Medqa Usmle
Papers and Code
Measuring Black-Box Confidence via Reasoning Trajectories: Geometry, Coverage, and Verbalization
May 07, 2026Reliable confidence estimation enables safe deployment of chain-of-thought (CoT) reasoning through text-only APIs. Yet the dominant black-box baseline, self-consistency over K samples, is linearly expensive and ignores the geometry of the trace. We propose a black-box trajectory-confidence score: we embed a CoT as a sliding-window trajectory and measure its convergence to external answer anchors with a one-parameter softmax. The method needs no logits, hidden states, or supervised calibrators. Across six (benchmark, reasoner) settings on MedQA-USMLE, GPQA Diamond, and MMLU-Pro with Gemini 3.1 Pro and Claude Sonnet 4.6, fusing this score with coverage and verbalized-confidence channels at K=4 yields Pareto improvements over self-consistency at K=8 in 6/6 settings (median AUC 0.78 vs 0.71, deltaAUC=+0.075). A fixed-pick control (+0.060) and E5 cross-embedder replication rule out answer switching and single-vendor artifacts. Geometry peaks in the penultimate window across benchmarks and reasoners, and inverts at the terminal window on GPQA Diamond. Three unscaffolded regimes separate black-box confidence into a judge-mediated Coverage prior (C), within-trace Geometry (G), and a conditional Verbalization channel (V). Across 18 benchmark x reasoner x proposer settings, C and G provide independent signal in 18/18 and 16/18, while V contributes residual signal in 6/18. Swapping the judge from GPT-5-mini to Claude Sonnet 4.6 leaves G-only AUC unchanged (|delta|<=0.013) and shifts C-only AUC by at most +/-0.02 (kappa=0.82). Fusion beats the best single channel in 17/18 settings (median AUC 0.78, max 0.92).
Domain Fine-Tuning vs. Retrieval-Augmented Generation for Medical Multiple-Choice Question Answering: A Controlled Comparison at the 4B-Parameter Scale
Apr 26, 2026Practitioners deploying small open-weight large language models (LLMs) for medical question answering face a recurring design choice: invest in a domain-fine-tuned model, or keep a general-purpose model and inject domain knowledge at inference time via retrieval-augmented generation (RAG). We isolate this trade-off by holding model size, prompt template, decoding temperature, retrieval pipeline, and evaluation protocol fixed, and varying only (i) whether the model has been domain-adapted (Gemma 3 4B vs. MedGemma 4B, both 4-bit quantized and served via Ollama) and (ii) whether retrieved passages from a medical knowledge corpus are inserted into the prompt. We evaluate all four cells of this 2x2 design on the full MedQA-USMLE 4-option test split (1,273 questions) with three repetitions per question (15,276 LLM calls). Domain fine-tuning yields a +6.8 percentage-point gain in majority-vote accuracy over the general 4B baseline (53.3% vs. 46.4%, McNemar p < 10^-4). RAG over MedMCQA explanations does not produce a statistically significant gain in either model, and in the domain-tuned model the point estimate is slightly negative (-1.9 pp, p = 0.16). At this scale and on this benchmark, domain knowledge encoded in weights dominates domain knowledge supplied in context. We release the full experiment code and JSONL traces to support replication.
A Systematic Study of Retrieval Pipeline Design for Retrieval-Augmented Medical Question Answering
Apr 08, 2026Large language models (LLMs) have demonstrated strong capabilities in medical question answering; however, purely parametric models often suffer from knowledge gaps and limited factual grounding. Retrieval-augmented generation (RAG) addresses this limitation by integrating external knowledge retrieval into the reasoning process. Despite increasing interest in RAG-based medical systems, the impact of individual retrieval components on performance remains insufficiently understood. This study presents a systematic evaluation of retrieval-augmented medical question answering using the MedQA USMLE benchmark and a structured textbook-based knowledge corpus. We analyze the interaction between language models, embedding models, retrieval strategies, query reformulation, and cross-encoder reranking within a unified experimental framework comprising forty configurations. Results show that retrieval augmentation significantly improves zero-shot medical question answering performance. The best-performing configuration was dense retrieval with query reformulation and reranking achieved 60.49% accuracy. Domain-specialized language models were also found to better utilize retrieved medical evidence than general-purpose models. The analysis further reveals a clear tradeoff between retrieval effectiveness and computational cost, with simpler dense retrieval configurations providing strong performance while maintaining higher throughput. All experiments were conducted on a single consumer-grade GPU, demonstrating that systematic evaluation of retrieval-augmented medical QA systems can be performed under modest computational resources.
Multi-Agent Reasoning with Consistency Verification Improves Uncertainty Calibration in Medical MCQA
Mar 25, 2026Miscalibrated confidence scores are a practical obstacle to deploying AI in clinical settings. A model that is always overconfident offers no useful signal for deferral. We present a multi-agent framework that combines domain-specific specialist agents with Two-Phase Verification and S-Score Weighted Fusion to improve both calibration and discrimination in medical multiple-choice question answering. Four specialist agents (respiratory, cardiology, neurology, gastroenterology) generate independent diagnoses using Qwen2.5-7B-Instruct. Each diagnosis is then subjected to a two-phase self-verification process that measures internal consistency and produces a Specialist Confidence Score (S-score). The S-scores drive a weighted fusion strategy that selects the final answer and calibrates the reported confidence. We evaluate across four experimental settings, covering 100-question and 250-question high-disagreement subsets of both MedQA-USMLE and MedMCQA. Calibration improvement is the central finding, with ECE reduced by 49-74% across all four settings, including the harder MedMCQA benchmark where these gains persist even when absolute accuracy is constrained by knowledge-intensive recall demands. On MedQA-250, the full system achieves ECE = 0.091 (74.4% reduction over the single-specialist baseline) and AUROC = 0.630 (+0.056) at 59.2% accuracy. Ablation analysis identifies Two-Phase Verification as the primary calibration driver and multi-agent reasoning as the primary accuracy driver. These results establish that consistency-based verification produces more reliable uncertainty estimates across diverse medical question types, providing a practical confidence signal for deferral in safety-critical clinical AI applications.
To Reason or Not to: Selective Chain-of-Thought in Medical Question Answering
Feb 23, 2026Objective: To improve the efficiency of medical question answering (MedQA) with large language models (LLMs) by avoiding unnecessary reasoning while maintaining accuracy. Methods: We propose Selective Chain-of-Thought (Selective CoT), an inference-time strategy that first predicts whether a question requires reasoning and generates a rationale only when needed. Two open-source LLMs (Llama-3.1-8B and Qwen-2.5-7B) were evaluated on four biomedical QA benchmarks-HeadQA, MedQA-USMLE, MedMCQA, and PubMedQA. Metrics included accuracy, total generated tokens, and inference time. Results: Selective CoT reduced inference time by 13-45% and token usage by 8-47% with minimal accuracy loss ($\leq$4\%). In some model-task pairs, it achieved both higher accuracy and greater efficiency than standard CoT. Compared with fixed-length CoT, Selective CoT reached similar or superior accuracy at substantially lower computational cost. Discussion: Selective CoT dynamically balances reasoning depth and efficiency by invoking explicit reasoning only when beneficial, reducing redundancy on recall-type questions while preserving interpretability. Conclusion: Selective CoT provides a simple, model-agnostic, and cost-effective approach for medical QA, aligning reasoning effort with question complexity to enhance real-world deployability of LLM-based clinical systems.
Capabilities of GPT-5 on Multimodal Medical Reasoning
Aug 13, 2025Recent advances in large language models (LLMs) have enabled general-purpose systems to perform increasingly complex domain-specific reasoning without extensive fine-tuning. In the medical domain, decision-making often requires integrating heterogeneous information sources, including patient narratives, structured data, and medical images. This study positions GPT-5 as a generalist multimodal reasoner for medical decision support and systematically evaluates its zero-shot chain-of-thought reasoning performance on both text-based question answering and visual question answering tasks under a unified protocol. We benchmark GPT-5, GPT-5-mini, GPT-5-nano, and GPT-4o-2024-11-20 against standardized splits of MedQA, MedXpertQA (text and multimodal), MMLU medical subsets, USMLE self-assessment exams, and VQA-RAD. Results show that GPT-5 consistently outperforms all baselines, achieving state-of-the-art accuracy across all QA benchmarks and delivering substantial gains in multimodal reasoning. On MedXpertQA MM, GPT-5 improves reasoning and understanding scores by +29.26% and +26.18% over GPT-4o, respectively, and surpasses pre-licensed human experts by +24.23% in reasoning and +29.40% in understanding. In contrast, GPT-4o remains below human expert performance in most dimensions. A representative case study demonstrates GPT-5's ability to integrate visual and textual cues into a coherent diagnostic reasoning chain, recommending appropriate high-stakes interventions. Our results show that, on these controlled multimodal reasoning benchmarks, GPT-5 moves from human-comparable to above human-expert performance. This improvement may substantially inform the design of future clinical decision-support systems.
WiNGPT-3.0 Technical Report
May 23, 2025



Current Large Language Models (LLMs) exhibit significant limitations, notably in structured, interpretable, and verifiable medical reasoning, alongside practical deployment challenges related to computational resources and data privacy. This report focused on the development of WiNGPT-3.0, the 32-billion parameter LLMs, engineered with the objective of enhancing its capacity for medical reasoning and exploring its potential for effective integration within healthcare IT infrastructures. The broader aim is to advance towards clinically applicable models. The approach involved a multi-stage training pipeline tailored for general, medical, and clinical reasoning. This pipeline incorporated supervised fine-tuning (SFT) and reinforcement learning (RL), leveraging curated Long Chain-of-Thought (CoT) datasets, auxiliary reward models, and an evidence-based diagnostic chain simulation. WiNGPT-3.0 demonstrated strong performance: specific model variants achieved scores of 66.6 on MedCalc and 87.1 on MedQA-USMLE. Furthermore, targeted training improved performance on a clinical reasoning task from a baseline score of 58.1 to 62.5. These findings suggest that reinforcement learning, even when applied with a limited dataset of only a few thousand examples, can enhance medical reasoning accuracy. Crucially, this demonstration of RL's efficacy with limited data and computation paves the way for more trustworthy and practically deployable LLMs within clinical workflows and health information infrastructures.
Disentangling Reasoning and Knowledge in Medical Large Language Models
May 16, 2025



Medical reasoning in large language models (LLMs) aims to emulate clinicians' diagnostic thinking, but current benchmarks such as MedQA-USMLE, MedMCQA, and PubMedQA often mix reasoning with factual recall. We address this by separating 11 biomedical QA benchmarks into reasoning- and knowledge-focused subsets using a PubMedBERT classifier that reaches 81 percent accuracy, comparable to human performance. Our analysis shows that only 32.8 percent of questions require complex reasoning. We evaluate biomedical models (HuatuoGPT-o1, MedReason, m1) and general-domain models (DeepSeek-R1, o4-mini, Qwen3), finding consistent gaps between knowledge and reasoning performance. For example, m1 scores 60.5 on knowledge but only 47.1 on reasoning. In adversarial tests where models are misled with incorrect initial reasoning, biomedical models degrade sharply, while larger or RL-trained general models show more robustness. To address this, we train BioMed-R1 using fine-tuning and reinforcement learning on reasoning-heavy examples. It achieves the strongest performance among similarly sized models. Further gains may come from incorporating clinical case reports and training with adversarial and backtracking scenarios.
Open-Medical-R1: How to Choose Data for RLVR Training at Medicine Domain
Apr 16, 2025



This paper explores optimal data selection strategies for Reinforcement Learning with Verified Rewards (RLVR) training in the medical domain. While RLVR has shown exceptional potential for enhancing reasoning capabilities in large language models, most prior implementations have focused on mathematics and logical puzzles, with limited exploration of domain-specific applications like medicine. We investigate four distinct data sampling strategies from MedQA-USMLE: random sampling (baseline), and filtering using Phi-4, Gemma-3-27b-it, and Gemma-3-12b-it models. Using Gemma-3-12b-it as our base model and implementing Group Relative Policy Optimization (GRPO), we evaluate performance across multiple benchmarks including MMLU, GSM8K, MMLU-Pro, and CMMLU. Our findings demonstrate that models trained on filtered data generally outperform those trained on randomly selected samples. Notably, training on self-filtered samples (using Gemma-3-12b-it for filtering) achieved superior performance in medical domains but showed reduced robustness across different benchmarks, while filtering with larger models from the same series yielded better overall robustness. These results provide valuable insights into effective data organization strategies for RLVR in specialized domains and highlight the importance of thoughtful data selection in achieving optimal performance. You can access our repository (https://github.com/Qsingle/open-medical-r1) to get the codes.
AfriMed-QA: A Pan-African, Multi-Specialty, Medical Question-Answering Benchmark Dataset
Nov 23, 2024

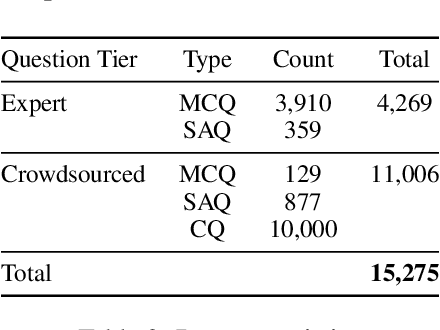

Recent advancements in large language model(LLM) performance on medical multiple choice question (MCQ) benchmarks have stimulated interest from healthcare providers and patients globally. Particularly in low-and middle-income countries (LMICs) facing acute physician shortages and lack of specialists, LLMs offer a potentially scalable pathway to enhance healthcare access and reduce costs. However, their effectiveness in the Global South, especially across the African continent, remains to be established. In this work, we introduce AfriMed-QA, the first large scale Pan-African English multi-specialty medical Question-Answering (QA) dataset, 15,000 questions (open and closed-ended) sourced from over 60 medical schools across 16 countries, covering 32 medical specialties. We further evaluate 30 LLMs across multiple axes including correctness and demographic bias. Our findings show significant performance variation across specialties and geographies, MCQ performance clearly lags USMLE (MedQA). We find that biomedical LLMs underperform general models and smaller edge-friendly LLMs struggle to achieve a passing score. Interestingly, human evaluations show a consistent consumer preference for LLM answers and explanations when compared with clinician answers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge