Attention-aware non-rigid image registration for accelerated MR imaging
Apr 26, 2024Aya Ghoul, Jiazhen Pan, Andreas Lingg, Jens Kübler, Patrick Krumm, Kerstin Hammernik, Daniel Rueckert, Sergios Gatidis, Thomas Küstner
Accurate motion estimation at high acceleration factors enables rapid motion-compensated reconstruction in Magnetic Resonance Imaging (MRI) without compromising the diagnostic image quality. In this work, we introduce an attention-aware deep learning-based framework that can perform non-rigid pairwise registration for fully sampled and accelerated MRI. We extract local visual representations to build similarity maps between the registered image pairs at multiple resolution levels and additionally leverage long-range contextual information using a transformer-based module to alleviate ambiguities in the presence of artifacts caused by undersampling. We combine local and global dependencies to perform simultaneous coarse and fine motion estimation. The proposed method was evaluated on in-house acquired fully sampled and accelerated data of 101 patients and 62 healthy subjects undergoing cardiac and thoracic MRI. The impact of motion estimation accuracy on the downstream task of motion-compensated reconstruction was analyzed. We demonstrate that our model derives reliable and consistent motion fields across different sampling trajectories (Cartesian and radial) and acceleration factors of up to 16x for cardiac motion and 30x for respiratory motion and achieves superior image quality in motion-compensated reconstruction qualitatively and quantitatively compared to conventional and recent deep learning-based approaches. The code is publicly available at https://github.com/lab-midas/GMARAFT.
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023Jianning Li, Antonio Pepe, Christina Gsaxner, Gijs Luijten, Yuan Jin, Narmada Ambigapathy, Enrico Nasca, Naida Solak, Gian Marco Melito, Viet Duc Vu, Afaque R. Memon, Xiaojun Chen, Jan Stefan Kirschke, Ezequiel de la Rosa, Patrick Ferdinand Christ, Hongwei Bran Li, David G. Ellis, Michele R. Aizenberg, Sergios Gatidis, Thomas Küstner, Nadya Shusharina, Nicholas Heller, Vincent Andrearczyk, Adrien Depeursinge, Mathieu Hatt, Anjany Sekuboyina, Maximilian Löffler, Hans Liebl, Reuben Dorent, Tom Vercauteren, Jonathan Shapey, Aaron Kujawa, Stefan Cornelissen, Patrick Langenhuizen, Achraf Ben-Hamadou, Ahmed Rekik, Sergi Pujades, Edmond Boyer, Federico Bolelli, Costantino Grana, Luca Lumetti, Hamidreza Salehi, Jun Ma, Yao Zhang, Ramtin Gharleghi, Susann Beier, Arcot Sowmya, Eduardo A. Garza-Villarreal, Thania Balducci, Diego Angeles-Valdez, Roberto Souza, Leticia Rittner, Richard Frayne, Yuanfeng Ji, Soumick Chatterjee, Florian Dubost, Stefanie Schreiber, Hendrik Mattern, Oliver Speck, Daniel Haehn, Christoph John, Andreas Nürnberger, João Pedrosa, Carlos Ferreira, Guilherme Aresta, António Cunha, Aurélio Campilho, Yannick Suter, Jose Garcia, Alain Lalande, Emmanuel Audenaert, Claudia Krebs, Timo Van Leeuwen, Evie Vereecke, Rainer Röhrig, Frank Hölzle, Vahid Badeli, Kathrin Krieger, Matthias Gunzer, Jianxu Chen, Amin Dada, Miriam Balzer, Jana Fragemann, Frederic Jonske, Moritz Rempe, Stanislav Malorodov, Fin H. Bahnsen, Constantin Seibold, Alexander Jaus, Ana Sofia Santos, Mariana Lindo, André Ferreira, Victor Alves, Michael Kamp, Amr Abourayya, Felix Nensa, Fabian Hörst, Alexander Brehmer, Lukas Heine, Lars E. Podleska, Matthias A. Fink, Julius Keyl, Konstantinos Tserpes, Moon-Sung Kim, Shireen Elhabian, Hans Lamecker, Dženan Zukić, Beatriz Paniagua, Christian Wachinger, Martin Urschler, Luc Duong, Jakob Wasserthal, Peter F. Hoyer, Oliver Basu, Thomas Maal, Max J. H. Witjes, Ti-chiun Chang, Seyed-Ahmad Ahmadi, Ping Luo, Bjoern Menze, Mauricio Reyes, Christos Davatzikos, Behrus Puladi, Jens Kleesiek, Jan Egger
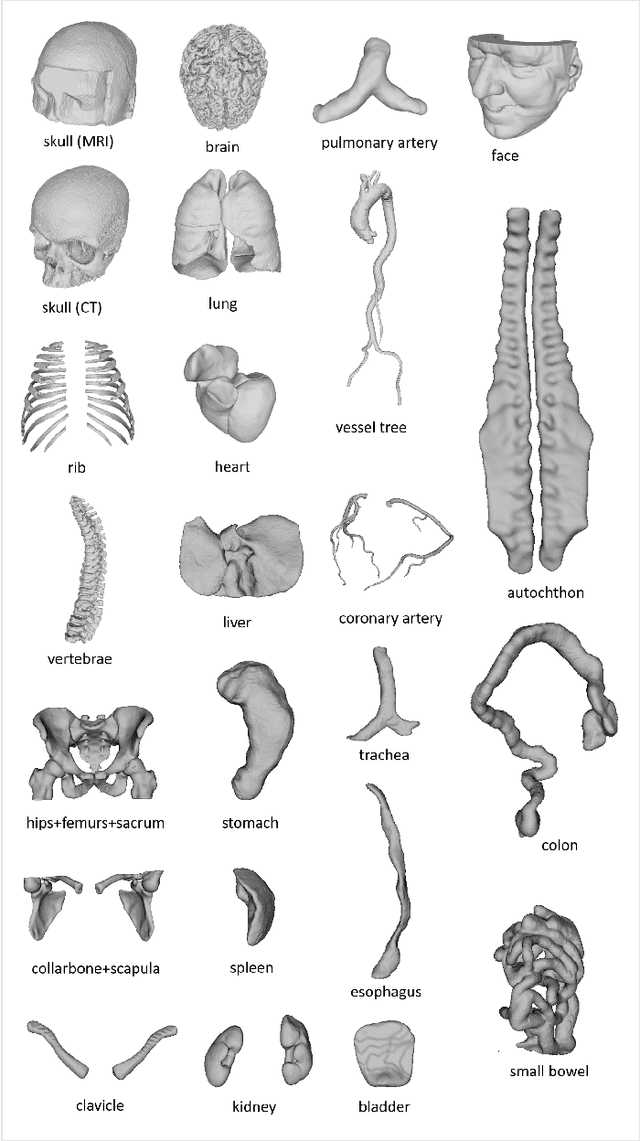

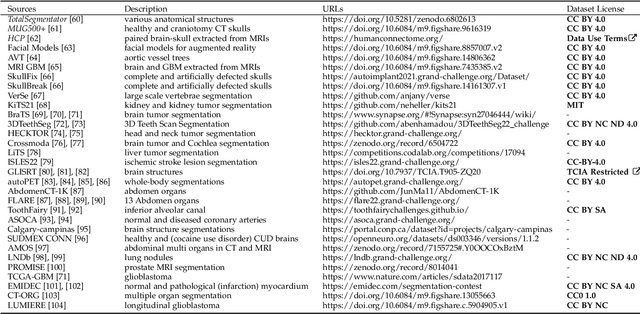
We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
Uncertainty Estimation and Propagation in Accelerated MRI Reconstruction
Aug 04, 2023Paul Fischer, Thomas Küstner, Christian F. Baumgartner
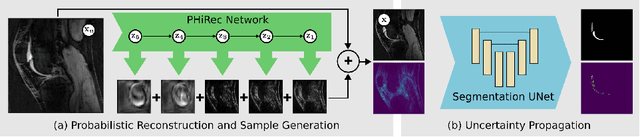
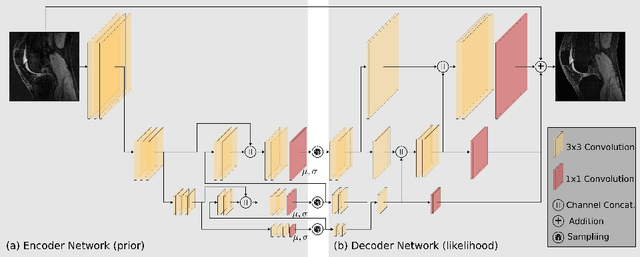
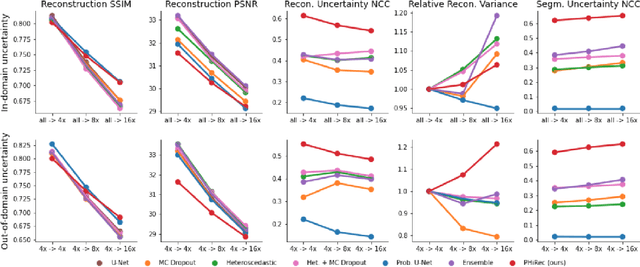
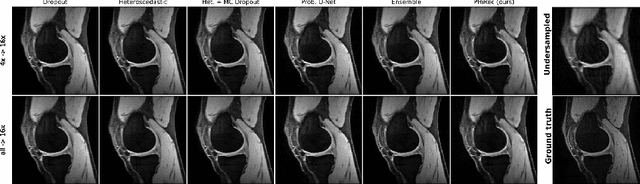
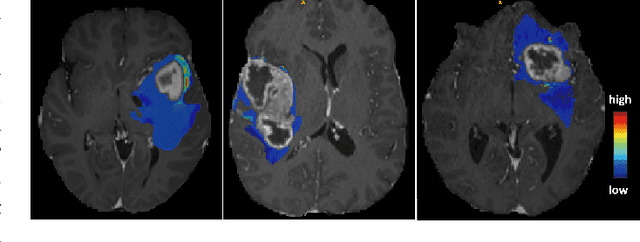
MRI reconstruction techniques based on deep learning have led to unprecedented reconstruction quality especially in highly accelerated settings. However, deep learning techniques are also known to fail unexpectedly and hallucinate structures. This is particularly problematic if reconstructions are directly used for downstream tasks such as real-time treatment guidance or automated extraction of clinical paramters (e.g. via segmentation). Well-calibrated uncertainty quantification will be a key ingredient for safe use of this technology in clinical practice. In this paper we propose a novel probabilistic reconstruction technique (PHiRec) building on the idea of conditional hierarchical variational autoencoders. We demonstrate that our proposed method produces high-quality reconstructions as well as uncertainty quantification that is substantially better calibrated than several strong baselines. We furthermore demonstrate how uncertainties arising in the MR econstruction can be propagated to a downstream segmentation task, and show that PHiRec also allows well-calibrated estimation of segmentation uncertainties that originated in the MR reconstruction process.
Global k-Space Interpolation for Dynamic MRI Reconstruction using Masked Image Modeling
Jul 24, 2023Jiazhen Pan, Suprosanna Shit, Özgün Turgut, Wenqi Huang, Hongwei Bran Li, Nil Stolt-Ansó, Thomas Küstner, Kerstin Hammernik, Daniel Rueckert
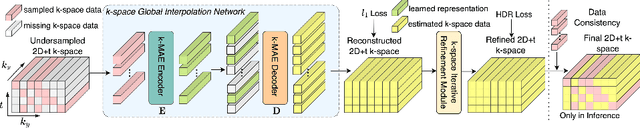
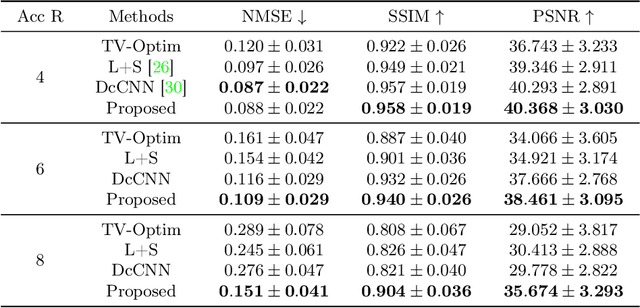
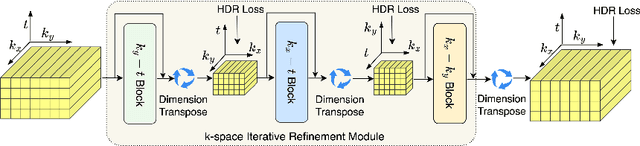
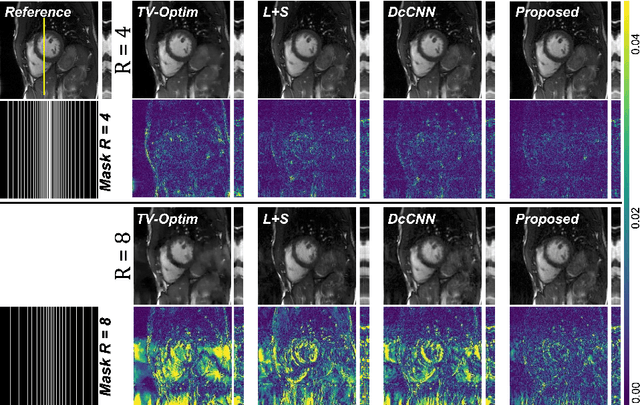
In dynamic Magnetic Resonance Imaging (MRI), k-space is typically undersampled due to limited scan time, resulting in aliasing artifacts in the image domain. Hence, dynamic MR reconstruction requires not only modeling spatial frequency components in the x and y directions of k-space but also considering temporal redundancy. Most previous works rely on image-domain regularizers (priors) to conduct MR reconstruction. In contrast, we focus on interpolating the undersampled k-space before obtaining images with Fourier transform. In this work, we connect masked image modeling with k-space interpolation and propose a novel Transformer-based k-space Global Interpolation Network, termed k-GIN. Our k-GIN learns global dependencies among low- and high-frequency components of 2D+t k-space and uses it to interpolate unsampled data. Further, we propose a novel k-space Iterative Refinement Module (k-IRM) to enhance the high-frequency components learning. We evaluate our approach on 92 in-house 2D+t cardiac MR subjects and compare it to MR reconstruction methods with image-domain regularizers. Experiments show that our proposed k-space interpolation method quantitatively and qualitatively outperforms baseline methods. Importantly, the proposed approach achieves substantially higher robustness and generalizability in cases of highly-undersampled MR data.
Reconstruction-driven motion estimation for motion-compensated MR CINE imaging
Feb 05, 2023Jiazhen Pan, Wenqi Huang, Daniel Rueckert, Thomas Küstner, Kerstin Hammernik
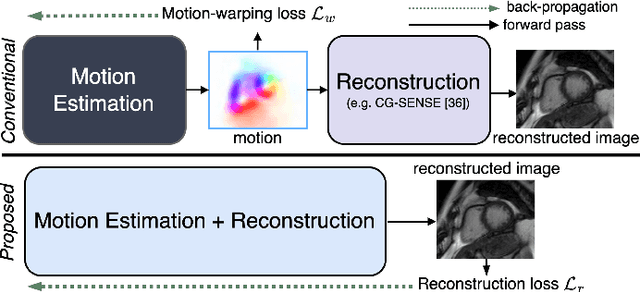
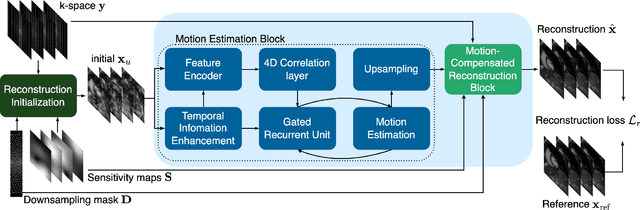
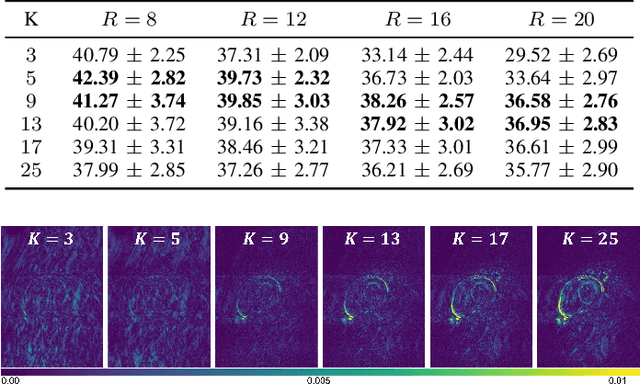
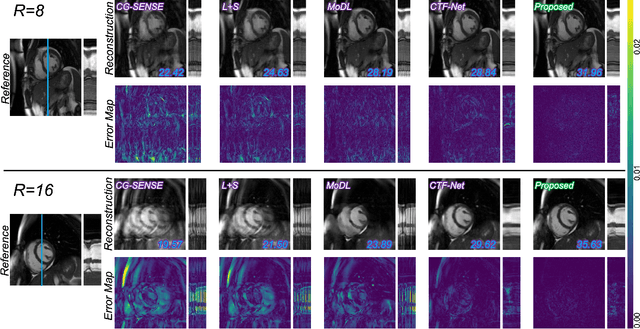
In cardiac CINE, motion-compensated MR reconstruction (MCMR) is an effective approach to address highly undersampled acquisitions by incorporating motion information between frames. In this work, we propose a deep learning-based framework to address the MCMR problem efficiently. Contrary to state-of-the-art (SOTA) MCMR methods which break the original problem into two sub-optimization problems, i.e. motion estimation and reconstruction, we formulate this problem as a single entity with one single optimization. We discard the canonical motion-warping loss (similarity measurement between motion-warped images and target images) to estimate the motion, but drive the motion estimation process directly by the final reconstruction performance. The higher reconstruction quality is achieved without using any smoothness loss terms and without iterative processing between motion estimation and reconstruction. Therefore, we avoid non-trivial loss weighting factors tuning and time-consuming iterative processing. Experiments on 43 in-house acquired 2D CINE datasets indicate that the proposed MCMR framework can deliver artifact-free motion estimation and high-quality MR images even for imaging accelerations up to 20x. The proposed framework is compared to SOTA non-MCMR and MCMR methods and outperforms these methods qualitatively and quantitatively in all applied metrics across all experiments with different acceleration rates.
Learning-based and unrolled motion-compensated reconstruction for cardiac MR CINE imaging
Sep 08, 2022Jiazhen Pan, Daniel Rueckert, Thomas Küstner, Kerstin Hammernik




Motion-compensated MR reconstruction (MCMR) is a powerful concept with considerable potential, consisting of two coupled sub-problems: Motion estimation, assuming a known image, and image reconstruction, assuming known motion. In this work, we propose a learning-based self-supervised framework for MCMR, to efficiently deal with non-rigid motion corruption in cardiac MR imaging. Contrary to conventional MCMR methods in which the motion is estimated prior to reconstruction and remains unchanged during the iterative optimization process, we introduce a dynamic motion estimation process and embed it into the unrolled optimization. We establish a cardiac motion estimation network that leverages temporal information via a group-wise registration approach, and carry out a joint optimization between the motion estimation and reconstruction. Experiments on 40 acquired 2D cardiac MR CINE datasets demonstrate that the proposed unrolled MCMR framework can reconstruct high quality MR images at high acceleration rates where other state-of-the-art methods fail. We also show that the joint optimization mechanism is mutually beneficial for both sub-tasks, i.e., motion estimation and image reconstruction, especially when the MR image is highly undersampled.
Physics-Driven Deep Learning for Computational Magnetic Resonance Imaging
Mar 23, 2022Kerstin Hammernik, Thomas Küstner, Burhaneddin Yaman, Zhengnan Huang, Daniel Rueckert, Florian Knoll, Mehmet Akçakaya
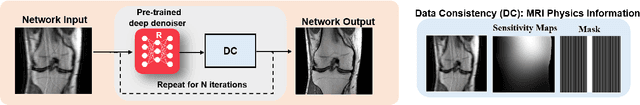
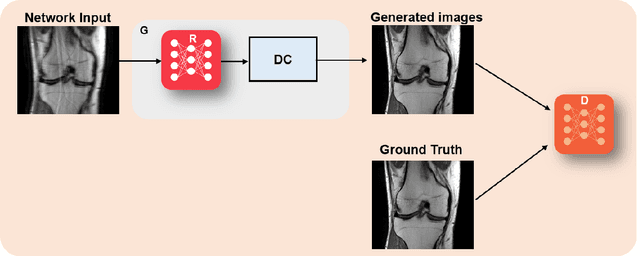
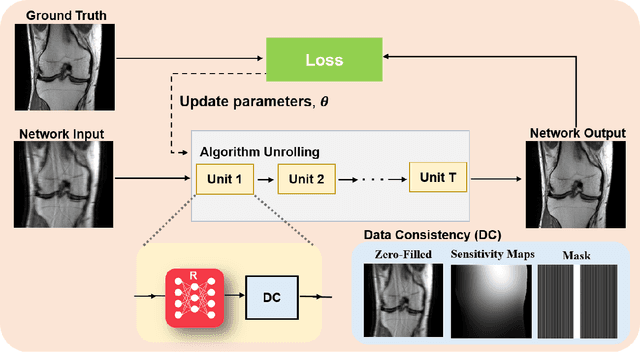
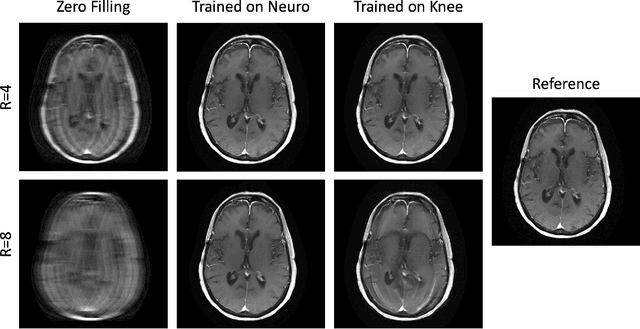
Physics-driven deep learning methods have emerged as a powerful tool for computational magnetic resonance imaging (MRI) problems, pushing reconstruction performance to new limits. This article provides an overview of the recent developments in incorporating physics information into learning-based MRI reconstruction. We consider inverse problems with both linear and non-linear forward models for computational MRI, and review the classical approaches for solving these. We then focus on physics-driven deep learning approaches, covering physics-driven loss functions, plug-and-play methods, generative models, and unrolled networks. We highlight domain-specific challenges such as real- and complex-valued building blocks of neural networks, and translational applications in MRI with linear and non-linear forward models. Finally, we discuss common issues and open challenges, and draw connections to the importance of physics-driven learning when combined with other downstream tasks in the medical imaging pipeline.
LAPNet: Non-rigid Registration derived in k-space for Magnetic Resonance Imaging
Jul 19, 2021Thomas Küstner, Jiazhen Pan, Haikun Qi, Gastao Cruz, Christopher Gilliam, Thierry Blu, Bin Yang, Sergios Gatidis, René Botnar, Claudia Prieto
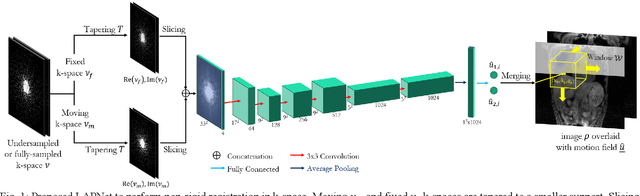
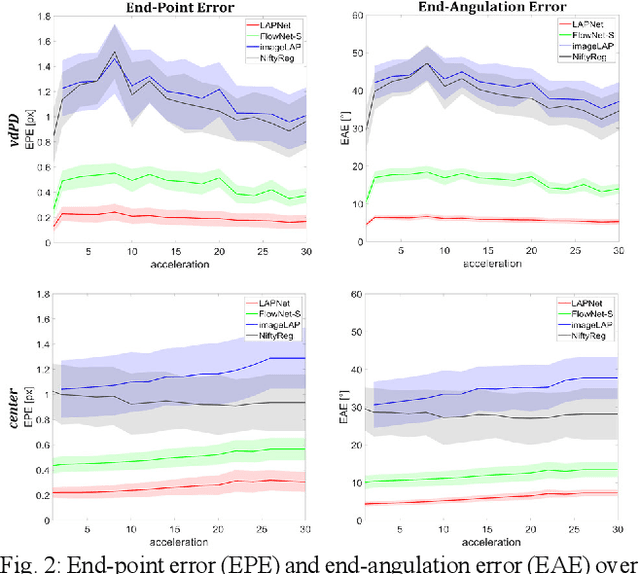
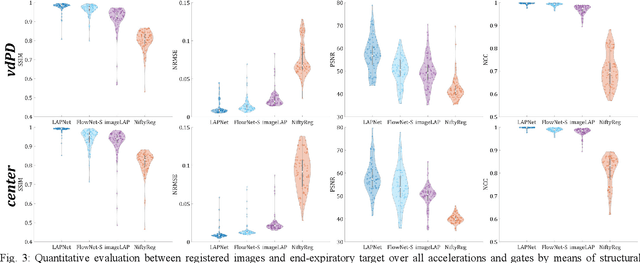
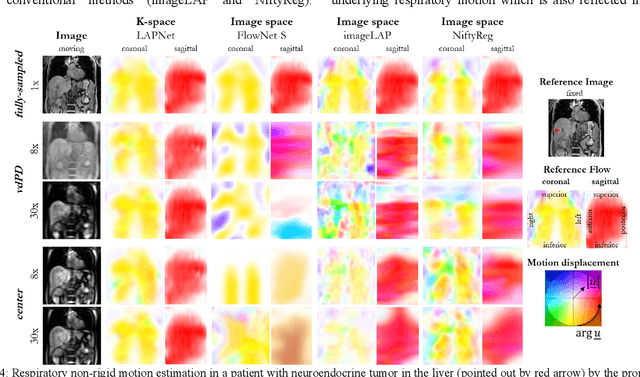
Physiological motion, such as cardiac and respiratory motion, during Magnetic Resonance (MR) image acquisition can cause image artifacts. Motion correction techniques have been proposed to compensate for these types of motion during thoracic scans, relying on accurate motion estimation from undersampled motion-resolved reconstruction. A particular interest and challenge lie in the derivation of reliable non-rigid motion fields from the undersampled motion-resolved data. Motion estimation is usually formulated in image space via diffusion, parametric-spline, or optical flow methods. However, image-based registration can be impaired by remaining aliasing artifacts due to the undersampled motion-resolved reconstruction. In this work, we describe a formalism to perform non-rigid registration directly in the sampled Fourier space, i.e. k-space. We propose a deep-learning based approach to perform fast and accurate non-rigid registration from the undersampled k-space data. The basic working principle originates from the Local All-Pass (LAP) technique, a recently introduced optical flow-based registration. The proposed LAPNet is compared against traditional and deep learning image-based registrations and tested on fully-sampled and highly-accelerated (with two undersampling strategies) 3D respiratory motion-resolved MR images in a cohort of 40 patients with suspected liver or lung metastases and 25 healthy subjects. The proposed LAPNet provided consistent and superior performance to image-based approaches throughout different sampling trajectories and acceleration factors.
Complementary Time-Frequency Domain Networks for Dynamic Parallel MR Image Reconstruction
Dec 22, 2020Chen Qin, Jinming Duan, Kerstin Hammernik, Jo Schlemper, Thomas Küstner, René Botnar, Claudia Prieto, Anthony N. Price, Joseph V. Hajnal, Daniel Rueckert
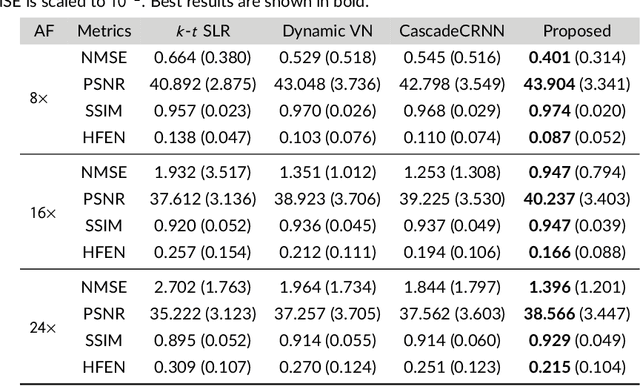
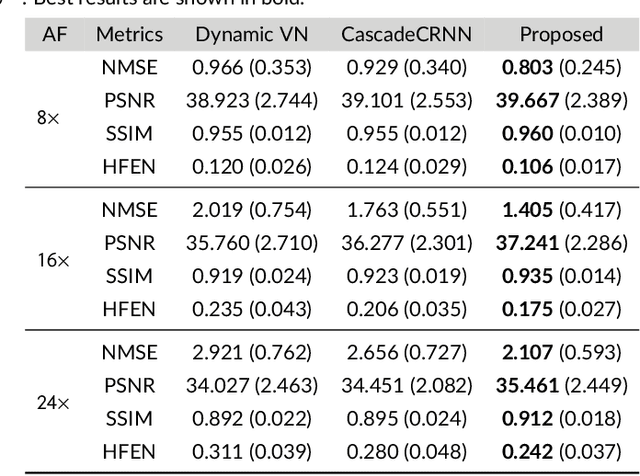
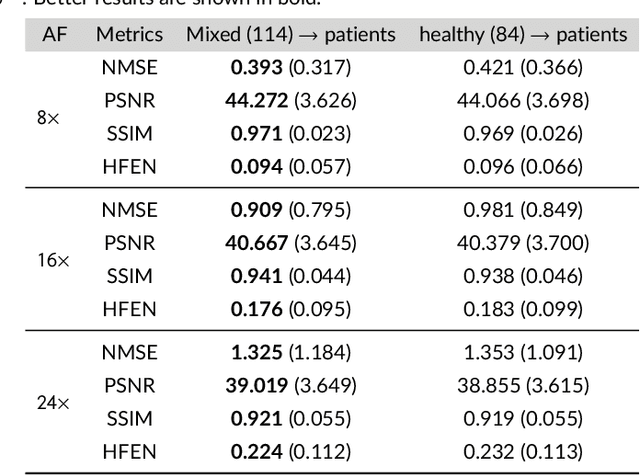
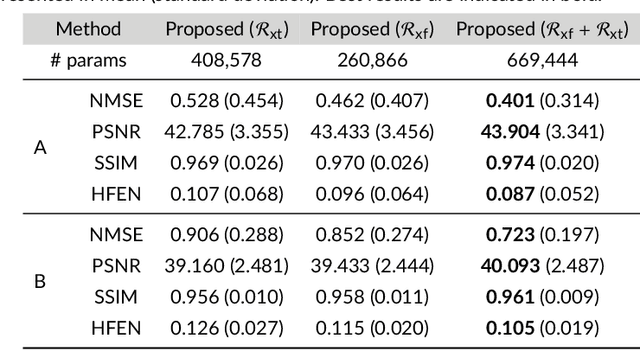
Purpose: To introduce a novel deep learning based approach for fast and high-quality dynamic multi-coil MR reconstruction by learning a complementary time-frequency domain network that exploits spatio-temporal correlations simultaneously from complementary domains. Theory and Methods: Dynamic parallel MR image reconstruction is formulated as a multi-variable minimisation problem, where the data is regularised in combined temporal Fourier and spatial (x-f) domain as well as in spatio-temporal image (x-t) domain. An iterative algorithm based on variable splitting technique is derived, which alternates among signal de-aliasing steps in x-f and x-t spaces, a closed-form point-wise data consistency step and a weighted coupling step. The iterative model is embedded into a deep recurrent neural network which learns to recover the image via exploiting spatio-temporal redundancies in complementary domains. Results: Experiments were performed on two datasets of highly undersampled multi-coil short-axis cardiac cine MRI scans. Results demonstrate that our proposed method outperforms the current state-of-the-art approaches both quantitatively and qualitatively. The proposed model can also generalise well to data acquired from a different scanner and data with pathologies that were not seen in the training set. Conclusion: The work shows the benefit of reconstructing dynamic parallel MRI in complementary time-frequency domains with deep neural networks. The method can effectively and robustly reconstruct high-quality images from highly undersampled dynamic multi-coil data ($16 \times$ and $24 \times$ yielding 15s and 10s scan times respectively) with fast reconstruction speed (2.8s). This could potentially facilitate achieving fast single-breath-hold clinical 2D cardiac cine imaging.
Age-Net: An MRI-Based Iterative Framework for Biological Age Estimation
Sep 22, 2020Karim Armanious, Sherif Abdulatif, Wenbin Shi, Shashank Salian, Thomas Küstner, Daniel Weiskopf, Tobias Hepp, Sergios Gatidis, Bin Yang
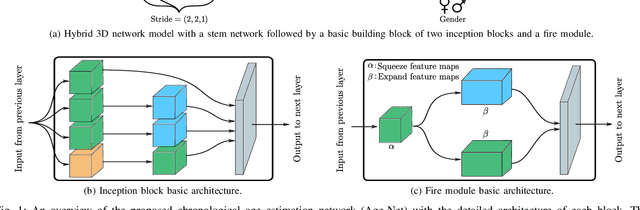
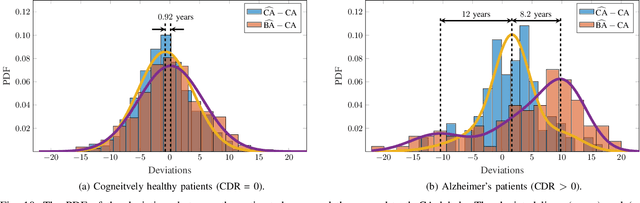
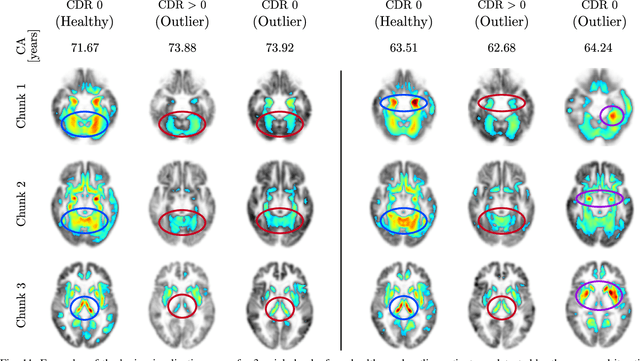
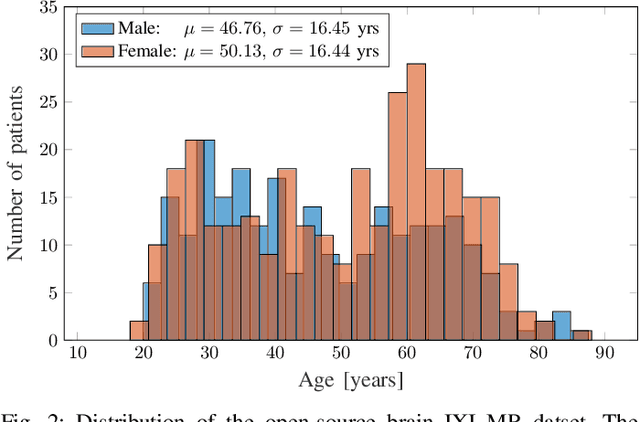
The concept of biological age (BA) - although important in clinical practice - is hard to grasp mainly due to lack of a clearly defined reference standard. For specific applications, especially in pediatrics, medical image data are used for BA estimation in a routine clinical context. Beyond this young age group, BA estimation is restricted to whole-body assessment using non-imaging indicators such as blood biomarkers, genetic and cellular data. However, various organ systems may exhibit different aging characteristics due to lifestyle and genetic factors. Thus, a whole-body assessment of the BA does not reflect the deviations of aging behavior between organs. To this end, we propose a new imaging-based framework for organ-specific BA estimation. As a first step, we introduce a chronological age (CA) estimation framework using deep convolutional neural networks (Age-Net). We quantitatively assess the performance of this framework in comparison to existing CA estimation approaches. Furthermore, we expand upon Age-Net with a novel iterative data-cleaning algorithm to segregate atypical-aging patients (BA $\not \approx$ CA) from the given population. In this manner, we hypothesize that the remaining population should approximate the true BA behaviour. For this initial study, we apply the proposed methodology on a brain magnetic resonance image (MRI) dataset containing healthy individuals as well as Alzheimer's patients with different dementia ratings. We demonstrate the correlation between the predicted BAs and the expected cognitive deterioration in Alzheimer's patients. A statistical and visualization-based analysis has provided evidence regarding the potential and current challenges of the proposed methodology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge