Nadya Shusharina
Treatment-wise Glioblastoma Survival Inference with Multi-parametric Preoperative MRI
Feb 10, 2024Abstract:In this work, we aim to predict the survival time (ST) of glioblastoma (GBM) patients undergoing different treatments based on preoperative magnetic resonance (MR) scans. The personalized and precise treatment planning can be achieved by comparing the ST of different treatments. It is well established that both the current status of the patient (as represented by the MR scans) and the choice of treatment are the cause of ST. While previous related MR-based glioblastoma ST studies have focused only on the direct mapping of MR scans to ST, they have not included the underlying causal relationship between treatments and ST. To address this limitation, we propose a treatment-conditioned regression model for glioblastoma ST that incorporates treatment information in addition to MR scans. Our approach allows us to effectively utilize the data from all of the treatments in a unified manner, rather than having to train separate models for each of the treatments. Furthermore, treatment can be effectively injected into each convolutional layer through the adaptive instance normalization we employ. We evaluate our framework on the BraTS20 ST prediction task. Three treatment options are considered: Gross Total Resection (GTR), Subtotal Resection (STR), and no resection. The evaluation results demonstrate the effectiveness of injecting the treatment for estimating GBM survival.
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023



Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
ACT: Semi-supervised Domain-adaptive Medical Image Segmentation with Asymmetric Co-training
Jun 09, 2022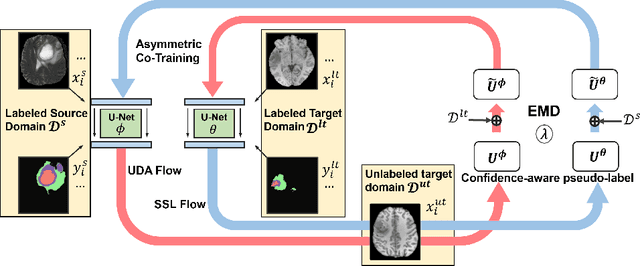
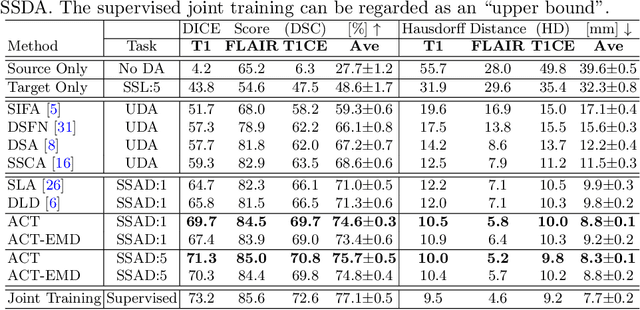
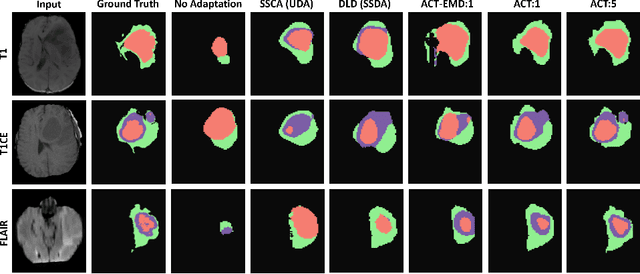
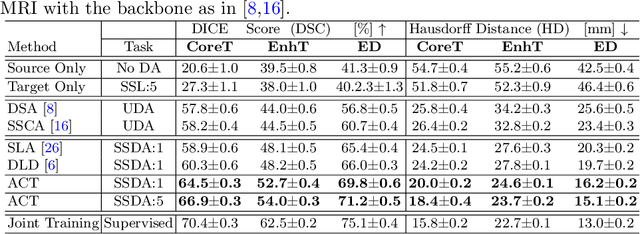
Abstract:Unsupervised domain adaptation (UDA) has been vastly explored to alleviate domain shifts between source and target domains, by applying a well-performed model in an unlabeled target domain via supervision of a labeled source domain. Recent literature, however, has indicated that the performance is still far from satisfactory in the presence of significant domain shifts. Nonetheless, delineating a few target samples is usually manageable and particularly worthwhile, due to the substantial performance gain. Inspired by this, we aim to develop semi-supervised domain adaptation (SSDA) for medical image segmentation, which is largely underexplored. We, thus, propose to exploit both labeled source and target domain data, in addition to unlabeled target data in a unified manner. Specifically, we present a novel asymmetric co-training (ACT) framework to integrate these subsets and avoid the domination of the source domain data. Following a divide-and-conquer strategy, we explicitly decouple the label supervisions in SSDA into two asymmetric sub-tasks, including semi-supervised learning (SSL) and UDA, and leverage different knowledge from two segmentors to take into account the distinction between the source and target label supervisions. The knowledge learned in the two modules is then adaptively integrated with ACT, by iteratively teaching each other, based on the confidence-aware pseudo-label. In addition, pseudo label noise is well-controlled with an exponential MixUp decay scheme for smooth propagation. Experiments on cross-modality brain tumor MRI segmentation tasks using the BraTS18 database showed, even with limited labeled target samples, ACT yielded marked improvements over UDA and state-of-the-art SSDA methods and approached an "upper bound" of supervised joint training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge