Guang Yang
Shanghai Key Laboratory of Magnetic Resonance, School of Physics and Electronic Science, East China Normal University, Shanghai, China, Institute of Magnetic Resonance and Molecular Imaging in Medicine, East China Normal University, Shanghai, China
Advancing oncology with federated learning: transcending boundaries in breast, lung, and prostate cancer. A systematic review
Aug 08, 2024Abstract:Federated Learning (FL) has emerged as a promising solution to address the limitations of centralised machine learning (ML) in oncology, particularly in overcoming privacy concerns and harnessing the power of diverse, multi-center data. This systematic review synthesises current knowledge on the state-of-the-art FL in oncology, focusing on breast, lung, and prostate cancer. Distinct from previous surveys, our comprehensive review critically evaluates the real-world implementation and impact of FL on cancer care, demonstrating its effectiveness in enhancing ML generalisability, performance and data privacy in clinical settings and data. We evaluated state-of-the-art advances in FL, demonstrating its growing adoption amid tightening data privacy regulations. FL outperformed centralised ML in 15 out of the 25 studies reviewed, spanning diverse ML models and clinical applications, and facilitating integration of multi-modal information for precision medicine. Despite the current challenges identified in reproducibility, standardisation and methodology across studies, the demonstrable benefits of FL in harnessing real-world data and addressing clinical needs highlight its significant potential for advancing cancer research. We propose that future research should focus on addressing these limitations and investigating further advanced FL methods, to fully harness data diversity and realise the transformative power of cutting-edge FL in cancer care.
CIResDiff: A Clinically-Informed Residual Diffusion Model for Predicting Idiopathic Pulmonary Fibrosis Progression
Aug 05, 2024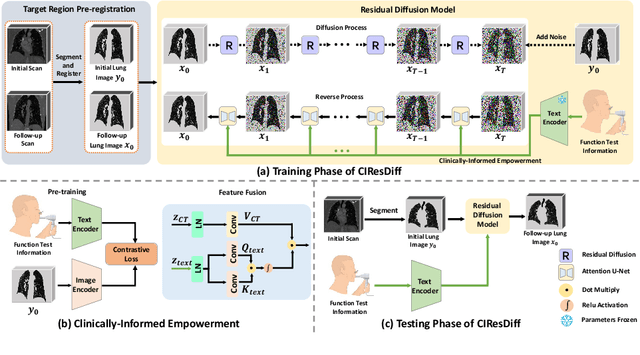
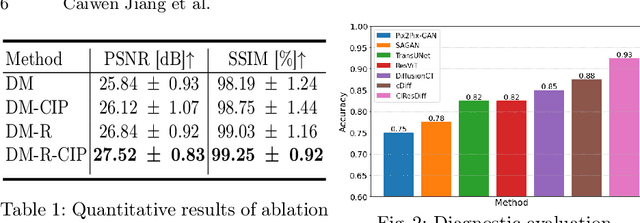
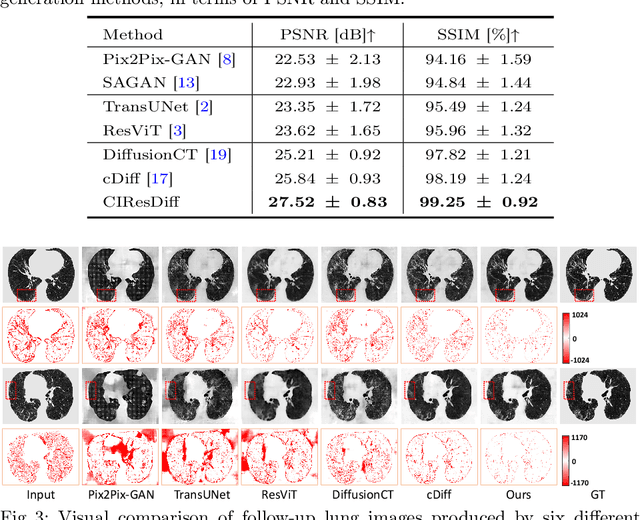
Abstract:The progression of Idiopathic Pulmonary Fibrosis (IPF) significantly correlates with higher patient mortality rates. Early detection of IPF progression is critical for initiating timely treatment, which can effectively slow down the advancement of the disease. However, the current clinical criteria define disease progression requiring two CT scans with a one-year interval, presenting a dilemma: a disease progression is identified only after the disease has already progressed. To this end, in this paper, we develop a novel diffusion model to accurately predict the progression of IPF by generating patient's follow-up CT scan from the initial CT scan. Specifically, from the clinical prior knowledge, we tailor improvements to the traditional diffusion model and propose a Clinically-Informed Residual Diffusion model, called CIResDiff. The key innovations of CIResDiff include 1) performing the target region pre-registration to align the lung regions of two CT scans at different time points for reducing the generation difficulty, 2) adopting the residual diffusion instead of traditional diffusion to enable the model focus more on differences (i.e., lesions) between the two CT scans rather than the largely identical anatomical content, and 3) designing the clinically-informed process based on CLIP technology to integrate lung function information which is highly relevant to diagnosis into the reverse process for assisting generation. Extensive experiments on clinical data demonstrate that our approach can outperform state-of-the-art methods and effectively predict the progression of IPF.
A dual-task mutual learning framework for predicting post-thrombectomy cerebral hemorrhage
Aug 01, 2024Abstract:Ischemic stroke is a severe condition caused by the blockage of brain blood vessels, and can lead to the death of brain tissue due to oxygen deprivation. Thrombectomy has become a common treatment choice for ischemic stroke due to its immediate effectiveness. But, it carries the risk of postoperative cerebral hemorrhage. Clinically, multiple CT scans within 0-72 hours post-surgery are used to monitor for hemorrhage. However, this approach exposes radiation dose to patients, and may delay the detection of cerebral hemorrhage. To address this dilemma, we propose a novel prediction framework for measuring postoperative cerebral hemorrhage using only the patient's initial CT scan. Specifically, we introduce a dual-task mutual learning framework to takes the initial CT scan as input and simultaneously estimates both the follow-up CT scan and prognostic label to predict the occurrence of postoperative cerebral hemorrhage. Our proposed framework incorporates two attention mechanisms, i.e., self-attention and interactive attention. Specifically, the self-attention mechanism allows the model to focus more on high-density areas in the image, which are critical for diagnosis (i.e., potential hemorrhage areas). The interactive attention mechanism further models the dependencies between the interrelated generation and classification tasks, enabling both tasks to perform better than the case when conducted individually. Validated on clinical data, our method can generate follow-up CT scans better than state-of-the-art methods, and achieves an accuracy of 86.37% in predicting follow-up prognostic labels. Thus, our work thus contributes to the timely screening of post-thrombectomy cerebral hemorrhage, and could significantly reform the clinical process of thrombectomy and other similar operations related to stroke.
Artificial Immunofluorescence in a Flash: Rapid Synthetic Imaging from Brightfield Through Residual Diffusion
Jul 25, 2024



Abstract:Immunofluorescent (IF) imaging is crucial for visualizing biomarker expressions, cell morphology and assessing the effects of drug treatments on sub-cellular components. IF imaging needs extra staining process and often requiring cell fixation, therefore it may also introduce artefects and alter endogenouous cell morphology. Some IF stains are expensive or not readily available hence hindering experiments. Recent diffusion models, which synthesise high-fidelity IF images from easy-to-acquire brightfield (BF) images, offer a promising solution but are hindered by training instability and slow inference times due to the noise diffusion process. This paper presents a novel method for the conditional synthesis of IF images directly from BF images along with cell segmentation masks. Our approach employs a Residual Diffusion process that enhances stability and significantly reduces inference time. We performed a critical evaluation against other image-to-image synthesis models, including UNets, GANs, and advanced diffusion models. Our model demonstrates significant improvements in image quality (p<0.05 in MSE, PSNR, and SSIM), inference speed (26 times faster than competing diffusion models), and accurate segmentation results for both nuclei and cell bodies (0.77 and 0.63 mean IOU for nuclei and cell true positives, respectively). This paper is a substantial advancement in the field, providing robust and efficient tools for cell image analysis.
Differentiable Voxelization and Mesh Morphing
Jul 15, 2024



Abstract:In this paper, we propose the differentiable voxelization of 3D meshes via the winding number and solid angles. The proposed approach achieves fast, flexible, and accurate voxelization of 3D meshes, admitting the computation of gradients with respect to the input mesh and GPU acceleration. We further demonstrate the application of the proposed voxelization in mesh morphing, where the voxelized mesh is deformed by a neural network. The proposed method is evaluated on the ShapeNet dataset and achieves state-of-the-art performance in terms of both accuracy and efficiency.
LGRNet: Local-Global Reciprocal Network for Uterine Fibroid Segmentation in Ultrasound Videos
Jul 08, 2024



Abstract:Regular screening and early discovery of uterine fibroid are crucial for preventing potential malignant transformations and ensuring timely, life-saving interventions. To this end, we collect and annotate the first ultrasound video dataset with 100 videos for uterine fibroid segmentation (UFUV). We also present Local-Global Reciprocal Network (LGRNet) to efficiently and effectively propagate the long-term temporal context which is crucial to help distinguish between uninformative noisy surrounding tissues and target lesion regions. Specifically, the Cyclic Neighborhood Propagation (CNP) is introduced to propagate the inter-frame local temporal context in a cyclic manner. Moreover, to aggregate global temporal context, we first condense each frame into a set of frame bottleneck queries and devise Hilbert Selective Scan (HilbertSS) to both efficiently path connect each frame and preserve the locality bias. A distribute layer is then utilized to disseminate back the global context for reciprocal refinement. Extensive experiments on UFUV and three public Video Polyp Segmentation (VPS) datasets demonstrate consistent improvements compared to state-of-the-art segmentation methods, indicating the effectiveness and versatility of LGRNet. Code, checkpoints, and dataset are available at https://github.com/bio-mlhui/LGRNet
Probing Perfection: The Relentless Art of Meddling for Pulmonary Airway Segmentation from HRCT via a Human-AI Collaboration Based Active Learning Method
Jul 03, 2024Abstract:In pulmonary tracheal segmentation, the scarcity of annotated data is a prevalent issue in medical segmentation. Additionally, Deep Learning (DL) methods face challenges: the opacity of 'black box' models and the need for performance enhancement. Our Human-Computer Interaction (HCI) based models (RS_UNet, LC_UNet, UUNet, and WD_UNet) address these challenges by combining diverse query strategies with various DL models. We train four HCI models and repeat these steps: (1) Query Strategy: The HCI models select samples that provide the most additional representative information when labeled in each iteration and identify unlabeled samples with the greatest predictive disparity using Wasserstein Distance, Least Confidence, Entropy Sampling, and Random Sampling. (2) Central line correction: Selected samples are used for expert correction of system-generated tracheal central lines in each training round. (3) Update training dataset: Experts update the training dataset after each DL model's training epoch, enhancing the trustworthiness and performance of the models. (4) Model training: The HCI model is trained using the updated dataset and an enhanced UNet version. Experimental results confirm the effectiveness of these HCI-based approaches, showing that WD-UNet, LC-UNet, UUNet, and RS-UNet achieve comparable or superior performance to state-of-the-art DL models. Notably, WD-UNet achieves this with only 15%-35% of the training data, reducing physician annotation time by 65%-85%.
CMRxRecon2024: A Multi-Modality, Multi-View K-Space Dataset Boosting Universal Machine Learning for Accelerated Cardiac MRI
Jun 27, 2024



Abstract:Cardiac magnetic resonance imaging (MRI) has emerged as a clinically gold-standard technique for diagnosing cardiac diseases, thanks to its ability to provide diverse information with multiple modalities and anatomical views. Accelerated cardiac MRI is highly expected to achieve time-efficient and patient-friendly imaging, and then advanced image reconstruction approaches are required to recover high-quality, clinically interpretable images from undersampled measurements. However, the lack of publicly available cardiac MRI k-space dataset in terms of both quantity and diversity has severely hindered substantial technological progress, particularly for data-driven artificial intelligence. Here, we provide a standardized, diverse, and high-quality CMRxRecon2024 dataset to facilitate the technical development, fair evaluation, and clinical transfer of cardiac MRI reconstruction approaches, towards promoting the universal frameworks that enable fast and robust reconstructions across different cardiac MRI protocols in clinical practice. To the best of our knowledge, the CMRxRecon2024 dataset is the largest and most diverse publicly available cardiac k-space dataset. It is acquired from 330 healthy volunteers, covering commonly used modalities, anatomical views, and acquisition trajectories in clinical cardiac MRI workflows. Besides, an open platform with tutorials, benchmarks, and data processing tools is provided to facilitate data usage, advanced method development, and fair performance evaluation.
SimsChat: A Customisable Persona-Driven Role-Playing Agent
Jun 25, 2024Abstract:Large Language Models (LLMs) possess the remarkable capability to understand human instructions and generate high-quality text, enabling them to act as agents that simulate human behaviours. This capability allows LLMs to emulate human beings in a more advanced manner, beyond merely replicating simple human behaviours. However, there is a lack of exploring into leveraging LLMs to craft characters from several aspects. In this work, we introduce the Customisable Conversation Agent Framework, which employs LLMs to simulate real-world characters that can be freely customised according to different user preferences. The customisable framework is helpful for designing customisable characters and role-playing agents according to human's preferences. We first propose the SimsConv dataset, which comprises 68 different customised characters, 1,360 multi-turn role-playing dialogues, and encompasses 13,971 interaction dialogues in total. The characters are created from several real-world elements, such as career, aspiration, trait, and skill. Building on these foundations, we present SimsChat, a freely customisable role-playing agent. It incorporates different real-world scenes and topic-specific character interaction dialogues, simulating characters' life experiences in various scenarios and topic-specific interactions with specific emotions. Experimental results show that our proposed framework achieves desirable performance and provides helpful guideline for building better simulacra of human beings in the future. Our data and code are available at https://github.com/Bernard-Yang/SimsChat.
Diff3Dformer: Leveraging Slice Sequence Diffusion for Enhanced 3D CT Classification with Transformer Networks
Jun 24, 2024Abstract:The manifestation of symptoms associated with lung diseases can vary in different depths for individual patients, highlighting the significance of 3D information in CT scans for medical image classification. While Vision Transformer has shown superior performance over convolutional neural networks in image classification tasks, their effectiveness is often demonstrated on sufficiently large 2D datasets and they easily encounter overfitting issues on small medical image datasets. To address this limitation, we propose a Diffusion-based 3D Vision Transformer (Diff3Dformer), which utilizes the latent space of the Diffusion model to form the slice sequence for 3D analysis and incorporates clustering attention into ViT to aggregate repetitive information within 3D CT scans, thereby harnessing the power of the advanced transformer in 3D classification tasks on small datasets. Our method exhibits improved performance on two different scales of small datasets of 3D lung CT scans, surpassing the state of the art 3D methods and other transformer-based approaches that emerged during the COVID-19 pandemic, demonstrating its robust and superior performance across different scales of data. Experimental results underscore the superiority of our proposed method, indicating its potential for enhancing medical image classification tasks in real-world scenarios.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge