Xiaodan Xing
Revisiting Medical Image Retrieval via Knowledge Consolidation
Mar 12, 2025Abstract:As artificial intelligence and digital medicine increasingly permeate healthcare systems, robust governance frameworks are essential to ensure ethical, secure, and effective implementation. In this context, medical image retrieval becomes a critical component of clinical data management, playing a vital role in decision-making and safeguarding patient information. Existing methods usually learn hash functions using bottleneck features, which fail to produce representative hash codes from blended embeddings. Although contrastive hashing has shown superior performance, current approaches often treat image retrieval as a classification task, using category labels to create positive/negative pairs. Moreover, many methods fail to address the out-of-distribution (OOD) issue when models encounter external OOD queries or adversarial attacks. In this work, we propose a novel method to consolidate knowledge of hierarchical features and optimisation functions. We formulate the knowledge consolidation by introducing Depth-aware Representation Fusion (DaRF) and Structure-aware Contrastive Hashing (SCH). DaRF adaptively integrates shallow and deep representations into blended features, and SCH incorporates image fingerprints to enhance the adaptability of positive/negative pairings. These blended features further facilitate OOD detection and content-based recommendation, contributing to a secure AI-driven healthcare environment. Moreover, we present a content-guided ranking to improve the robustness and reproducibility of retrieval results. Our comprehensive assessments demonstrate that the proposed method could effectively recognise OOD samples and significantly outperform existing approaches in medical image retrieval (p<0.05). In particular, our method achieves a 5.6-38.9% improvement in mean Average Precision on the anatomical radiology dataset.
Anatomy-Guided Radiology Report Generation with Pathology-Aware Regional Prompts
Nov 16, 2024



Abstract:Radiology reporting generative AI holds significant potential to alleviate clinical workloads and streamline medical care. However, achieving high clinical accuracy is challenging, as radiological images often feature subtle lesions and intricate structures. Existing systems often fall short, largely due to their reliance on fixed size, patch-level image features and insufficient incorporation of pathological information. This can result in the neglect of such subtle patterns and inconsistent descriptions of crucial pathologies. To address these challenges, we propose an innovative approach that leverages pathology-aware regional prompts to explicitly integrate anatomical and pathological information of various scales, significantly enhancing the precision and clinical relevance of generated reports. We develop an anatomical region detector that extracts features from distinct anatomical areas, coupled with a novel multi-label lesion detector that identifies global pathologies. Our approach emulates the diagnostic process of radiologists, producing clinically accurate reports with comprehensive diagnostic capabilities. Experimental results show that our model outperforms previous state-of-the-art methods on most natural language generation and clinical efficacy metrics, with formal expert evaluations affirming its potential to enhance radiology practice.
Deep Generative Models Unveil Patterns in Medical Images Through Vision-Language Conditioning
Oct 17, 2024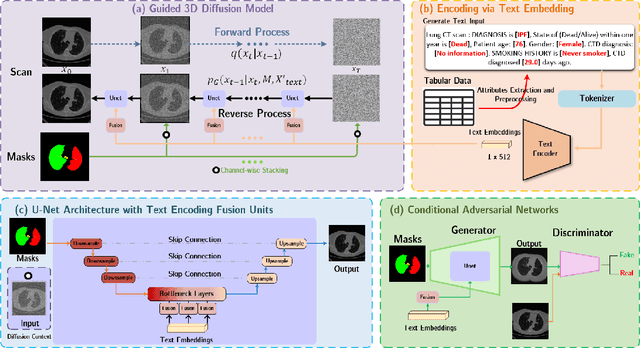
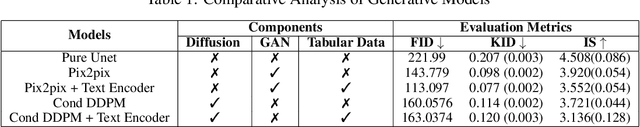
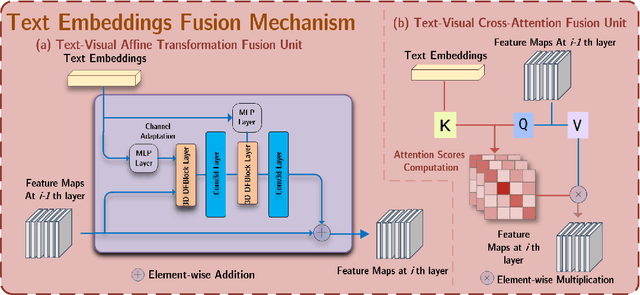
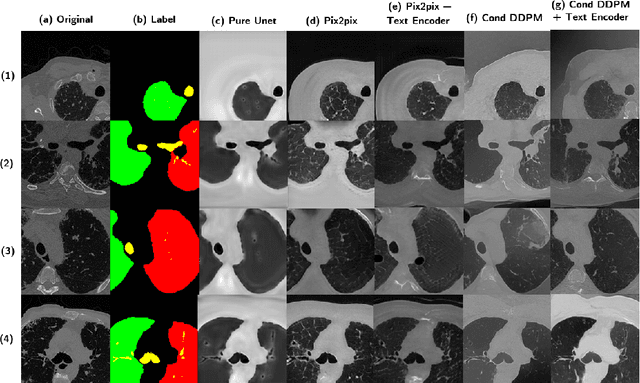
Abstract:Deep generative models have significantly advanced medical imaging analysis by enhancing dataset size and quality. Beyond mere data augmentation, our research in this paper highlights an additional, significant capacity of deep generative models: their ability to reveal and demonstrate patterns in medical images. We employ a generative structure with hybrid conditions, combining clinical data and segmentation masks to guide the image synthesis process. Furthermore, we innovatively transformed the tabular clinical data into textual descriptions. This approach simplifies the handling of missing values and also enables us to leverage large pre-trained vision-language models that investigate the relations between independent clinical entries and comprehend general terms, such as gender and smoking status. Our approach differs from and presents a more challenging task than traditional medical report-guided synthesis due to the less visual correlation of our clinical information with the images. To overcome this, we introduce a text-visual embedding mechanism that strengthens the conditions, ensuring the network effectively utilizes the provided information. Our pipeline is generalizable to both GAN-based and diffusion models. Experiments on chest CT, particularly focusing on the smoking status, demonstrated a consistent intensity shift in the lungs which is in agreement with clinical observations, indicating the effectiveness of our method in capturing and visualizing the impact of specific attributes on medical image patterns. Our methods offer a new avenue for the early detection and precise visualization of complex clinical conditions with deep generative models. All codes are https://github.com/junzhin/DGM-VLC.
Preserving Cardiac Integrity: A Topology-Infused Approach to Whole Heart Segmentation
Oct 14, 2024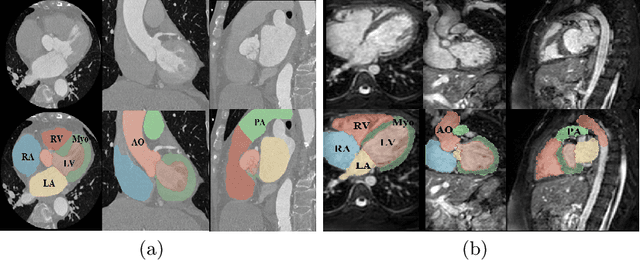
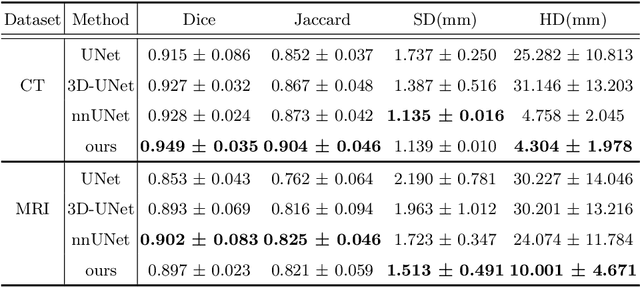
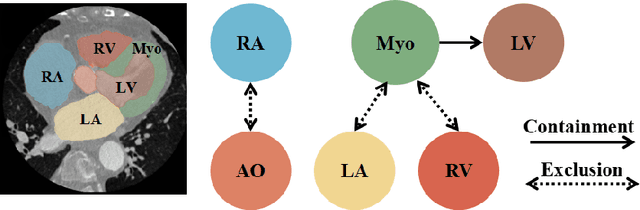
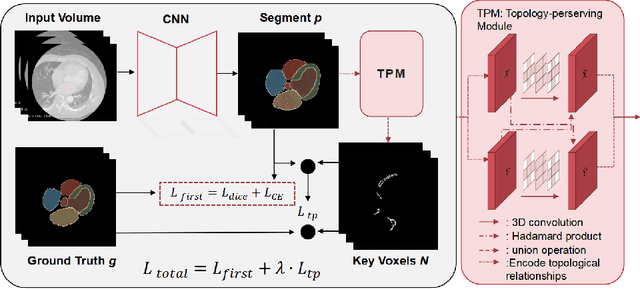
Abstract:Whole heart segmentation (WHS) supports cardiovascular disease (CVD) diagnosis, disease monitoring, treatment planning, and prognosis. Deep learning has become the most widely used method for WHS applications in recent years. However, segmentation of whole-heart structures faces numerous challenges including heart shape variability during the cardiac cycle, clinical artifacts like motion and poor contrast-to-noise ratio, domain shifts in multi-center data, and the distinct modalities of CT and MRI. To address these limitations and improve segmentation quality, this paper introduces a new topology-preserving module that is integrated into deep neural networks. The implementation achieves anatomically plausible segmentation by using learned topology-preserving fields, which are based entirely on 3D convolution and are therefore very effective for 3D voxel data. We incorporate natural constraints between structures into the end-to-end training and enrich the feature representation of the neural network. The effectiveness of the proposed method is validated on an open-source medical heart dataset, specifically using the WHS++ data. The results demonstrate that the architecture performs exceptionally well, achieving a Dice coefficient of 0.939 during testing. This indicates full topology preservation for individual structures and significantly outperforms other baselines in preserving the overall scene topology.
MONICA: Benchmarking on Long-tailed Medical Image Classification
Oct 02, 2024



Abstract:Long-tailed learning is considered to be an extremely challenging problem in data imbalance learning. It aims to train well-generalized models from a large number of images that follow a long-tailed class distribution. In the medical field, many diagnostic imaging exams such as dermoscopy and chest radiography yield a long-tailed distribution of complex clinical findings. Recently, long-tailed learning in medical image analysis has garnered significant attention. However, the field currently lacks a unified, strictly formulated, and comprehensive benchmark, which often leads to unfair comparisons and inconclusive results. To help the community improve the evaluation and advance, we build a unified, well-structured codebase called Medical OpeN-source Long-taIled ClassifiCAtion (MONICA), which implements over 30 methods developed in relevant fields and evaluated on 12 long-tailed medical datasets covering 6 medical domains. Our work provides valuable practical guidance and insights for the field, offering detailed analysis and discussion on the effectiveness of individual components within the inbuilt state-of-the-art methodologies. We hope this codebase serves as a comprehensive and reproducible benchmark, encouraging further advancements in long-tailed medical image learning. The codebase is publicly available on https://github.com/PyJulie/MONICA.
Coupling AI and Citizen Science in Creation of Enhanced Training Dataset for Medical Image Segmentation
Sep 04, 2024



Abstract:Recent advancements in medical imaging and artificial intelligence (AI) have greatly enhanced diagnostic capabilities, but the development of effective deep learning (DL) models is still constrained by the lack of high-quality annotated datasets. The traditional manual annotation process by medical experts is time- and resource-intensive, limiting the scalability of these datasets. In this work, we introduce a robust and versatile framework that combines AI and crowdsourcing to improve both the quality and quantity of medical image datasets across different modalities. Our approach utilises a user-friendly online platform that enables a diverse group of crowd annotators to label medical images efficiently. By integrating the MedSAM segmentation AI with this platform, we accelerate the annotation process while maintaining expert-level quality through an algorithm that merges crowd-labelled images. Additionally, we employ pix2pixGAN, a generative AI model, to expand the training dataset with synthetic images that capture realistic morphological features. These methods are combined into a cohesive framework designed to produce an enhanced dataset, which can serve as a universal pre-processing pipeline to boost the training of any medical deep learning segmentation model. Our results demonstrate that this framework significantly improves model performance, especially when training data is limited.
Beyond the Hype: A dispassionate look at vision-language models in medical scenario
Aug 16, 2024



Abstract:Recent advancements in Large Vision-Language Models (LVLMs) have demonstrated remarkable capabilities across diverse tasks, garnering significant attention in AI communities. However, their performance and reliability in specialized domains such as medicine remain insufficiently assessed. In particular, most assessments over-concentrate in evaluating VLMs based on simple Visual Question Answering (VQA) on multi-modality data, while ignoring the in-depth characteristic of LVLMs. In this study, we introduce RadVUQA, a novel Radiological Visual Understanding and Question Answering benchmark, to comprehensively evaluate existing LVLMs. RadVUQA mainly validates LVLMs across five dimensions: 1) Anatomical understanding, assessing the models' ability to visually identify biological structures; 2) Multimodal comprehension, which involves the capability of interpreting linguistic and visual instructions to produce desired outcomes; 3) Quantitative and spatial reasoning, evaluating the models' spatial awareness and proficiency in combining quantitative analysis with visual and linguistic information; 4) Physiological knowledge, measuring the models' capability to comprehend functions and mechanisms of organs and systems; and 5) Robustness, which assesses the models' capabilities against unharmonised and synthetic data. The results indicate that both generalized LVLMs and medical-specific LVLMs have critical deficiencies with weak multimodal comprehension and quantitative reasoning capabilities. Our findings reveal the large gap between existing LVLMs and clinicians, highlighting the urgent need for more robust and intelligent LVLMs. The code and dataset will be available after the acceptance of this paper.
CIResDiff: A Clinically-Informed Residual Diffusion Model for Predicting Idiopathic Pulmonary Fibrosis Progression
Aug 05, 2024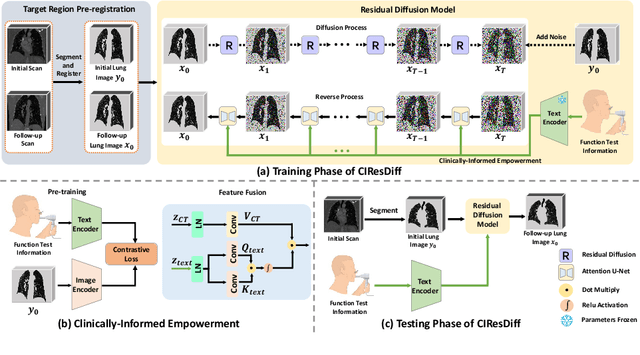
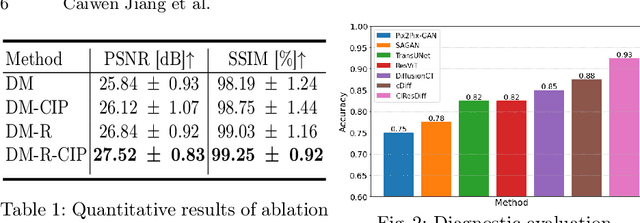
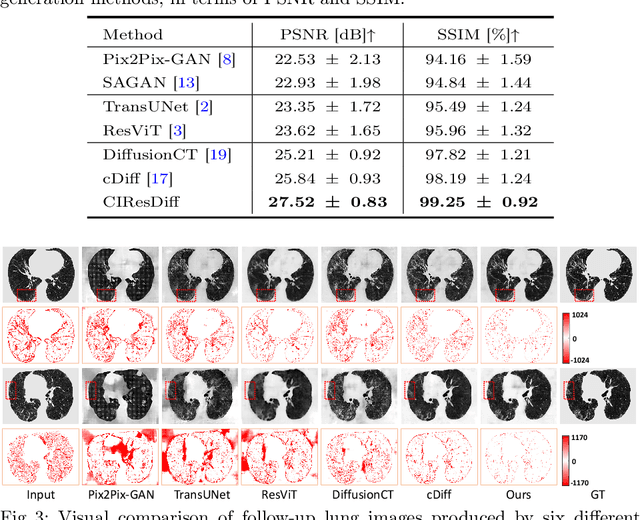
Abstract:The progression of Idiopathic Pulmonary Fibrosis (IPF) significantly correlates with higher patient mortality rates. Early detection of IPF progression is critical for initiating timely treatment, which can effectively slow down the advancement of the disease. However, the current clinical criteria define disease progression requiring two CT scans with a one-year interval, presenting a dilemma: a disease progression is identified only after the disease has already progressed. To this end, in this paper, we develop a novel diffusion model to accurately predict the progression of IPF by generating patient's follow-up CT scan from the initial CT scan. Specifically, from the clinical prior knowledge, we tailor improvements to the traditional diffusion model and propose a Clinically-Informed Residual Diffusion model, called CIResDiff. The key innovations of CIResDiff include 1) performing the target region pre-registration to align the lung regions of two CT scans at different time points for reducing the generation difficulty, 2) adopting the residual diffusion instead of traditional diffusion to enable the model focus more on differences (i.e., lesions) between the two CT scans rather than the largely identical anatomical content, and 3) designing the clinically-informed process based on CLIP technology to integrate lung function information which is highly relevant to diagnosis into the reverse process for assisting generation. Extensive experiments on clinical data demonstrate that our approach can outperform state-of-the-art methods and effectively predict the progression of IPF.
A dual-task mutual learning framework for predicting post-thrombectomy cerebral hemorrhage
Aug 01, 2024Abstract:Ischemic stroke is a severe condition caused by the blockage of brain blood vessels, and can lead to the death of brain tissue due to oxygen deprivation. Thrombectomy has become a common treatment choice for ischemic stroke due to its immediate effectiveness. But, it carries the risk of postoperative cerebral hemorrhage. Clinically, multiple CT scans within 0-72 hours post-surgery are used to monitor for hemorrhage. However, this approach exposes radiation dose to patients, and may delay the detection of cerebral hemorrhage. To address this dilemma, we propose a novel prediction framework for measuring postoperative cerebral hemorrhage using only the patient's initial CT scan. Specifically, we introduce a dual-task mutual learning framework to takes the initial CT scan as input and simultaneously estimates both the follow-up CT scan and prognostic label to predict the occurrence of postoperative cerebral hemorrhage. Our proposed framework incorporates two attention mechanisms, i.e., self-attention and interactive attention. Specifically, the self-attention mechanism allows the model to focus more on high-density areas in the image, which are critical for diagnosis (i.e., potential hemorrhage areas). The interactive attention mechanism further models the dependencies between the interrelated generation and classification tasks, enabling both tasks to perform better than the case when conducted individually. Validated on clinical data, our method can generate follow-up CT scans better than state-of-the-art methods, and achieves an accuracy of 86.37% in predicting follow-up prognostic labels. Thus, our work thus contributes to the timely screening of post-thrombectomy cerebral hemorrhage, and could significantly reform the clinical process of thrombectomy and other similar operations related to stroke.
Artificial Immunofluorescence in a Flash: Rapid Synthetic Imaging from Brightfield Through Residual Diffusion
Jul 25, 2024



Abstract:Immunofluorescent (IF) imaging is crucial for visualizing biomarker expressions, cell morphology and assessing the effects of drug treatments on sub-cellular components. IF imaging needs extra staining process and often requiring cell fixation, therefore it may also introduce artefects and alter endogenouous cell morphology. Some IF stains are expensive or not readily available hence hindering experiments. Recent diffusion models, which synthesise high-fidelity IF images from easy-to-acquire brightfield (BF) images, offer a promising solution but are hindered by training instability and slow inference times due to the noise diffusion process. This paper presents a novel method for the conditional synthesis of IF images directly from BF images along with cell segmentation masks. Our approach employs a Residual Diffusion process that enhances stability and significantly reduces inference time. We performed a critical evaluation against other image-to-image synthesis models, including UNets, GANs, and advanced diffusion models. Our model demonstrates significant improvements in image quality (p<0.05 in MSE, PSNR, and SSIM), inference speed (26 times faster than competing diffusion models), and accurate segmentation results for both nuclei and cell bodies (0.77 and 0.63 mean IOU for nuclei and cell true positives, respectively). This paper is a substantial advancement in the field, providing robust and efficient tools for cell image analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge