Khaled Saab
A prospective clinical feasibility study of a conversational diagnostic AI in an ambulatory primary care clinic
Mar 10, 2026Abstract:Large language model (LLM)-based AI systems have shown promise for patient-facing diagnostic and management conversations in simulated settings. Translating these systems into clinical practice requires assessment in real-world workflows with rigorous safety oversight. We report a prospective, single-arm feasibility study of an LLM-based conversational AI, the Articulate Medical Intelligence Explorer (AMIE), conducting clinical history taking and presentation of potential diagnoses for patients to discuss with their provider at urgent care appointments at a leading academic medical center. 100 adult patients completed an AMIE text-chat interaction up to 5 days before their appointment. We sought to assess the conversational safety and quality, patient and clinician experience, and clinical reasoning capabilities compared to primary care providers (PCPs). Human safety supervisors monitored all patient-AMIE interactions in real time and did not need to intervene to stop any consultations based on pre-defined criteria. Patients reported high satisfaction and their attitudes towards AI improved after interacting with AMIE (p < 0.001). PCPs found AMIE's output useful with a positive impact on preparedness. AMIE's differential diagnosis (DDx) included the final diagnosis, per chart review 8 weeks post-encounter, in 90% of cases, with 75% top-3 accuracy. Blinded assessment of AMIE and PCP DDx and management (Mx) plans suggested similar overall DDx and Mx plan quality, without significant differences for DDx (p = 0.6) and appropriateness and safety of Mx (p = 0.1 and 1.0, respectively). PCPs outperformed AMIE in the practicality (p = 0.003) and cost effectiveness (p = 0.004) of Mx. While further research is needed, this study demonstrates the initial feasibility, safety, and user acceptance of conversational AI in a real-world setting, representing crucial steps towards clinical translation.
Advancing Conversational Diagnostic AI with Multimodal Reasoning
May 06, 2025



Abstract:Large Language Models (LLMs) have demonstrated great potential for conducting diagnostic conversations but evaluation has been largely limited to language-only interactions, deviating from the real-world requirements of remote care delivery. Instant messaging platforms permit clinicians and patients to upload and discuss multimodal medical artifacts seamlessly in medical consultation, but the ability of LLMs to reason over such data while preserving other attributes of competent diagnostic conversation remains unknown. Here we advance the conversational diagnosis and management performance of the Articulate Medical Intelligence Explorer (AMIE) through a new capability to gather and interpret multimodal data, and reason about this precisely during consultations. Leveraging Gemini 2.0 Flash, our system implements a state-aware dialogue framework, where conversation flow is dynamically controlled by intermediate model outputs reflecting patient states and evolving diagnoses. Follow-up questions are strategically directed by uncertainty in such patient states, leading to a more structured multimodal history-taking process that emulates experienced clinicians. We compared AMIE to primary care physicians (PCPs) in a randomized, blinded, OSCE-style study of chat-based consultations with patient actors. We constructed 105 evaluation scenarios using artifacts like smartphone skin photos, ECGs, and PDFs of clinical documents across diverse conditions and demographics. Our rubric assessed multimodal capabilities and other clinically meaningful axes like history-taking, diagnostic accuracy, management reasoning, communication, and empathy. Specialist evaluation showed AMIE to be superior to PCPs on 7/9 multimodal and 29/32 non-multimodal axes (including diagnostic accuracy). The results show clear progress in multimodal conversational diagnostic AI, but real-world translation needs further research.
Systematic Evaluation of Large Vision-Language Models for Surgical Artificial Intelligence
Apr 03, 2025Abstract:Large Vision-Language Models offer a new paradigm for AI-driven image understanding, enabling models to perform tasks without task-specific training. This flexibility holds particular promise across medicine, where expert-annotated data is scarce. Yet, VLMs' practical utility in intervention-focused domains--especially surgery, where decision-making is subjective and clinical scenarios are variable--remains uncertain. Here, we present a comprehensive analysis of 11 state-of-the-art VLMs across 17 key visual understanding tasks in surgical AI--from anatomy recognition to skill assessment--using 13 datasets spanning laparoscopic, robotic, and open procedures. In our experiments, VLMs demonstrate promising generalizability, at times outperforming supervised models when deployed outside their training setting. In-context learning, incorporating examples during testing, boosted performance up to three-fold, suggesting adaptability as a key strength. Still, tasks requiring spatial or temporal reasoning remained difficult. Beyond surgery, our findings offer insights into VLMs' potential for tackling complex and dynamic scenarios in clinical and broader real-world applications.
Towards Conversational AI for Disease Management
Mar 08, 2025



Abstract:While large language models (LLMs) have shown promise in diagnostic dialogue, their capabilities for effective management reasoning - including disease progression, therapeutic response, and safe medication prescription - remain under-explored. We advance the previously demonstrated diagnostic capabilities of the Articulate Medical Intelligence Explorer (AMIE) through a new LLM-based agentic system optimised for clinical management and dialogue, incorporating reasoning over the evolution of disease and multiple patient visit encounters, response to therapy, and professional competence in medication prescription. To ground its reasoning in authoritative clinical knowledge, AMIE leverages Gemini's long-context capabilities, combining in-context retrieval with structured reasoning to align its output with relevant and up-to-date clinical practice guidelines and drug formularies. In a randomized, blinded virtual Objective Structured Clinical Examination (OSCE) study, AMIE was compared to 21 primary care physicians (PCPs) across 100 multi-visit case scenarios designed to reflect UK NICE Guidance and BMJ Best Practice guidelines. AMIE was non-inferior to PCPs in management reasoning as assessed by specialist physicians and scored better in both preciseness of treatments and investigations, and in its alignment with and grounding of management plans in clinical guidelines. To benchmark medication reasoning, we developed RxQA, a multiple-choice question benchmark derived from two national drug formularies (US, UK) and validated by board-certified pharmacists. While AMIE and PCPs both benefited from the ability to access external drug information, AMIE outperformed PCPs on higher difficulty questions. While further research would be needed before real-world translation, AMIE's strong performance across evaluations marks a significant step towards conversational AI as a tool in disease management.
Towards an AI co-scientist
Feb 26, 2025Abstract:Scientific discovery relies on scientists generating novel hypotheses that undergo rigorous experimental validation. To augment this process, we introduce an AI co-scientist, a multi-agent system built on Gemini 2.0. The AI co-scientist is intended to help uncover new, original knowledge and to formulate demonstrably novel research hypotheses and proposals, building upon prior evidence and aligned to scientist-provided research objectives and guidance. The system's design incorporates a generate, debate, and evolve approach to hypothesis generation, inspired by the scientific method and accelerated by scaling test-time compute. Key contributions include: (1) a multi-agent architecture with an asynchronous task execution framework for flexible compute scaling; (2) a tournament evolution process for self-improving hypotheses generation. Automated evaluations show continued benefits of test-time compute, improving hypothesis quality. While general purpose, we focus development and validation in three biomedical areas: drug repurposing, novel target discovery, and explaining mechanisms of bacterial evolution and anti-microbial resistance. For drug repurposing, the system proposes candidates with promising validation findings, including candidates for acute myeloid leukemia that show tumor inhibition in vitro at clinically applicable concentrations. For novel target discovery, the AI co-scientist proposed new epigenetic targets for liver fibrosis, validated by anti-fibrotic activity and liver cell regeneration in human hepatic organoids. Finally, the AI co-scientist recapitulated unpublished experimental results via a parallel in silico discovery of a novel gene transfer mechanism in bacterial evolution. These results, detailed in separate, co-timed reports, demonstrate the potential to augment biomedical and scientific discovery and usher an era of AI empowered scientists.
Advancing Multimodal Medical Capabilities of Gemini
May 06, 2024



Abstract:Many clinical tasks require an understanding of specialized data, such as medical images and genomics, which is not typically found in general-purpose large multimodal models. Building upon Gemini's multimodal models, we develop several models within the new Med-Gemini family that inherit core capabilities of Gemini and are optimized for medical use via fine-tuning with 2D and 3D radiology, histopathology, ophthalmology, dermatology and genomic data. Med-Gemini-2D sets a new standard for AI-based chest X-ray (CXR) report generation based on expert evaluation, exceeding previous best results across two separate datasets by an absolute margin of 1% and 12%, where 57% and 96% of AI reports on normal cases, and 43% and 65% on abnormal cases, are evaluated as "equivalent or better" than the original radiologists' reports. We demonstrate the first ever large multimodal model-based report generation for 3D computed tomography (CT) volumes using Med-Gemini-3D, with 53% of AI reports considered clinically acceptable, although additional research is needed to meet expert radiologist reporting quality. Beyond report generation, Med-Gemini-2D surpasses the previous best performance in CXR visual question answering (VQA) and performs well in CXR classification and radiology VQA, exceeding SoTA or baselines on 17 of 20 tasks. In histopathology, ophthalmology, and dermatology image classification, Med-Gemini-2D surpasses baselines across 18 out of 20 tasks and approaches task-specific model performance. Beyond imaging, Med-Gemini-Polygenic outperforms the standard linear polygenic risk score-based approach for disease risk prediction and generalizes to genetically correlated diseases for which it has never been trained. Although further development and evaluation are necessary in the safety-critical medical domain, our results highlight the potential of Med-Gemini across a wide range of medical tasks.
Capabilities of Gemini Models in Medicine
May 01, 2024



Abstract:Excellence in a wide variety of medical applications poses considerable challenges for AI, requiring advanced reasoning, access to up-to-date medical knowledge and understanding of complex multimodal data. Gemini models, with strong general capabilities in multimodal and long-context reasoning, offer exciting possibilities in medicine. Building on these core strengths of Gemini, we introduce Med-Gemini, a family of highly capable multimodal models that are specialized in medicine with the ability to seamlessly use web search, and that can be efficiently tailored to novel modalities using custom encoders. We evaluate Med-Gemini on 14 medical benchmarks, establishing new state-of-the-art (SoTA) performance on 10 of them, and surpass the GPT-4 model family on every benchmark where a direct comparison is viable, often by a wide margin. On the popular MedQA (USMLE) benchmark, our best-performing Med-Gemini model achieves SoTA performance of 91.1% accuracy, using a novel uncertainty-guided search strategy. On 7 multimodal benchmarks including NEJM Image Challenges and MMMU (health & medicine), Med-Gemini improves over GPT-4V by an average relative margin of 44.5%. We demonstrate the effectiveness of Med-Gemini's long-context capabilities through SoTA performance on a needle-in-a-haystack retrieval task from long de-identified health records and medical video question answering, surpassing prior bespoke methods using only in-context learning. Finally, Med-Gemini's performance suggests real-world utility by surpassing human experts on tasks such as medical text summarization, alongside demonstrations of promising potential for multimodal medical dialogue, medical research and education. Taken together, our results offer compelling evidence for Med-Gemini's potential, although further rigorous evaluation will be crucial before real-world deployment in this safety-critical domain.
Towards Conversational Diagnostic AI
Jan 11, 2024Abstract:At the heart of medicine lies the physician-patient dialogue, where skillful history-taking paves the way for accurate diagnosis, effective management, and enduring trust. Artificial Intelligence (AI) systems capable of diagnostic dialogue could increase accessibility, consistency, and quality of care. However, approximating clinicians' expertise is an outstanding grand challenge. Here, we introduce AMIE (Articulate Medical Intelligence Explorer), a Large Language Model (LLM) based AI system optimized for diagnostic dialogue. AMIE uses a novel self-play based simulated environment with automated feedback mechanisms for scaling learning across diverse disease conditions, specialties, and contexts. We designed a framework for evaluating clinically-meaningful axes of performance including history-taking, diagnostic accuracy, management reasoning, communication skills, and empathy. We compared AMIE's performance to that of primary care physicians (PCPs) in a randomized, double-blind crossover study of text-based consultations with validated patient actors in the style of an Objective Structured Clinical Examination (OSCE). The study included 149 case scenarios from clinical providers in Canada, the UK, and India, 20 PCPs for comparison with AMIE, and evaluations by specialist physicians and patient actors. AMIE demonstrated greater diagnostic accuracy and superior performance on 28 of 32 axes according to specialist physicians and 24 of 26 axes according to patient actors. Our research has several limitations and should be interpreted with appropriate caution. Clinicians were limited to unfamiliar synchronous text-chat which permits large-scale LLM-patient interactions but is not representative of usual clinical practice. While further research is required before AMIE could be translated to real-world settings, the results represent a milestone towards conversational diagnostic AI.
Towards trustworthy seizure onset detection using workflow notes
Jun 14, 2023



Abstract:A major barrier to deploying healthcare AI models is their trustworthiness. One form of trustworthiness is a model's robustness across different subgroups: while existing models may exhibit expert-level performance on aggregate metrics, they often rely on non-causal features, leading to errors in hidden subgroups. To take a step closer towards trustworthy seizure onset detection from EEG, we propose to leverage annotations that are produced by healthcare personnel in routine clinical workflows -- which we refer to as workflow notes -- that include multiple event descriptions beyond seizures. Using workflow notes, we first show that by scaling training data to an unprecedented level of 68,920 EEG hours, seizure onset detection performance significantly improves (+12.3 AUROC points) compared to relying on smaller training sets with expensive manual gold-standard labels. Second, we reveal that our binary seizure onset detection model underperforms on clinically relevant subgroups (e.g., up to a margin of 6.5 AUROC points between pediatrics and adults), while having significantly higher false positives on EEG clips showing non-epileptiform abnormalities compared to any EEG clip (+19 FPR points). To improve model robustness to hidden subgroups, we train a multilabel model that classifies 26 attributes other than seizures, such as spikes, slowing, and movement artifacts. We find that our multilabel model significantly improves overall seizure onset detection performance (+5.9 AUROC points) while greatly improving performance among subgroups (up to +8.3 AUROC points), and decreases false positives on non-epileptiform abnormalities by 8 FPR points. Finally, we propose a clinical utility metric based on false positives per 24 EEG hours and find that our multilabel model improves this clinical utility metric by a factor of 2x across different clinical settings.
The Importance of Background Information for Out of Distribution Generalization
Jun 17, 2022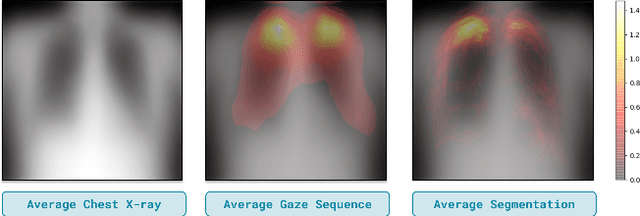
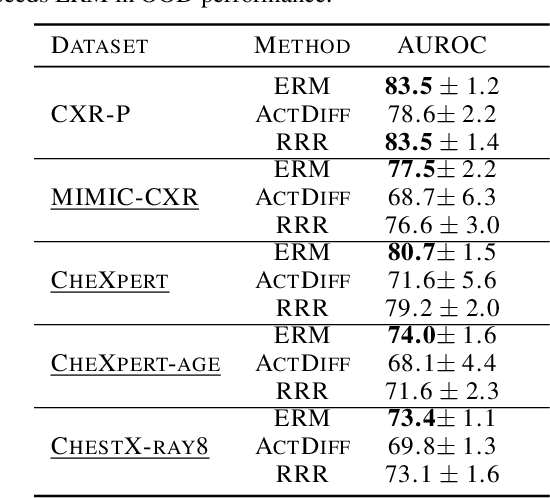

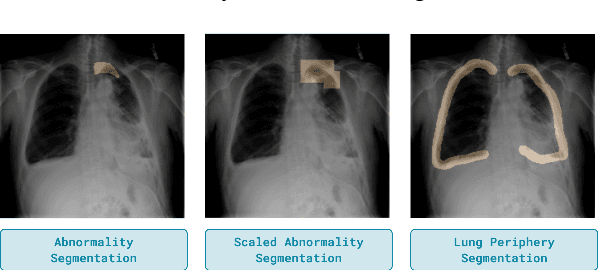
Abstract:Domain generalization in medical image classification is an important problem for trustworthy machine learning to be deployed in healthcare. We find that existing approaches for domain generalization which utilize ground-truth abnormality segmentations to control feature attributions have poor out-of-distribution (OOD) performance relative to the standard baseline of empirical risk minimization (ERM). We investigate what regions of an image are important for medical image classification and show that parts of the background, that which is not contained in the abnormality segmentation, provides helpful signal. We then develop a new task-specific mask which covers all relevant regions. Utilizing this new segmentation mask significantly improves the performance of the existing methods on the OOD test sets. To obtain better generalization results than ERM, we find it necessary to scale up the training data size in addition to the usage of these task-specific masks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge