilage graph representation, we design a non-Euclidean deep learning network with the self-attention mechanism, to extract cartilage features in the local and global, and to derive the final assessment with a visualized result. Our comprehensive experiments show that the proposed method yields superior performance in knee cartilage defect assessment, plus its convenient 3D visualization for interpretability.
SurvODE: Extrapolating Gene Expression Distribution for Early Cancer Identification
Nov 30, 2021Tong Chen, Sheng Wang
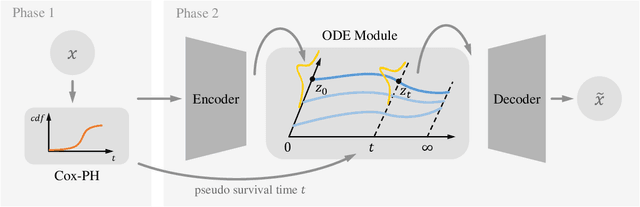
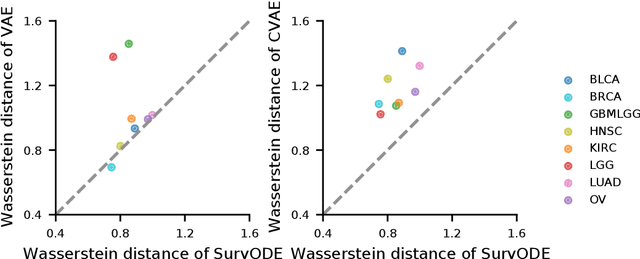
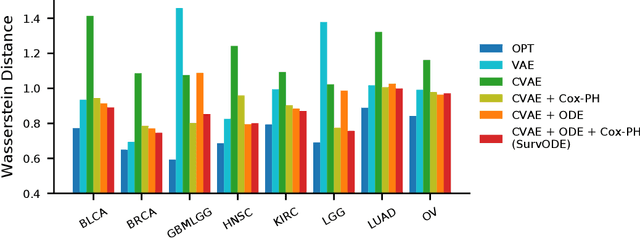
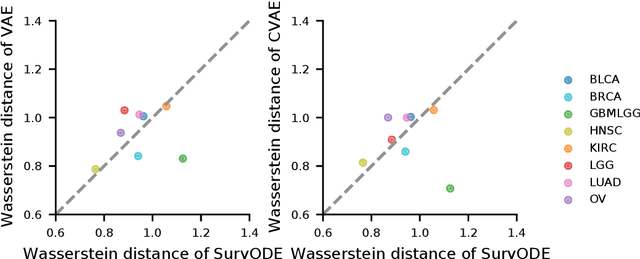
With the increasingly available large-scale cancer genomics datasets, machine learning approaches have played an important role in revealing novel insights into cancer development. Existing methods have shown encouraging performance in identifying genes that are predictive for cancer survival, but are still limited in modeling the distribution over genes. Here, we proposed a novel method that can simulate the gene expression distribution at any given time point, including those that are out of the range of the observed time points. In order to model the irregular time series where each patient is one observation, we integrated a neural ordinary differential equation (neural ODE) with cox regression into our framework. We evaluated our method on eight cancer types on TCGA and observed a substantial improvement over existing approaches. Our visualization results and further analysis indicate how our method can be used to simulate expression at the early cancer stage, offering the possibility for early cancer identification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge