Shuo Wang
State Key Laboratory of Management and Control for Complex Systems, Institute of Automation, Chinese Academy of Sciences, China
Graph-Based Spectral Decomposition for Parameter Coordination in Language Model Fine-Tuning
Apr 28, 2025Abstract:This paper proposes a parameter collaborative optimization algorithm for large language models, enhanced with graph spectral analysis. The goal is to improve both fine-tuning efficiency and structural awareness during training. In the proposed method, the parameters of a pre-trained language model are treated as nodes in a graph. A weighted graph is constructed, and Laplacian spectral decomposition is applied to enable frequency-domain modeling and structural representation of the parameter space. Based on this structure, a joint loss function is designed. It combines the task loss with a spectral regularization term to facilitate collaborative updates among parameters. In addition, a spectral filtering mechanism is introduced during the optimization phase. This mechanism adjusts gradients in a structure-aware manner, enhancing the model's training stability and convergence behavior. The method is evaluated on multiple tasks, including traditional fine-tuning comparisons, few-shot generalization tests, and convergence speed analysis. In all settings, the proposed approach demonstrates superior performance. The experimental results confirm that the spectral collaborative optimization framework effectively reduces parameter perturbations and improves fine-tuning quality while preserving overall model performance. This work contributes significantly to the field of artificial intelligence by advancing parameter-efficient training methodologies for large-scale models, reinforcing the importance of structural signal processing in deep learning optimization, and offering a robust, generalizable framework for enhancing language model adaptability and performance.
Diffusion-Driven Universal Model Inversion Attack for Face Recognition
Apr 25, 2025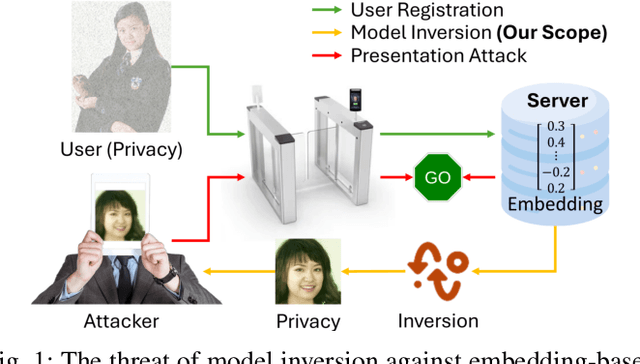
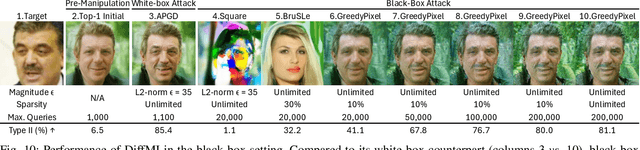
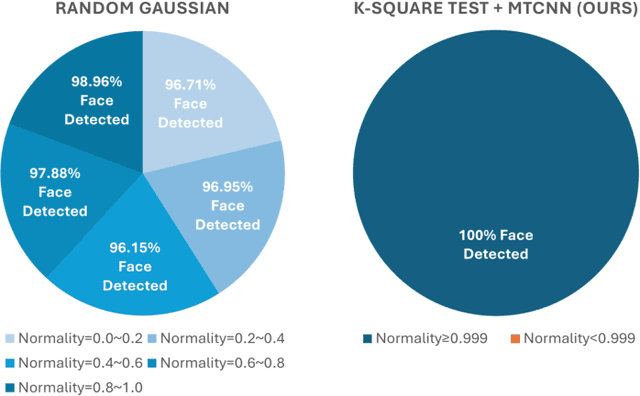
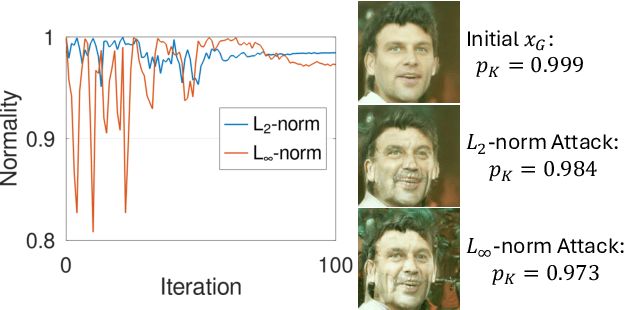
Abstract:Facial recognition technology poses significant privacy risks, as it relies on biometric data that is inherently sensitive and immutable if compromised. To mitigate these concerns, face recognition systems convert raw images into embeddings, traditionally considered privacy-preserving. However, model inversion attacks pose a significant privacy threat by reconstructing these private facial images, making them a crucial tool for evaluating the privacy risks of face recognition systems. Existing methods usually require training individual generators for each target model, a computationally expensive process. In this paper, we propose DiffUMI, a training-free diffusion-driven universal model inversion attack for face recognition systems. DiffUMI is the first approach to apply a diffusion model for unconditional image generation in model inversion. Unlike other methods, DiffUMI is universal, eliminating the need for training target-specific generators. It operates within a fixed framework and pretrained diffusion model while seamlessly adapting to diverse target identities and models. DiffUMI breaches privacy-preserving face recognition systems with state-of-the-art success, demonstrating that an unconditional diffusion model, coupled with optimized adversarial search, enables efficient and high-fidelity facial reconstruction. Additionally, we introduce a novel application of out-of-domain detection (OODD), marking the first use of model inversion to distinguish non-face inputs from face inputs based solely on embeddings.
RT-DATR:Real-time Unsupervised Domain Adaptive Detection Transformer with Adversarial Feature Learning
Apr 12, 2025



Abstract:Despite domain-adaptive object detectors based on CNN and transformers have made significant progress in cross-domain detection tasks, it is regrettable that domain adaptation for real-time transformer-based detectors has not yet been explored. Directly applying existing domain adaptation algorithms has proven to be suboptimal. In this paper, we propose RT-DATR, a simple and efficient real-time domain adaptive detection transformer. Building on RT-DETR as our base detector, we first introduce a local object-level feature alignment module to significantly enhance the feature representation of domain invariance during object transfer. Additionally, we introduce a scene semantic feature alignment module designed to boost cross-domain detection performance by aligning scene semantic features. Finally, we introduced a domain query and decoupled it from the object query to further align the instance feature distribution within the decoder layer, reduce the domain gap, and maintain discriminative ability. Experimental results on various benchmarks demonstrate that our method outperforms current state-of-the-art approaches. Our code will be released soon.
LLM$\times$MapReduce-V2: Entropy-Driven Convolutional Test-Time Scaling for Generating Long-Form Articles from Extremely Long Resources
Apr 08, 2025Abstract:Long-form generation is crucial for a wide range of practical applications, typically categorized into short-to-long and long-to-long generation. While short-to-long generations have received considerable attention, generating long texts from extremely long resources remains relatively underexplored. The primary challenge in long-to-long generation lies in effectively integrating and analyzing relevant information from extensive inputs, which remains difficult for current large language models (LLMs). In this paper, we propose LLM$\times$MapReduce-V2, a novel test-time scaling strategy designed to enhance the ability of LLMs to process extremely long inputs. Drawing inspiration from convolutional neural networks, which iteratively integrate local features into higher-level global representations, LLM$\times$MapReduce-V2 utilizes stacked convolutional scaling layers to progressively expand the understanding of input materials. Both quantitative and qualitative experimental results demonstrate that our approach substantially enhances the ability of LLMs to process long inputs and generate coherent, informative long-form articles, outperforming several representative baselines.
Improving Harmful Text Detection with Joint Retrieval and External Knowledge
Apr 03, 2025Abstract:Harmful text detection has become a crucial task in the development and deployment of large language models, especially as AI-generated content continues to expand across digital platforms. This study proposes a joint retrieval framework that integrates pre-trained language models with knowledge graphs to improve the accuracy and robustness of harmful text detection. Experimental results demonstrate that the joint retrieval approach significantly outperforms single-model baselines, particularly in low-resource training scenarios and multilingual environments. The proposed method effectively captures nuanced harmful content by leveraging external contextual information, addressing the limitations of traditional detection models. Future research should focus on optimizing computational efficiency, enhancing model interpretability, and expanding multimodal detection capabilities to better tackle evolving harmful content patterns. This work contributes to the advancement of AI safety, ensuring more trustworthy and reliable content moderation systems.
Benchmarking Federated Machine Unlearning methods for Tabular Data
Apr 01, 2025



Abstract:Machine unlearning, which enables a model to forget specific data upon request, is increasingly relevant in the era of privacy-centric machine learning, particularly within federated learning (FL) environments. This paper presents a pioneering study on benchmarking machine unlearning methods within a federated setting for tabular data, addressing the unique challenges posed by cross-silo FL where data privacy and communication efficiency are paramount. We explore unlearning at the feature and instance levels, employing both machine learning, random forest and logistic regression models. Our methodology benchmarks various unlearning algorithms, including fine-tuning and gradient-based approaches, across multiple datasets, with metrics focused on fidelity, certifiability, and computational efficiency. Experiments demonstrate that while fidelity remains high across methods, tree-based models excel in certifiability, ensuring exact unlearning, whereas gradient-based methods show improved computational efficiency. This study provides critical insights into the design and selection of unlearning algorithms tailored to the FL environment, offering a foundation for further research in privacy-preserving machine learning.
Why Stop at One Error? Benchmarking LLMs as Data Science Code Debuggers for Multi-Hop and Multi-Bug Errors
Mar 28, 2025Abstract:LLMs are transforming software development, yet current code generation and code repair benchmarks mainly assess syntactic and functional correctness in simple, single-error cases. LLMs' capabilities to autonomously find and fix runtime logical errors in complex data science code remain largely unexplored. To address this gap, we introduce DSDBench: the Data Science Debugging Benchmark, the first benchmark for systematic evaluation of LLMs on multi-hop error tracing and multi-bug detection in data science code debugging. DSDBench adapts datasets from existing data science task benchmarks, such as DABench and MatPlotBench, featuring realistic data science debugging tasks with automatically synthesized multi-hop, multi-bug code snippets. DSDBench includes 1,117 annotated samples with 741 cause-effect error pairs and runtime error messages. Evaluations of state-of-the-art LLMs on DSDBench show significant performance gaps, highlighting challenges in debugging logical runtime errors in data science code. DSDBench offers a crucial resource to evaluate and improve LLMs' debugging and reasoning capabilities, enabling more reliable AI-assisted data science in the future.DSDBench is publicly available at https://github.com/KevinCL16/DSDBench.
Exploring CLIP's Dense Knowledge for Weakly Supervised Semantic Segmentation
Mar 26, 2025



Abstract:Weakly Supervised Semantic Segmentation (WSSS) with image-level labels aims to achieve pixel-level predictions using Class Activation Maps (CAMs). Recently, Contrastive Language-Image Pre-training (CLIP) has been introduced in WSSS. However, recent methods primarily focus on image-text alignment for CAM generation, while CLIP's potential in patch-text alignment remains unexplored. In this work, we propose ExCEL to explore CLIP's dense knowledge via a novel patch-text alignment paradigm for WSSS. Specifically, we propose Text Semantic Enrichment (TSE) and Visual Calibration (VC) modules to improve the dense alignment across both text and vision modalities. To make text embeddings semantically informative, our TSE module applies Large Language Models (LLMs) to build a dataset-wide knowledge base and enriches the text representations with an implicit attribute-hunting process. To mine fine-grained knowledge from visual features, our VC module first proposes Static Visual Calibration (SVC) to propagate fine-grained knowledge in a non-parametric manner. Then Learnable Visual Calibration (LVC) is further proposed to dynamically shift the frozen features towards distributions with diverse semantics. With these enhancements, ExCEL not only retains CLIP's training-free advantages but also significantly outperforms other state-of-the-art methods with much less training cost on PASCAL VOC and MS COCO.
RoMA: Scaling up Mamba-based Foundation Models for Remote Sensing
Mar 13, 2025Abstract:Recent advances in self-supervised learning for Vision Transformers (ViTs) have fueled breakthroughs in remote sensing (RS) foundation models. However, the quadratic complexity of self-attention poses a significant barrier to scalability, particularly for large models and high-resolution images. While the linear-complexity Mamba architecture offers a promising alternative, existing RS applications of Mamba remain limited to supervised tasks on small, domain-specific datasets. To address these challenges, we propose RoMA, a framework that enables scalable self-supervised pretraining of Mamba-based RS foundation models using large-scale, diverse, unlabeled data. RoMA enhances scalability for high-resolution images through a tailored auto-regressive learning strategy, incorporating two key innovations: 1) a rotation-aware pretraining mechanism combining adaptive cropping with angular embeddings to handle sparsely distributed objects with arbitrary orientations, and 2) multi-scale token prediction objectives that address the extreme variations in object scales inherent to RS imagery. Systematic empirical studies validate that Mamba adheres to RS data and parameter scaling laws, with performance scaling reliably as model and data size increase. Furthermore, experiments across scene classification, object detection, and semantic segmentation tasks demonstrate that RoMA-pretrained Mamba models consistently outperform ViT-based counterparts in both accuracy and computational efficiency. The source code and pretrained models will be released at https://github.com/MiliLab/RoMA.
TLA: Tactile-Language-Action Model for Contact-Rich Manipulation
Mar 11, 2025Abstract:Significant progress has been made in vision-language models. However, language-conditioned robotic manipulation for contact-rich tasks remains underexplored, particularly in terms of tactile sensing. To address this gap, we introduce the Tactile-Language-Action (TLA) model, which effectively processes sequential tactile feedback via cross-modal language grounding to enable robust policy generation in contact-intensive scenarios. In addition, we construct a comprehensive dataset that contains 24k pairs of tactile action instruction data, customized for fingertip peg-in-hole assembly, providing essential resources for TLA training and evaluation. Our results show that TLA significantly outperforms traditional imitation learning methods (e.g., diffusion policy) in terms of effective action generation and action accuracy, while demonstrating strong generalization capabilities by achieving over 85\% success rate on previously unseen assembly clearances and peg shapes. We publicly release all data and code in the hope of advancing research in language-conditioned tactile manipulation skill learning. Project website: https://sites.google.com/view/tactile-language-action/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge