Shen Yuan
MoORE: SVD-based Model MoE-ization for Conflict- and Oblivion-Resistant Multi-Task Adaptation
Jun 17, 2025Abstract:Adapting large-scale foundation models in multi-task scenarios often suffers from task conflict and oblivion. To mitigate such issues, we propose a novel ''model MoE-ization'' strategy that leads to a conflict- and oblivion-resistant multi-task adaptation method. Given a weight matrix of a pre-trained model, our method applies SVD to it and introduces a learnable router to adjust its singular values based on tasks and samples. Accordingly, the weight matrix becomes a Mixture of Orthogonal Rank-one Experts (MoORE), in which each expert corresponds to the outer product of a left singular vector and the corresponding right one. We can improve the model capacity by imposing a learnable orthogonal transform on the right singular vectors. Unlike low-rank adaptation (LoRA) and its MoE-driven variants, MoORE guarantees the experts' orthogonality and maintains the column space of the original weight matrix. These two properties make the adapted model resistant to the conflicts among the new tasks and the oblivion of its original tasks, respectively. Experiments on various datasets demonstrate that MoORE outperforms existing multi-task adaptation methods consistently, showing its superiority in terms of conflict- and oblivion-resistance. The code of the experiments is available at https://github.com/DaShenZi721/MoORE.
Towards Better Multi-head Attention via Channel-wise Sample Permutation
Oct 14, 2024



Abstract:Transformer plays a central role in many fundamental deep learning models, e.g., the ViT in computer vision and the BERT and GPT in natural language processing, whose effectiveness is mainly attributed to its multi-head attention (MHA) mechanism. In this study, we propose a simple and novel channel-wise sample permutation (CSP) operator, achieving a new structured MHA with fewer parameters and lower complexity. Given an input matrix, CSP circularly shifts the samples of different channels with various steps and then sorts grouped samples of each channel. This operator is equivalent to implicitly implementing cross-channel attention maps as permutation matrices, which achieves linear complexity and suppresses the risk of rank collapse when representing data. We replace the MHA of some representative models with CSP and test the CSP-based models in several discriminative tasks, including image classification and long sequence analysis. Experiments show that the CSP-based models achieve comparable or better performance with fewer parameters and lower computational costs than the classic Transformer and its state-of-the-art variants. The code is available at https://github.com/DaShenZi721/CSP.
Predicting Polymer Properties Based on Multimodal Multitask Pretraining
Jun 07, 2024Abstract:In the past few decades, polymers, high-molecular-weight compounds formed by bonding numerous identical or similar monomers covalently, have played an essential role in various scientific fields. In this context, accurate prediction of their properties is becoming increasingly crucial. Typically, the properties of a polymer, such as plasticity, conductivity, bio-compatibility, and so on, are highly correlated with its 3D structure. However, current methods for predicting polymer properties heavily rely on information from polymer SMILES sequences (P-SMILES strings) while ignoring crucial 3D structural information, leading to sub-optimal performance. In this work, we propose MMPolymer, a novel multimodal multitask pretraining framework incorporating both polymer 1D sequential information and 3D structural information to enhance downstream polymer property prediction tasks. Besides, to overcome the limited availability of polymer 3D data, we further propose the "Star Substitution" strategy to extract 3D structural information effectively. During pretraining, MMPolymer not only predicts masked tokens and recovers 3D coordinates but also achieves the cross-modal alignment of latent representation. Subsequently, we further fine-tune the pretrained MMPolymer for downstream polymer property prediction tasks in the supervised learning paradigm. Experimental results demonstrate that MMPolymer achieves state-of-the-art performance in various polymer property prediction tasks. Moreover, leveraging the pretrained MMPolymer and using only one modality (either P-SMILES string or 3D conformation) during fine-tuning can also surpass existing polymer property prediction methods, highlighting the exceptional capability of MMPolymer in polymer feature extraction and utilization. Our online platform for polymer property prediction is available at https://app.bohrium.dp.tech/mmpolymer.
Bridging The Gap between Low-rank and Orthogonal Adaptation via Householder Reflection Adaptation
May 24, 2024



Abstract:While following different technical routes, both low-rank and orthogonal adaptation techniques can efficiently adapt large-scale pre-training models in specific tasks or domains based on a small piece of trainable parameters. In this study, we bridge the gap between these two techniques, proposing a simple but effective adaptation method based on Householder reflections. Given a pre-trained model, our method fine-tunes its layers by multiplying each frozen weight matrix with an orthogonal matrix constructed by a chain of learnable Householder reflections (HRs). This HR-based orthogonal fine-tuning is equivalent to an adaptive low-rank adaptation. Moreover, we show that the orthogonality of the reflection planes corresponding to the HRs impacts the model capacity and regularity. The analysis motivates us to regularize the orthogonality of the HRs, leading to different implementations of the proposed Householder reflection adaptation (HRA) method. Compared with state-of-the-art methods, HRA achieves superior performance with fewer learnable parameters when adapting large language models and conditional image generators. The code is available at https://github.com/DaShenZi721/HRA
Sliceformer: Make Multi-head Attention as Simple as Sorting in Discriminative Tasks
Oct 26, 2023



Abstract:As one of the most popular neural network modules, Transformer plays a central role in many fundamental deep learning models, e.g., the ViT in computer vision and the BERT and GPT in natural language processing. The effectiveness of the Transformer is often attributed to its multi-head attention (MHA) mechanism. In this study, we discuss the limitations of MHA, including the high computational complexity due to its ``query-key-value'' architecture and the numerical issue caused by its softmax operation. Considering the above problems and the recent development tendency of the attention layer, we propose an effective and efficient surrogate of the Transformer, called Sliceformer. Our Sliceformer replaces the classic MHA mechanism with an extremely simple ``slicing-sorting'' operation, i.e., projecting inputs linearly to a latent space and sorting them along different feature dimensions (or equivalently, called channels). For each feature dimension, the sorting operation implicitly generates an implicit attention map with sparse, full-rank, and doubly-stochastic structures. We consider different implementations of the slicing-sorting operation and analyze their impacts on the Sliceformer. We test the Sliceformer in the Long-Range Arena benchmark, image classification, text classification, and molecular property prediction, demonstrating its advantage in computational complexity and universal effectiveness in discriminative tasks. Our Sliceformer achieves comparable or better performance with lower memory cost and faster speed than the Transformer and its variants. Moreover, the experimental results reveal that applying our Sliceformer can empirically suppress the risk of mode collapse when representing data. The code is available at \url{https://github.com/SDS-Lab/sliceformer}.
ImDrug: A Benchmark for Deep Imbalanced Learning in AI-aided Drug Discovery
Sep 16, 2022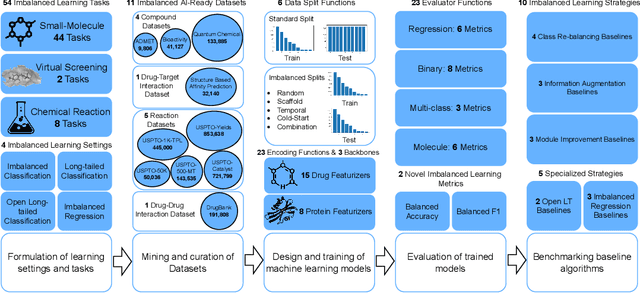

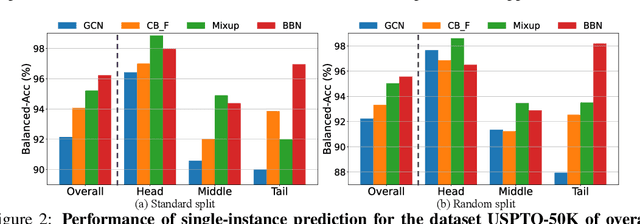
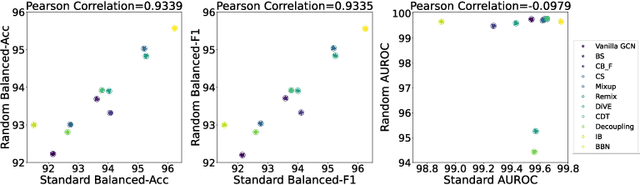
Abstract:The last decade has witnessed a prosperous development of computational methods and dataset curation for AI-aided drug discovery (AIDD). However, real-world pharmaceutical datasets often exhibit highly imbalanced distribution, which is largely overlooked by the current literature but may severely compromise the fairness and generalization of machine learning applications. Motivated by this observation, we introduce ImDrug, a comprehensive benchmark with an open-source Python library which consists of 4 imbalance settings, 11 AI-ready datasets, 54 learning tasks and 16 baseline algorithms tailored for imbalanced learning. It provides an accessible and customizable testbed for problems and solutions spanning a broad spectrum of the drug discovery pipeline such as molecular modeling, drug-target interaction and retrosynthesis. We conduct extensive empirical studies with novel evaluation metrics, to demonstrate that the existing algorithms fall short of solving medicinal and pharmaceutical challenges in the data imbalance scenario. We believe that ImDrug opens up avenues for future research and development, on real-world challenges at the intersection of AIDD and deep imbalanced learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge