Pheng-Ann Heng
DR-Label: Improving GNN Models for Catalysis Systems by Label Deconstruction and Reconstruction
Mar 06, 2023Abstract:Attaining the equilibrium state of a catalyst-adsorbate system is key to fundamentally assessing its effective properties, such as adsorption energy. Machine learning methods with finer supervision strategies have been applied to boost and guide the relaxation process of an atomic system and better predict its properties at the equilibrium state. In this paper, we present a novel graph neural network (GNN) supervision and prediction strategy DR-Label. The method enhances the supervision signal, reduces the multiplicity of solutions in edge representation, and encourages the model to provide node predictions that are graph structural variation robust. DR-Label first Deconstructs finer-grained equilibrium state information to the model by projecting the node-level supervision signal to each edge. Reversely, the model Reconstructs a more robust equilibrium state prediction by transforming edge-level predictions to node-level with a sphere-fitting algorithm. The DR-Label strategy was applied to three radically distinct models, each of which displayed consistent performance enhancements. Based on the DR-Label strategy, we further proposed DRFormer, which achieved a new state-of-the-art performance on the Open Catalyst 2020 (OC20) dataset and the Cu-based single-atom-alloyed CO adsorption (SAA) dataset. We expect that our work will highlight crucial steps for the development of a more accurate model in equilibrium state property prediction of a catalysis system.
RepMode: Learning to Re-parameterize Diverse Experts for Subcellular Structure Prediction
Dec 20, 2022



Abstract:In subcellular biological research, fluorescence staining is a key technique to reveal the locations and morphology of subcellular structures. However, fluorescence staining is slow, expensive, and harmful to cells. In this paper, we treat it as a deep learning task termed subcellular structure prediction (SSP), aiming to predict the 3D fluorescent images of multiple subcellular structures from a 3D transmitted-light image. Unfortunately, due to the limitations of current biotechnology, each image is partially labeled in SSP. Besides, naturally, the subcellular structures vary considerably in size, which causes the multi-scale issue in SSP. However, traditional solutions can not address SSP well since they organize network parameters inefficiently and inflexibly. To overcome these challenges, we propose Re-parameterizing Mixture-of-Diverse-Experts (RepMode), a network that dynamically organizes its parameters with task-aware priors to handle specified single-label prediction tasks of SSP. In RepMode, the Mixture-of-Diverse-Experts (MoDE) block is designed to learn the generalized parameters for all tasks, and gating re-parameterization (GatRep) is performed to generate the specialized parameters for each task, by which RepMode can maintain a compact practical topology exactly like a plain network, and meanwhile achieves a powerful theoretical topology. Comprehensive experiments show that RepMode outperforms existing methods on ten of twelve prediction tasks of SSP and achieves state-of-the-art overall performance.
Video Instance Shadow Detection
Nov 23, 2022



Abstract:Video instance shadow detection aims to simultaneously detect, segment, associate, and track paired shadow-object associations in videos. This work has three key contributions to the task. First, we design SSIS-Track, a new framework to extract shadow-object associations in videos with paired tracking and without category specification; especially, we strive to maintain paired tracking even the objects/shadows are temporarily occluded for several frames. Second, we leverage both labeled images and unlabeled videos, and explore temporal coherence by augmenting the tracking ability via an association cycle consistency loss to optimize SSIS-Track's performance. Last, we build $\textit{SOBA-VID}$, a new dataset with 232 unlabeled videos of ${5,863}$ frames for training and 60 labeled videos of ${1,182}$ frames for testing. Experimental results show that SSIS-Track surpasses baselines built from SOTA video tracking and instance-shadow-detection methods by a large margin. In the end, we showcase several video-level applications.
Dual Multi-scale Mean Teacher Network for Semi-supervised Infection Segmentation in Chest CT Volume for COVID-19
Nov 10, 2022Abstract:Automated detecting lung infections from computed tomography (CT) data plays an important role for combating COVID-19. However, there are still some challenges for developing AI system. 1) Most current COVID-19 infection segmentation methods mainly relied on 2D CT images, which lack 3D sequential constraint. 2) Existing 3D CT segmentation methods focus on single-scale representations, which do not achieve the multiple level receptive field sizes on 3D volume. 3) The emergent breaking out of COVID-19 makes it hard to annotate sufficient CT volumes for training deep model. To address these issues, we first build a multiple dimensional-attention convolutional neural network (MDA-CNN) to aggregate multi-scale information along different dimension of input feature maps and impose supervision on multiple predictions from different CNN layers. Second, we assign this MDA-CNN as a basic network into a novel dual multi-scale mean teacher network (DM${^2}$T-Net) for semi-supervised COVID-19 lung infection segmentation on CT volumes by leveraging unlabeled data and exploring the multi-scale information. Our DM${^2}$T-Net encourages multiple predictions at different CNN layers from the student and teacher networks to be consistent for computing a multi-scale consistency loss on unlabeled data, which is then added to the supervised loss on the labeled data from multiple predictions of MDA-CNN. Third, we collect two COVID-19 segmentation datasets to evaluate our method. The experimental results show that our network consistently outperforms the compared state-of-the-art methods.
Domain-incremental Cardiac Image Segmentation with Style-oriented Replay and Domain-sensitive Feature Whitening
Nov 09, 2022Abstract:Contemporary methods have shown promising results on cardiac image segmentation, but merely in static learning, i.e., optimizing the network once for all, ignoring potential needs for model updating. In real-world scenarios, new data continues to be gathered from multiple institutions over time and new demands keep growing to pursue more satisfying performance. The desired model should incrementally learn from each incoming dataset and progressively update with improved functionality as time goes by. As the datasets sequentially delivered from multiple sites are normally heterogenous with domain discrepancy, each updated model should not catastrophically forget previously learned domains while well generalizing to currently arrived domains or even unseen domains. In medical scenarios, this is particularly challenging as accessing or storing past data is commonly not allowed due to data privacy. To this end, we propose a novel domain-incremental learning framework to recover past domain inputs first and then regularly replay them during model optimization. Particularly, we first present a style-oriented replay module to enable structure-realistic and memory-efficient reproduction of past data, and then incorporate the replayed past data to jointly optimize the model with current data to alleviate catastrophic forgetting. During optimization, we additionally perform domain-sensitive feature whitening to suppress model's dependency on features that are sensitive to domain changes (e.g., domain-distinctive style features) to assist domain-invariant feature exploration and gradually improve the generalization performance of the network. We have extensively evaluated our approach with the M&Ms Dataset in single-domain and compound-domain incremental learning settings with improved performance over other comparison approaches.
A Survey on Generative Diffusion Model
Sep 21, 2022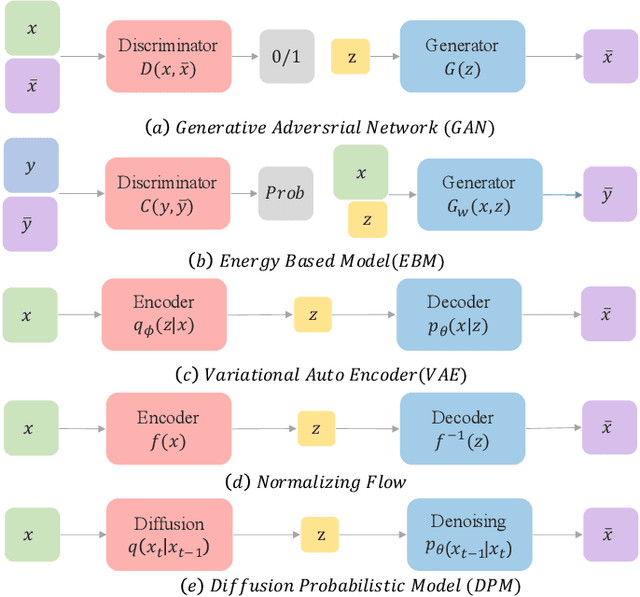
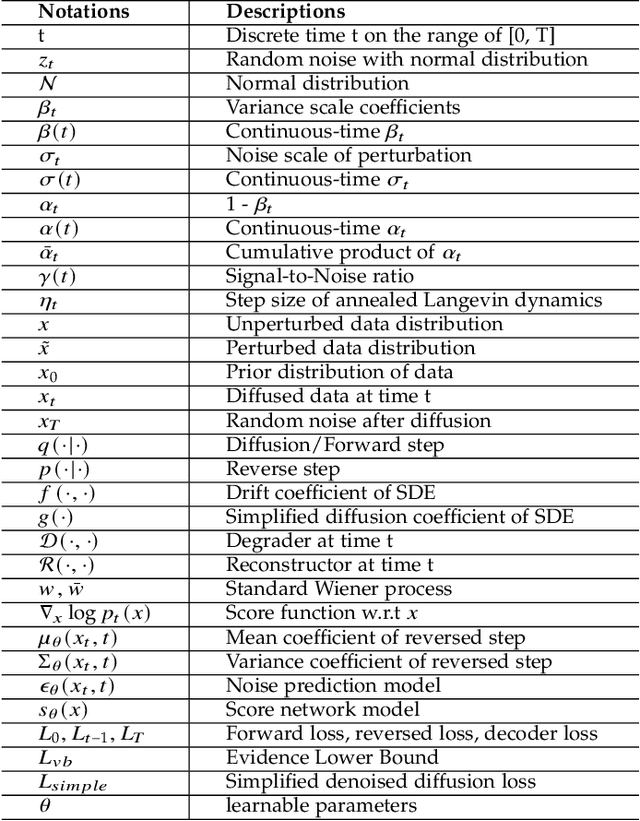
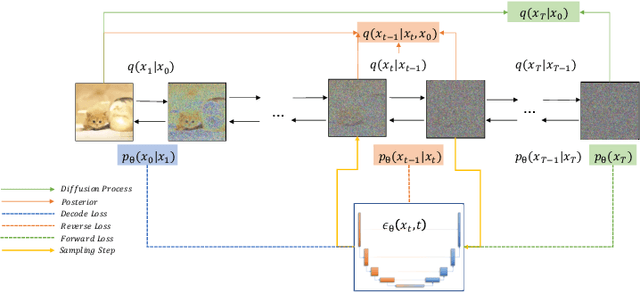
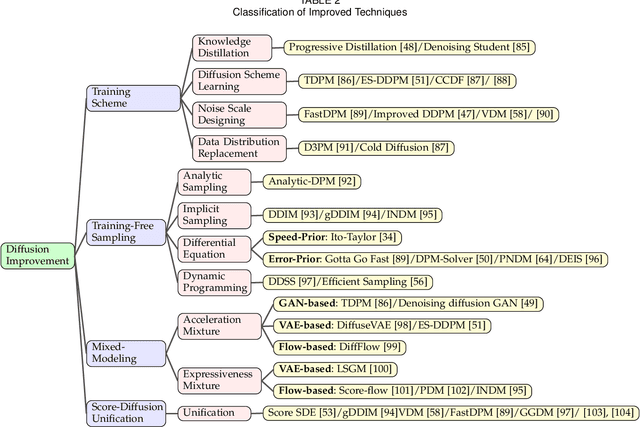
Abstract:Deep learning shows great potential in generation tasks thanks to deep latent representation. Generative models are classes of models that can generate observations randomly with respect to certain implied parameters. Recently, the diffusion Model becomes a raising class of generative models by virtue of its power-generating ability. Nowadays, great achievements have been reached. More applications except for computer vision, speech generation, bioinformatics, and natural language processing are to be explored in this field. However, the diffusion model has its natural drawback of a slow generation process, leading to many enhanced works. This survey makes a summary of the field of the diffusion model. We firstly state the main problem with two landmark works - DDPM and DSM. Then, we present a diverse range of advanced techniques to speed up the diffusion models - training schedule, training-free sampling, mixed-modeling, and score & diffusion unification. Regarding existing models, we also provide a benchmark of FID score, IS, and NLL according to specific NFE. Moreover, applications with diffusion models are introduced including computer vision, sequence modeling, audio, and AI for science. Finally, there is a summarization of this field together with limitations & further directions.
ImDrug: A Benchmark for Deep Imbalanced Learning in AI-aided Drug Discovery
Sep 16, 2022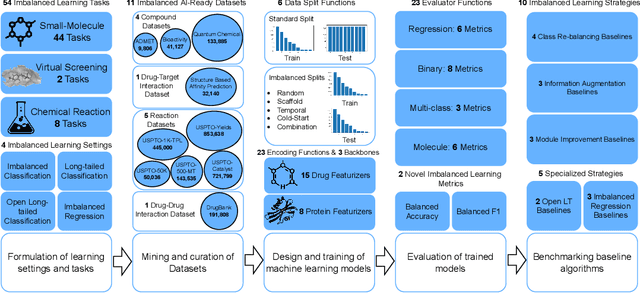

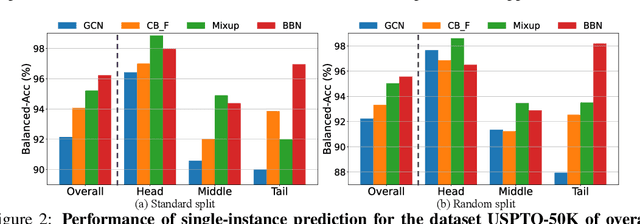
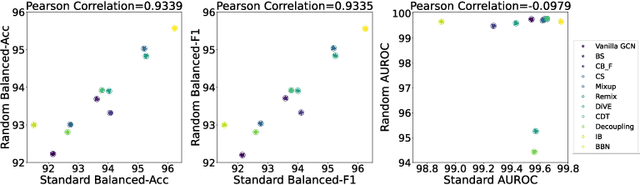
Abstract:The last decade has witnessed a prosperous development of computational methods and dataset curation for AI-aided drug discovery (AIDD). However, real-world pharmaceutical datasets often exhibit highly imbalanced distribution, which is largely overlooked by the current literature but may severely compromise the fairness and generalization of machine learning applications. Motivated by this observation, we introduce ImDrug, a comprehensive benchmark with an open-source Python library which consists of 4 imbalance settings, 11 AI-ready datasets, 54 learning tasks and 16 baseline algorithms tailored for imbalanced learning. It provides an accessible and customizable testbed for problems and solutions spanning a broad spectrum of the drug discovery pipeline such as molecular modeling, drug-target interaction and retrosynthesis. We conduct extensive empirical studies with novel evaluation metrics, to demonstrate that the existing algorithms fall short of solving medicinal and pharmaceutical challenges in the data imbalance scenario. We believe that ImDrug opens up avenues for future research and development, on real-world challenges at the intersection of AIDD and deep imbalanced learning.
Multi-Task Mixture Density Graph Neural Networks for Predicting Cu-based Single-Atom Alloy Catalysts for CO2 Reduction Reaction
Sep 15, 2022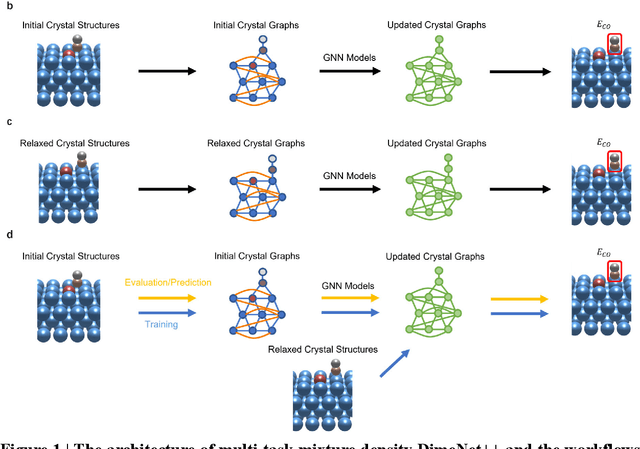
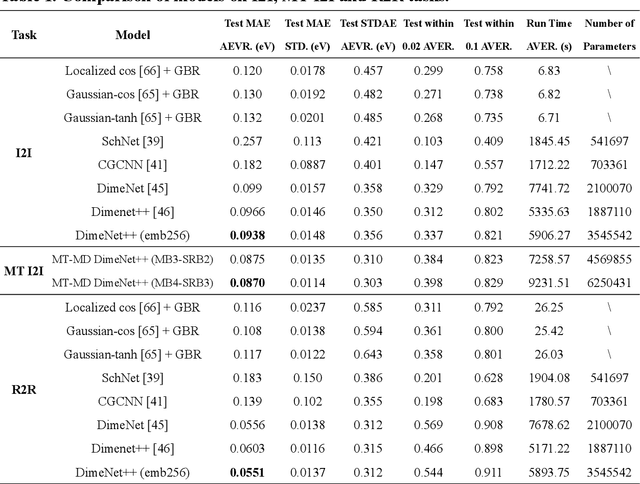
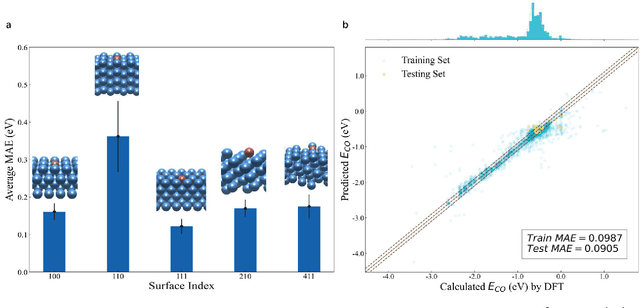
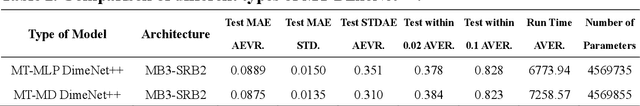
Abstract:Graph neural networks (GNNs) have drawn more and more attention from material scientists and demonstrated a high capacity to establish connections between the structure and properties. However, with only unrelaxed structures provided as input, few GNN models can predict the thermodynamic properties of relaxed configurations with an acceptable level of error. In this work, we develop a multi-task (MT) architecture based on DimeNet++ and mixture density networks to improve the performance of such task. Taking CO adsorption on Cu-based single-atom alloy catalysts as an illustration, we show that our method can reliably estimate CO adsorption energy with a mean absolute error of 0.087 eV from the initial CO adsorption structures without costly first-principles calculations. Further, compared to other state-of-the-art GNN methods, our model exhibits improved generalization ability when predicting catalytic performance of out-of-domain configurations, built with either unseen substrate surfaces or doping species. We show that the proposed MT GNN strategy can facilitate catalyst discovery.
MetaRF: Differentiable Random Forest for Reaction Yield Prediction with a Few Trails
Aug 22, 2022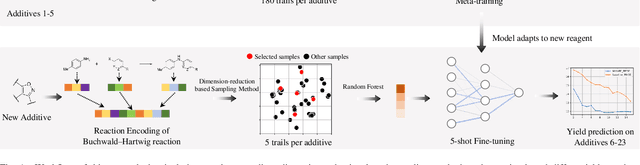
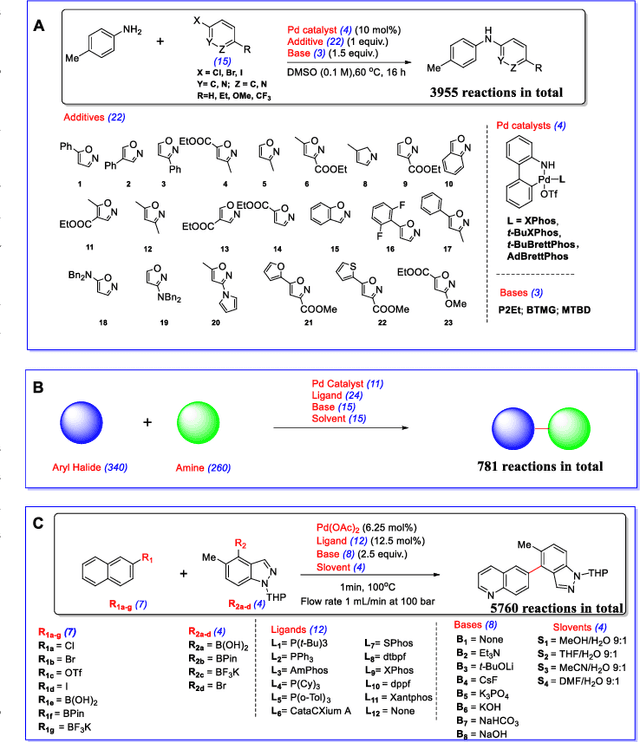
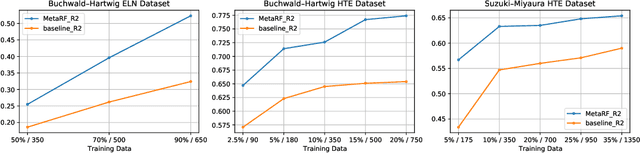
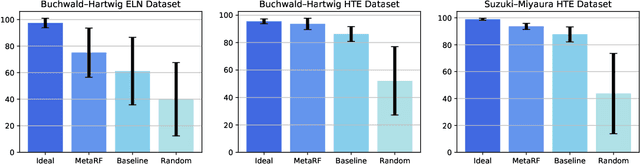
Abstract:Artificial intelligence has deeply revolutionized the field of medicinal chemistry with many impressive applications, but the success of these applications requires a massive amount of training samples with high-quality annotations, which seriously limits the wide usage of data-driven methods. In this paper, we focus on the reaction yield prediction problem, which assists chemists in selecting high-yield reactions in a new chemical space only with a few experimental trials. To attack this challenge, we first put forth MetaRF, an attention-based differentiable random forest model specially designed for the few-shot yield prediction, where the attention weight of a random forest is automatically optimized by the meta-learning framework and can be quickly adapted to predict the performance of new reagents while given a few additional samples. To improve the few-shot learning performance, we further introduce a dimension-reduction based sampling method to determine valuable samples to be experimentally tested and then learned. Our methodology is evaluated on three different datasets and acquires satisfactory performance on few-shot prediction. In high-throughput experimentation (HTE) datasets, the average yield of our methodology's top 10 high-yield reactions is relatively close to the results of ideal yield selection.
Pseudo-label Guided Cross-video Pixel Contrast for Robotic Surgical Scene Segmentation with Limited Annotations
Jul 20, 2022



Abstract:Surgical scene segmentation is fundamentally crucial for prompting cognitive assistance in robotic surgery. However, pixel-wise annotating surgical video in a frame-by-frame manner is expensive and time consuming. To greatly reduce the labeling burden, in this work, we study semi-supervised scene segmentation from robotic surgical video, which is practically essential yet rarely explored before. We consider a clinically suitable annotation situation under the equidistant sampling. We then propose PGV-CL, a novel pseudo-label guided cross-video contrast learning method to boost scene segmentation. It effectively leverages unlabeled data for a trusty and global model regularization that produces more discriminative feature representation. Concretely, for trusty representation learning, we propose to incorporate pseudo labels to instruct the pair selection, obtaining more reliable representation pairs for pixel contrast. Moreover, we expand the representation learning space from previous image-level to cross-video, which can capture the global semantics to benefit the learning process. We extensively evaluate our method on a public robotic surgery dataset EndoVis18 and a public cataract dataset CaDIS. Experimental results demonstrate the effectiveness of our method, consistently outperforming the state-of-the-art semi-supervised methods under different labeling ratios, and even surpassing fully supervised training on EndoVis18 with 10.1% labeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge