Chunhui Lin
Dolphin: Document Image Parsing via Heterogeneous Anchor Prompting
May 20, 2025Abstract:Document image parsing is challenging due to its complexly intertwined elements such as text paragraphs, figures, formulas, and tables. Current approaches either assemble specialized expert models or directly generate page-level content autoregressively, facing integration overhead, efficiency bottlenecks, and layout structure degradation despite their decent performance. To address these limitations, we present \textit{Dolphin} (\textit{\textbf{Do}cument Image \textbf{P}arsing via \textbf{H}eterogeneous Anchor Prompt\textbf{in}g}), a novel multimodal document image parsing model following an analyze-then-parse paradigm. In the first stage, Dolphin generates a sequence of layout elements in reading order. These heterogeneous elements, serving as anchors and coupled with task-specific prompts, are fed back to Dolphin for parallel content parsing in the second stage. To train Dolphin, we construct a large-scale dataset of over 30 million samples, covering multi-granularity parsing tasks. Through comprehensive evaluations on both prevalent benchmarks and self-constructed ones, Dolphin achieves state-of-the-art performance across diverse page-level and element-level settings, while ensuring superior efficiency through its lightweight architecture and parallel parsing mechanism. The code and pre-trained models are publicly available at https://github.com/ByteDance/Dolphin
OCRBench v2: An Improved Benchmark for Evaluating Large Multimodal Models on Visual Text Localization and Reasoning
Dec 31, 2024



Abstract:Scoring the Optical Character Recognition (OCR) capabilities of Large Multimodal Models (LMMs) has witnessed growing interest recently. Existing benchmarks have highlighted the impressive performance of LMMs in text recognition; however, their abilities on certain challenging tasks, such as text localization, handwritten content extraction, and logical reasoning, remain underexplored. To bridge this gap, we introduce OCRBench v2, a large-scale bilingual text-centric benchmark with currently the most comprehensive set of tasks (4x more tasks than the previous multi-scene benchmark OCRBench), the widest coverage of scenarios (31 diverse scenarios including street scene, receipt, formula, diagram, and so on), and thorough evaluation metrics, with a total of 10,000 human-verified question-answering pairs and a high proportion of difficult samples. After carefully benchmarking state-of-the-art LMMs on OCRBench v2, we find that 20 out of 22 LMMs score below 50 (100 in total) and suffer from five-type limitations, including less frequently encountered text recognition, fine-grained perception, layout perception, complex element parsing, and logical reasoning. The benchmark and evaluation scripts are available at https://github.com/Yuliang-liu/MultimodalOCR.
Harmonizing Visual Text Comprehension and Generation
Jul 23, 2024



Abstract:In this work, we present TextHarmony, a unified and versatile multimodal generative model proficient in comprehending and generating visual text. Simultaneously generating images and texts typically results in performance degradation due to the inherent inconsistency between vision and language modalities. To overcome this challenge, existing approaches resort to modality-specific data for supervised fine-tuning, necessitating distinct model instances. We propose Slide-LoRA, which dynamically aggregates modality-specific and modality-agnostic LoRA experts, partially decoupling the multimodal generation space. Slide-LoRA harmonizes the generation of vision and language within a singular model instance, thereby facilitating a more unified generative process. Additionally, we develop a high-quality image caption dataset, DetailedTextCaps-100K, synthesized with a sophisticated closed-source MLLM to enhance visual text generation capabilities further. Comprehensive experiments across various benchmarks demonstrate the effectiveness of the proposed approach. Empowered by Slide-LoRA, TextHarmony achieves comparable performance to modality-specific fine-tuning results with only a 2% increase in parameters and shows an average improvement of 2.5% in visual text comprehension tasks and 4.0% in visual text generation tasks. Our work delineates the viability of an integrated approach to multimodal generation within the visual text domain, setting a foundation for subsequent inquiries.
MTVQA: Benchmarking Multilingual Text-Centric Visual Question Answering
May 20, 2024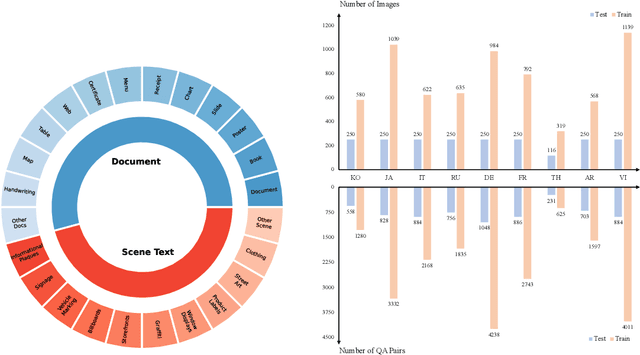
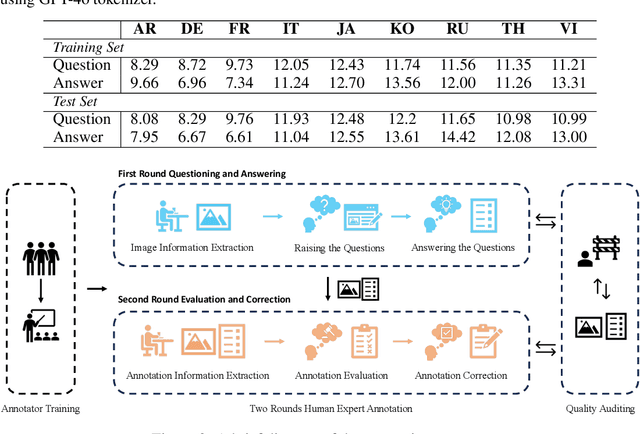

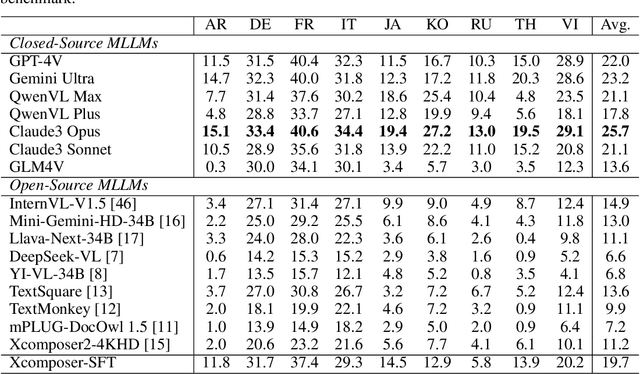
Abstract:Text-Centric Visual Question Answering (TEC-VQA) in its proper format not only facilitates human-machine interaction in text-centric visual environments but also serves as a de facto gold proxy to evaluate AI models in the domain of text-centric scene understanding. However, most TEC-VQA benchmarks have focused on high-resource languages like English and Chinese. Despite pioneering works to expand multilingual QA pairs in non-text-centric VQA datasets using translation engines, the translation-based protocol encounters a substantial ``Visual-textual misalignment'' problem when applied to TEC-VQA. Specifically, it prioritizes the text in question-answer pairs while disregarding the visual text present in images. Furthermore, it does not adequately tackle challenges related to nuanced meaning, contextual distortion, language bias, and question-type diversity. In this work, we address the task of multilingual TEC-VQA and provide a benchmark with high-quality human expert annotations in 9 diverse languages, called MTVQA. To our knowledge, MTVQA is the first multilingual TEC-VQA benchmark to provide human expert annotations for text-centric scenarios. Further, by evaluating several state-of-the-art Multimodal Large Language Models (MLLMs), including GPT-4V, on our MTVQA dataset, it is evident that there is still room for performance improvement, underscoring the value of our dataset. We hope this dataset will provide researchers with fresh perspectives and inspiration within the community. The MTVQA dataset will be available at https://huggingface.co/datasets/ByteDance/MTVQA.
TextSquare: Scaling up Text-Centric Visual Instruction Tuning
Apr 19, 2024



Abstract:Text-centric visual question answering (VQA) has made great strides with the development of Multimodal Large Language Models (MLLMs), yet open-source models still fall short of leading models like GPT4V and Gemini, partly due to a lack of extensive, high-quality instruction tuning data. To this end, we introduce a new approach for creating a massive, high-quality instruction-tuning dataset, Square-10M, which is generated using closed-source MLLMs. The data construction process, termed Square, consists of four steps: Self-Questioning, Answering, Reasoning, and Evaluation. Our experiments with Square-10M led to three key findings: 1) Our model, TextSquare, considerably surpasses open-source previous state-of-the-art Text-centric MLLMs and sets a new standard on OCRBench(62.2%). It even outperforms top-tier models like GPT4V and Gemini in 6 of 10 text-centric benchmarks. 2) Additionally, we demonstrate the critical role of VQA reasoning data in offering comprehensive contextual insights for specific questions. This not only improves accuracy but also significantly mitigates hallucinations. Specifically, TextSquare scores an average of 75.1% across four general VQA and hallucination evaluation datasets, outperforming previous state-of-the-art models. 3) Notably, the phenomenon observed in scaling text-centric VQA datasets reveals a vivid pattern: the exponential increase of instruction tuning data volume is directly proportional to the improvement in model performance, thereby validating the necessity of the dataset scale and the high quality of Square-10M.
Multi-modal In-Context Learning Makes an Ego-evolving Scene Text Recognizer
Nov 23, 2023



Abstract:Scene text recognition (STR) in the wild frequently encounters challenges when coping with domain variations, font diversity, shape deformations, etc. A straightforward solution is performing model fine-tuning tailored to a specific scenario, but it is computationally intensive and requires multiple model copies for various scenarios. Recent studies indicate that large language models (LLMs) can learn from a few demonstration examples in a training-free manner, termed "In-Context Learning" (ICL). Nevertheless, applying LLMs as a text recognizer is unacceptably resource-consuming. Moreover, our pilot experiments on LLMs show that ICL fails in STR, mainly attributed to the insufficient incorporation of contextual information from diverse samples in the training stage. To this end, we introduce E$^2$STR, a STR model trained with context-rich scene text sequences, where the sequences are generated via our proposed in-context training strategy. E$^2$STR demonstrates that a regular-sized model is sufficient to achieve effective ICL capabilities in STR. Extensive experiments show that E$^2$STR exhibits remarkable training-free adaptation in various scenarios and outperforms even the fine-tuned state-of-the-art approaches on public benchmarks.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
WSSS4LUAD: Grand Challenge on Weakly-supervised Tissue Semantic Segmentation for Lung Adenocarcinoma
Apr 14, 2022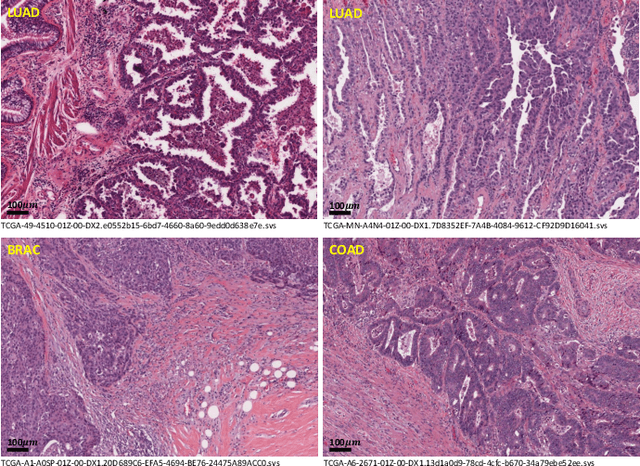
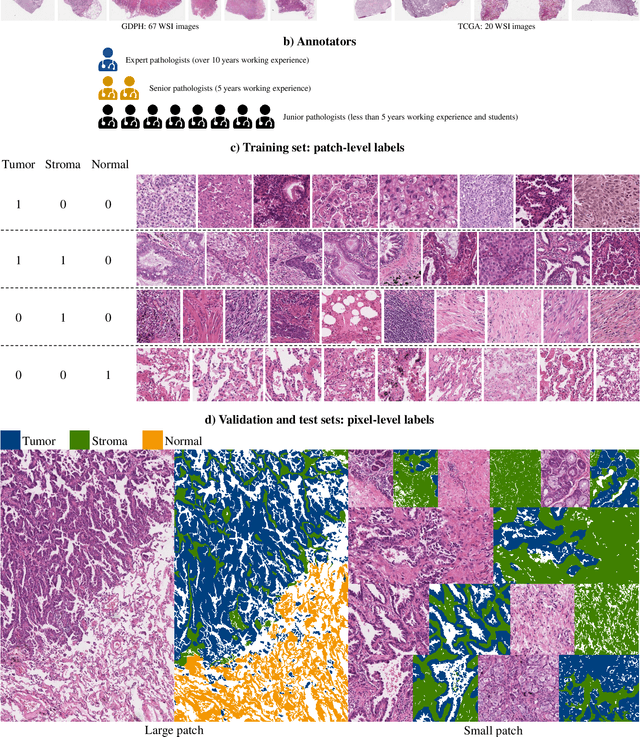
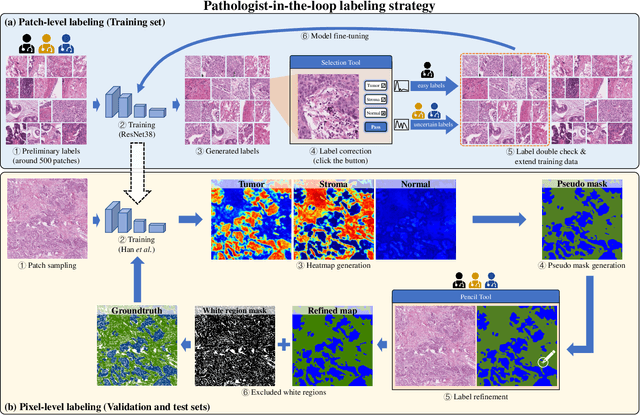
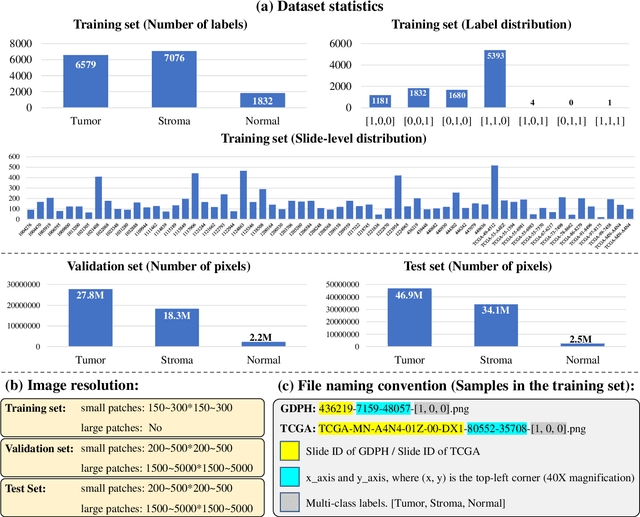
Abstract:Lung cancer is the leading cause of cancer death worldwide, and adenocarcinoma (LUAD) is the most common subtype. Exploiting the potential value of the histopathology images can promote precision medicine in oncology. Tissue segmentation is the basic upstream task of histopathology image analysis. Existing deep learning models have achieved superior segmentation performance but require sufficient pixel-level annotations, which is time-consuming and expensive. To enrich the label resources of LUAD and to alleviate the annotation efforts, we organize this challenge WSSS4LUAD to call for the outstanding weakly-supervised semantic segmentation (WSSS) techniques for histopathology images of LUAD. Participants have to design the algorithm to segment tumor epithelial, tumor-associated stroma and normal tissue with only patch-level labels. This challenge includes 10,091 patch-level annotations (the training set) and over 130 million labeled pixels (the validation and test sets), from 87 WSIs (67 from GDPH, 20 from TCGA). All the labels were generated by a pathologist-in-the-loop pipeline with the help of AI models and checked by the label review board. Among 532 registrations, 28 teams submitted the results in the test phase with over 1,000 submissions. Finally, the first place team achieved mIoU of 0.8413 (tumor: 0.8389, stroma: 0.7931, normal: 0.8919). According to the technical reports of the top-tier teams, CAM is still the most popular approach in WSSS. Cutmix data augmentation has been widely adopted to generate more reliable samples. With the success of this challenge, we believe that WSSS approaches with patch-level annotations can be a complement to the traditional pixel annotations while reducing the annotation efforts. The entire dataset has been released to encourage more researches on computational pathology in LUAD and more novel WSSS techniques.
Separable-HoverNet and Instance-YOLO for Colon Nuclei Identification and Counting
Mar 01, 2022Abstract:Nuclear segmentation, classification and quantification within Haematoxylin & Eosin stained histology images enables the extraction of interpretable cell-based features that can be used in downstream explainable models in computational pathology (CPath). However, automatic recognition of different nuclei is faced with a major challenge in that there are several different types of nuclei, some of them exhibiting large intraclass variability. In this work, we propose an approach that combine Separable-HoverNet and Instance-YOLOv5 to indentify colon nuclei small and unbalanced. Our approach can achieve mPQ+ 0.389 on the Segmentation and Classification-Preliminary Test Dataset and r2 0.599 on the Cellular Composition-Preliminary Test Dataset on ISBI 2022 CoNIC Challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge