Chu Han
BRIGHT: A Collaborative Generalist-Specialist Foundation Model for Breast Pathology
Mar 03, 2026Abstract:Generalist pathology foundation models (PFMs), pretrained on large-scale multi-organ datasets, have demonstrated remarkable predictive capabilities across diverse clinical applications. However, their proficiency on the full spectrum of clinically essential tasks within a specific organ system remains an open question due to the lack of large-scale validation cohorts for a single organ as well as the absence of a tailored training paradigm that can effectively translate broad histomorphological knowledge into the organ-specific expertise required for specialist-level interpretation. In this study, we propose BRIGHT, the first PFM specifically designed for breast pathology, trained on approximately 210 million histopathology tiles from over 51,000 breast whole-slide images derived from a cohort of over 40,000 patients across 19 hospitals. BRIGHT employs a collaborative generalist-specialist framework to capture both universal and organ-specific features. To comprehensively evaluate the performance of PFMs on breast oncology, we curate the largest multi-institutional cohorts to date for downstream task development and evaluation, comprising over 25,000 WSIs across 10 hospitals. The validation cohorts cover the full spectrum of breast pathology across 24 distinct clinical tasks spanning diagnosis, biomarker prediction, treatment response and survival prediction. Extensive experiments demonstrate that BRIGHT outperforms three leading generalist PFMs, achieving state-of-the-art (SOTA) performance in 21 of 24 internal validation tasks and in 5 of 10 external validation tasks with excellent heatmap interpretability. By evaluating on large-scale validation cohorts, this study not only demonstrates BRIGHT's clinical utility in breast oncology but also validates a collaborative generalist-specialist paradigm, providing a scalable template for developing PFMs on a specific organ system.
OMGs: A multi-agent system supporting MDT decision-making across the ovarian tumour care continuum
Feb 14, 2026Abstract:Ovarian tumour management has increasingly relied on multidisciplinary tumour board (MDT) deliberation to address treatment complexity and disease heterogeneity. However, most patients worldwide lack access to timely expert consensus, particularly in resource-constrained centres where MDT resources are scarce or unavailable. Here we present OMGs (Ovarian tumour Multidisciplinary intelligent aGent System), a multi-agent AI framework where domain-specific agents deliberate collaboratively to integrate multidisciplinary evidence and generate MDT-style recommendations with transparent rationales. To systematically evaluate MDT recommendation quality, we developed SPEAR (Safety, Personalization, Evidence, Actionability, Robustness) and validated OMGs across diverse clinical scenarios spanning the care continuum. In multicentre re-evaluation, OMGs achieved performance comparable to expert MDT consensus ($4.45 \pm 0.30$ versus $4.53 \pm 0.23$), with higher Evidence scores (4.57 versus 3.92). In prospective multicentre evaluation (59 patients), OMGs demonstrated high concordance with routine MDT decisions. Critically, in paired human-AI studies, OMGs most substantially enhanced clinicians' recommendations in Evidence and Robustness, the dimensions most compromised when multidisciplinary expertise is unavailable. These findings suggest that multi-agent deliberative systems can achieve performance comparable to expert MDT consensus, with potential to expand access to specialized oncology expertise in resource-limited settings.
Towards Computation- and Communication-efficient Computational Pathology
Apr 03, 2025Abstract:Despite the impressive performance across a wide range of applications, current computational pathology models face significant diagnostic efficiency challenges due to their reliance on high-magnification whole-slide image analysis. This limitation severely compromises their clinical utility, especially in time-sensitive diagnostic scenarios and situations requiring efficient data transfer. To address these issues, we present a novel computation- and communication-efficient framework called Magnification-Aligned Global-Local Transformer (MAGA-GLTrans). Our approach significantly reduces computational time, file transfer requirements, and storage overhead by enabling effective analysis using low-magnification inputs rather than high-magnification ones. The key innovation lies in our proposed magnification alignment (MAGA) mechanism, which employs self-supervised learning to bridge the information gap between low and high magnification levels by effectively aligning their feature representations. Through extensive evaluation across various fundamental CPath tasks, MAGA-GLTrans demonstrates state-of-the-art classification performance while achieving remarkable efficiency gains: up to 10.7 times reduction in computational time and over 20 times reduction in file transfer and storage requirements. Furthermore, we highlight the versatility of our MAGA framework through two significant extensions: (1) its applicability as a feature extractor to enhance the efficiency of any CPath architecture, and (2) its compatibility with existing foundation models and histopathology-specific encoders, enabling them to process low-magnification inputs with minimal information loss. These advancements position MAGA-GLTrans as a particularly promising solution for time-sensitive applications, especially in the context of intraoperative frozen section diagnosis where both accuracy and efficiency are paramount.
iMD4GC: Incomplete Multimodal Data Integration to Advance Precise Treatment Response Prediction and Survival Analysis for Gastric Cancer
Apr 01, 2024



Abstract:Gastric cancer (GC) is a prevalent malignancy worldwide, ranking as the fifth most common cancer with over 1 million new cases and 700 thousand deaths in 2020. Locally advanced gastric cancer (LAGC) accounts for approximately two-thirds of GC diagnoses, and neoadjuvant chemotherapy (NACT) has emerged as the standard treatment for LAGC. However, the effectiveness of NACT varies significantly among patients, with a considerable subset displaying treatment resistance. Ineffective NACT not only leads to adverse effects but also misses the optimal therapeutic window, resulting in lower survival rate. However, existing multimodal learning methods assume the availability of all modalities for each patient, which does not align with the reality of clinical practice. The limited availability of modalities for each patient would cause information loss, adversely affecting predictive accuracy. In this study, we propose an incomplete multimodal data integration framework for GC (iMD4GC) to address the challenges posed by incomplete multimodal data, enabling precise response prediction and survival analysis. Specifically, iMD4GC incorporates unimodal attention layers for each modality to capture intra-modal information. Subsequently, the cross-modal interaction layers explore potential inter-modal interactions and capture complementary information across modalities, thereby enabling information compensation for missing modalities. To evaluate iMD4GC, we collected three multimodal datasets for GC study: GastricRes (698 cases) for response prediction, GastricSur (801 cases) for survival analysis, and TCGA-STAD (400 cases) for survival analysis. The scale of our datasets is significantly larger than previous studies. The iMD4GC achieved impressive performance with an 80.2% AUC on GastricRes, 71.4% C-index on GastricSur, and 66.1% C-index on TCGA-STAD, significantly surpassing other compared methods.
BroadCAM: Outcome-agnostic Class Activation Mapping for Small-scale Weakly Supervised Applications
Sep 07, 2023



Abstract:Class activation mapping~(CAM), a visualization technique for interpreting deep learning models, is now commonly used for weakly supervised semantic segmentation~(WSSS) and object localization~(WSOL). It is the weighted aggregation of the feature maps by activating the high class-relevance ones. Current CAM methods achieve it relying on the training outcomes, such as predicted scores~(forward information), gradients~(backward information), etc. However, when with small-scale data, unstable training may lead to less effective model outcomes and generate unreliable weights, finally resulting in incorrect activation and noisy CAM seeds. In this paper, we propose an outcome-agnostic CAM approach, called BroadCAM, for small-scale weakly supervised applications. Since broad learning system (BLS) is independent to the model learning, BroadCAM can avoid the weights being affected by the unreliable model outcomes when with small-scale data. By evaluating BroadCAM on VOC2012 (natural images) and BCSS-WSSS (medical images) for WSSS and OpenImages30k for WSOL, BroadCAM demonstrates superior performance than existing CAM methods with small-scale data (less than 5\%) in different CNN architectures. It also achieves SOTA performance with large-scale training data. Extensive qualitative comparisons are conducted to demonstrate how BroadCAM activates the high class-relevance feature maps and generates reliable CAMs when with small-scale training data.
Treatment Outcome Prediction for Intracerebral Hemorrhage via Generative Prognostic Model with Imaging and Tabular Data
Jul 24, 2023


Abstract:Intracerebral hemorrhage (ICH) is the second most common and deadliest form of stroke. Despite medical advances, predicting treat ment outcomes for ICH remains a challenge. This paper proposes a novel prognostic model that utilizes both imaging and tabular data to predict treatment outcome for ICH. Our model is trained on observational data collected from non-randomized controlled trials, providing reliable predictions of treatment success. Specifically, we propose to employ a variational autoencoder model to generate a low-dimensional prognostic score, which can effectively address the selection bias resulting from the non-randomized controlled trials. Importantly, we develop a variational distributions combination module that combines the information from imaging data, non-imaging clinical data, and treatment assignment to accurately generate the prognostic score. We conducted extensive experiments on a real-world clinical dataset of intracerebral hemorrhage. Our proposed method demonstrates a substantial improvement in treatment outcome prediction compared to existing state-of-the-art approaches. Code is available at https://github.com/med-air/TOP-GPM
Rethinking Mitosis Detection: Towards Diverse Data and Feature Representation
Jul 12, 2023Abstract:Mitosis detection is one of the fundamental tasks in computational pathology, which is extremely challenging due to the heterogeneity of mitotic cell. Most of the current studies solve the heterogeneity in the technical aspect by increasing the model complexity. However, lacking consideration of the biological knowledge and the complex model design may lead to the overfitting problem while limited the generalizability of the detection model. In this paper, we systematically study the morphological appearances in different mitotic phases as well as the ambiguous non-mitotic cells and identify that balancing the data and feature diversity can achieve better generalizability. Based on this observation, we propose a novel generalizable framework (MitDet) for mitosis detection. The data diversity is considered by the proposed diversity-guided sample balancing (DGSB). And the feature diversity is preserved by inter- and intra- class feature diversity-preserved module (InCDP). Stain enhancement (SE) module is introduced to enhance the domain-relevant diversity of both data and features simultaneously. Extensive experiments have demonstrated that our proposed model outperforms all the SOTA approaches in several popular mitosis detection datasets in both internal and external test sets using minimal annotation efforts with point annotations only. Comprehensive ablation studies have also proven the effectiveness of the rethinking of data and feature diversity balancing. By analyzing the results quantitatively and qualitatively, we believe that our proposed model not only achieves SOTA performance but also might inspire the future studies in new perspectives. Source code is at https://github.com/Onehour0108/MitDet.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
FedDBL: Communication and Data Efficient Federated Deep-Broad Learning for Histopathological Tissue Classification
Feb 24, 2023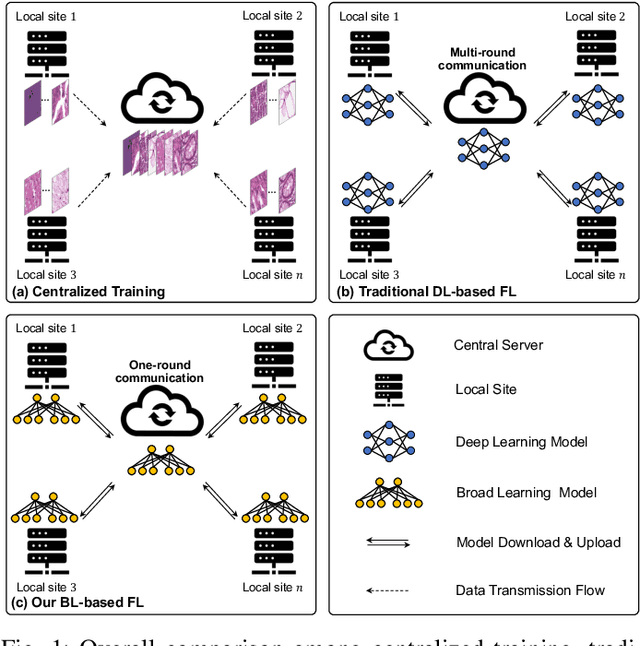
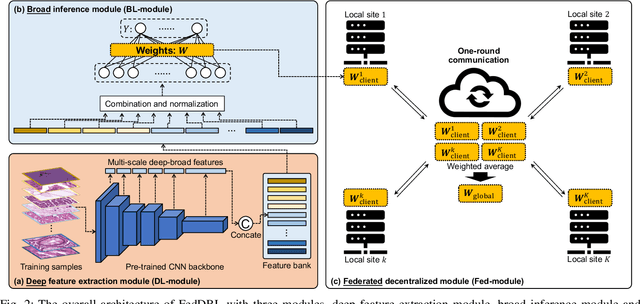
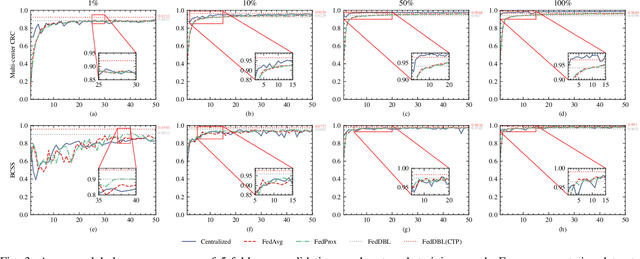
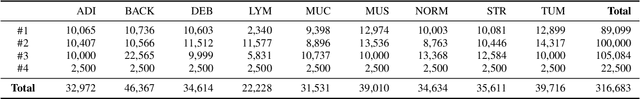
Abstract:Histopathological tissue classification is a fundamental task in computational pathology. Deep learning-based models have achieved superior performance but centralized training with data centralization suffers from the privacy leakage problem. Federated learning (FL) can safeguard privacy by keeping training samples locally, but existing FL-based frameworks require a large number of well-annotated training samples and numerous rounds of communication which hinder their practicability in the real-world clinical scenario. In this paper, we propose a universal and lightweight federated learning framework, named Federated Deep-Broad Learning (FedDBL), to achieve superior classification performance with limited training samples and only one-round communication. By simply associating a pre-trained deep learning feature extractor, a fast and lightweight broad learning inference system and a classical federated aggregation approach, FedDBL can dramatically reduce data dependency and improve communication efficiency. Five-fold cross-validation demonstrates that FedDBL greatly outperforms the competitors with only one-round communication and limited training samples, while it even achieves comparable performance with the ones under multiple-round communications. Furthermore, due to the lightweight design and one-round communication, FedDBL reduces the communication burden from 4.6GB to only 276.5KB per client using the ResNet-50 backbone at 50-round training. Since no data or deep model sharing across different clients, the privacy issue is well-solved and the model security is guaranteed with no model inversion attack risk. Code is available at https://github.com/tianpeng-deng/FedDBL.
CKD-TransBTS: Clinical Knowledge-Driven Hybrid Transformer with Modality-Correlated Cross-Attention for Brain Tumor Segmentation
Jul 15, 2022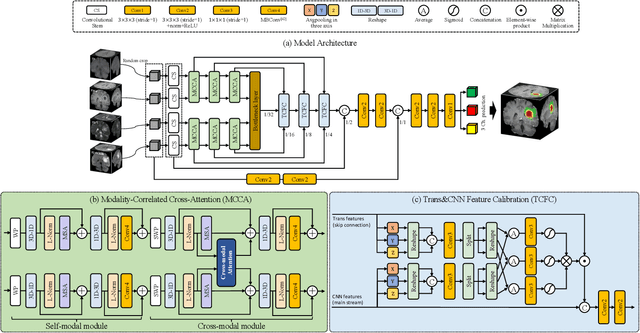
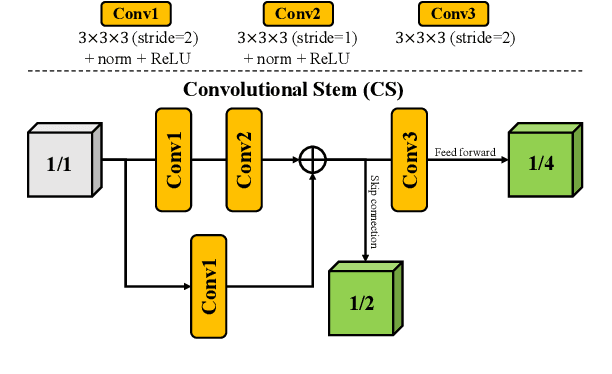
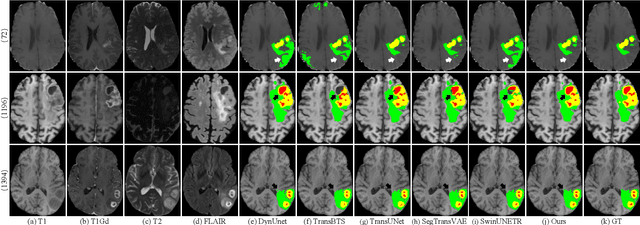
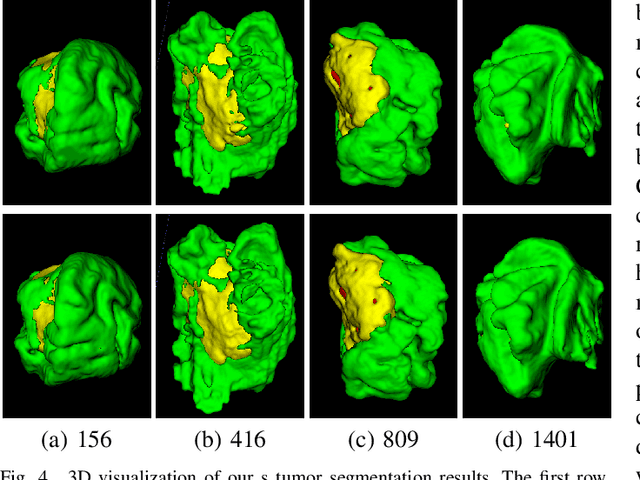
Abstract:Brain tumor segmentation (BTS) in magnetic resonance image (MRI) is crucial for brain tumor diagnosis, cancer management and research purposes. With the great success of the ten-year BraTS challenges as well as the advances of CNN and Transformer algorithms, a lot of outstanding BTS models have been proposed to tackle the difficulties of BTS in different technical aspects. However, existing studies hardly consider how to fuse the multi-modality images in a reasonable manner. In this paper, we leverage the clinical knowledge of how radiologists diagnose brain tumors from multiple MRI modalities and propose a clinical knowledge-driven brain tumor segmentation model, called CKD-TransBTS. Instead of directly concatenating all the modalities, we re-organize the input modalities by separating them into two groups according to the imaging principle of MRI. A dual-branch hybrid encoder with the proposed modality-correlated cross-attention block (MCCA) is designed to extract the multi-modality image features. The proposed model inherits the strengths from both Transformer and CNN with the local feature representation ability for precise lesion boundaries and long-range feature extraction for 3D volumetric images. To bridge the gap between Transformer and CNN features, we propose a Trans&CNN Feature Calibration block (TCFC) in the decoder. We compare the proposed model with five CNN-based models and six transformer-based models on the BraTS 2021 challenge dataset. Extensive experiments demonstrate that the proposed model achieves state-of-the-art brain tumor segmentation performance compared with all the competitors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge