Yoshua Bengio
DIRO
FL Games: A Federated Learning Framework for Distribution Shifts
Oct 31, 2022



Abstract:Federated learning aims to train predictive models for data that is distributed across clients, under the orchestration of a server. However, participating clients typically each hold data from a different distribution, which can yield to catastrophic generalization on data from a different client, which represents a new domain. In this work, we argue that in order to generalize better across non-i.i.d. clients, it is imperative to only learn correlations that are stable and invariant across domains. We propose FL GAMES, a game-theoretic framework for federated learning that learns causal features that are invariant across clients. While training to achieve the Nash equilibrium, the traditional best response strategy suffers from high-frequency oscillations. We demonstrate that FL GAMES effectively resolves this challenge and exhibits smooth performance curves. Further, FL GAMES scales well in the number of clients, requires significantly fewer communication rounds, and is agnostic to device heterogeneity. Through empirical evaluation, we demonstrate that FL GAMES achieves high out-of-distribution performance on various benchmarks.
Multi-Objective GFlowNets
Oct 23, 2022Abstract:In many applications of machine learning, like drug discovery and material design, the goal is to generate candidates that simultaneously maximize a set of objectives. As these objectives are often conflicting, there is no single candidate that simultaneously maximizes all objectives, but rather a set of Pareto-optimal candidates where one objective cannot be improved without worsening another. Moreover, in practice, these objectives are often under-specified, making the diversity of candidates a key consideration. The existing multi-objective optimization methods focus predominantly on covering the Pareto front, failing to capture diversity in the space of candidates. Motivated by the success of GFlowNets for generation of diverse candidates in a single objective setting, in this paper we consider Multi-Objective GFlowNets (MOGFNs). MOGFNs consist of a novel Conditional GFlowNet which models a family of single-objective sub-problems derived by decomposing the multi-objective optimization problem. Our work is the first to empirically demonstrate conditional GFlowNets. Through a series of experiments on synthetic and benchmark tasks, we empirically demonstrate that MOGFNs outperform existing methods in terms of Hypervolume, R2-distance and candidate diversity. We also demonstrate the effectiveness of MOGFNs over existing methods in active learning settings. Finally, we supplement our empirical results with a careful analysis of each component of MOGFNs.
Neural Attentive Circuits
Oct 19, 2022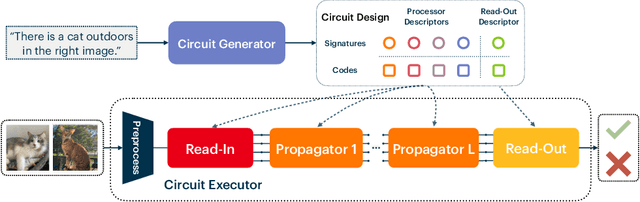
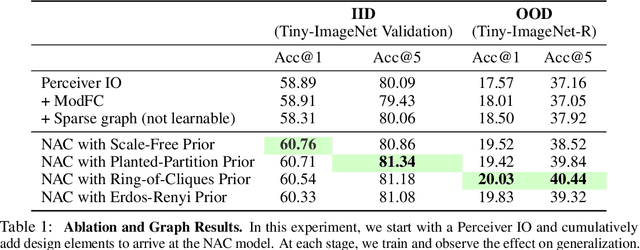
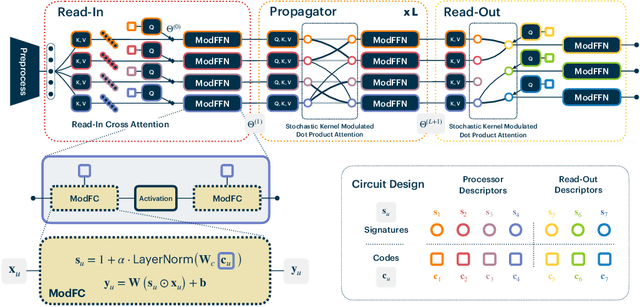

Abstract:Recent work has seen the development of general purpose neural architectures that can be trained to perform tasks across diverse data modalities. General purpose models typically make few assumptions about the underlying data-structure and are known to perform well in the large-data regime. At the same time, there has been growing interest in modular neural architectures that represent the data using sparsely interacting modules. These models can be more robust out-of-distribution, computationally efficient, and capable of sample-efficient adaptation to new data. However, they tend to make domain-specific assumptions about the data, and present challenges in how module behavior (i.e., parameterization) and connectivity (i.e., their layout) can be jointly learned. In this work, we introduce a general purpose, yet modular neural architecture called Neural Attentive Circuits (NACs) that jointly learns the parameterization and a sparse connectivity of neural modules without using domain knowledge. NACs are best understood as the combination of two systems that are jointly trained end-to-end: one that determines the module configuration and the other that executes it on an input. We demonstrate qualitatively that NACs learn diverse and meaningful module configurations on the NLVR2 dataset without additional supervision. Quantitatively, we show that by incorporating modularity in this way, NACs improve upon a strong non-modular baseline in terms of low-shot adaptation on CIFAR and CUBs dataset by about 10%, and OOD robustness on Tiny ImageNet-R by about 2.5%. Further, we find that NACs can achieve an 8x speedup at inference time while losing less than 3% performance. Finally, we find NACs to yield competitive results on diverse data modalities spanning point-cloud classification, symbolic processing and text-classification from ASCII bytes, thereby confirming its general purpose nature.
Toward Next-Generation Artificial Intelligence: Catalyzing the NeuroAI Revolution
Oct 15, 2022Abstract:Neuroscience has long been an important driver of progress in artificial intelligence (AI). We propose that to accelerate progress in AI, we must invest in fundamental research in NeuroAI.
Contrastive introspection (ConSpec) to rapidly identify invariant steps for success
Oct 12, 2022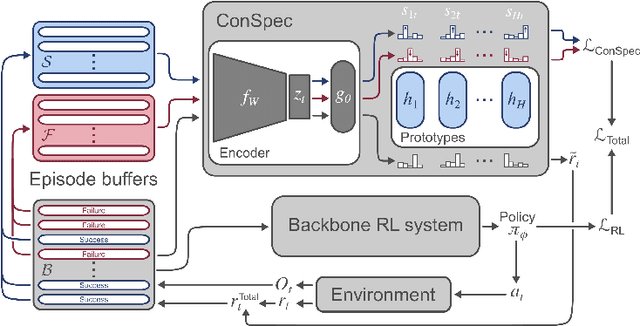
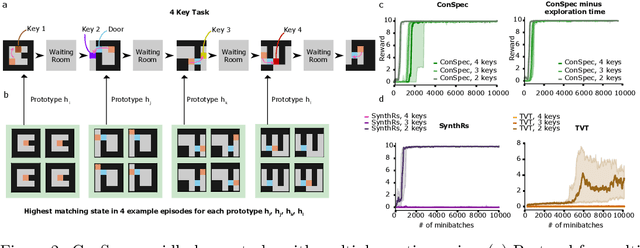
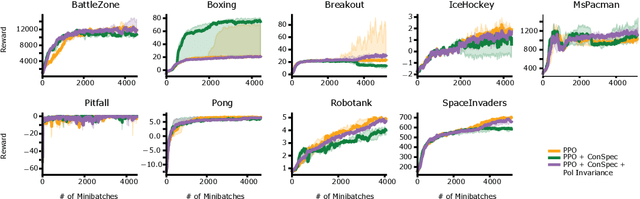
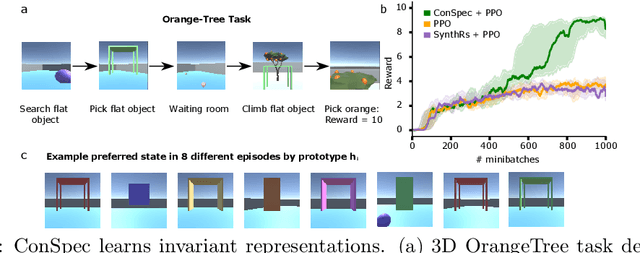
Abstract:Reinforcement learning (RL) algorithms have achieved notable success in recent years, but still struggle with fundamental issues in long-term credit assignment. It remains difficult to learn in situations where success is contingent upon multiple critical steps that are distant in time from each other and from a sparse reward; as is often the case in real life. Moreover, how RL algorithms assign credit in these difficult situations is typically not coded in a way that can rapidly generalize to new situations. Here, we present an approach using offline contrastive learning, which we call contrastive introspection (ConSpec), that can be added to any existing RL algorithm and addresses both issues. In ConSpec, a contrastive loss is used during offline replay to identify invariances among successful episodes. This takes advantage of the fact that it is easier to retrospectively identify the small set of steps that success is contingent upon than it is to prospectively predict reward at every step taken in the environment. ConSpec stores this knowledge in a collection of prototypes summarizing the intermediate states required for success. During training, arrival at any state that matches these prototypes generates an intrinsic reward that is added to any external rewards. As well, the reward shaping provided by ConSpec can be made to preserve the optimal policy of the underlying RL agent. The prototypes in ConSpec provide two key benefits for credit assignment: (1) They enable rapid identification of all the critical states. (2) They do so in a readily interpretable manner, enabling out of distribution generalization when sensory features are altered. In summary, ConSpec is a modular system that can be added to any existing RL algorithm to improve its long-term credit assignment.
MAgNet: Mesh Agnostic Neural PDE Solver
Oct 11, 2022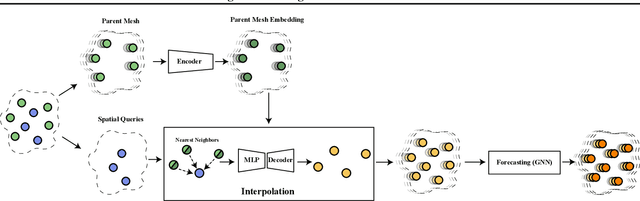
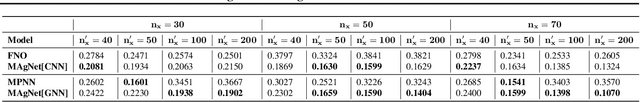
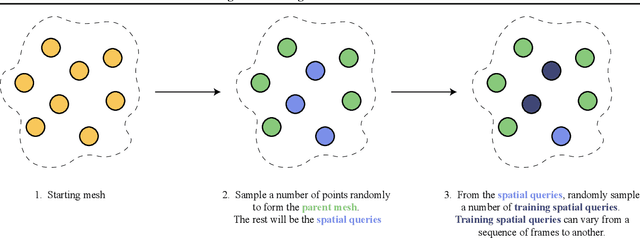
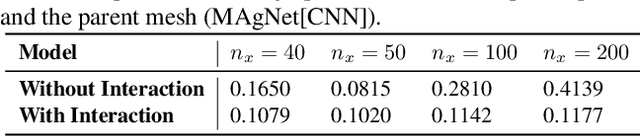
Abstract:The computational complexity of classical numerical methods for solving Partial Differential Equations (PDE) scales significantly as the resolution increases. As an important example, climate predictions require fine spatio-temporal resolutions to resolve all turbulent scales in the fluid simulations. This makes the task of accurately resolving these scales computationally out of reach even with modern supercomputers. As a result, current numerical modelers solve PDEs on grids that are too coarse (3km to 200km on each side), which hinders the accuracy and usefulness of the predictions. In this paper, we leverage the recent advances in Implicit Neural Representations (INR) to design a novel architecture that predicts the spatially continuous solution of a PDE given a spatial position query. By augmenting coordinate-based architectures with Graph Neural Networks (GNN), we enable zero-shot generalization to new non-uniform meshes and long-term predictions up to 250 frames ahead that are physically consistent. Our Mesh Agnostic Neural PDE Solver (MAgNet) is able to make accurate predictions across a variety of PDE simulation datasets and compares favorably with existing baselines. Moreover, MAgNet generalizes well to different meshes and resolutions up to four times those trained on.
Robust and Controllable Object-Centric Learning through Energy-based Models
Oct 11, 2022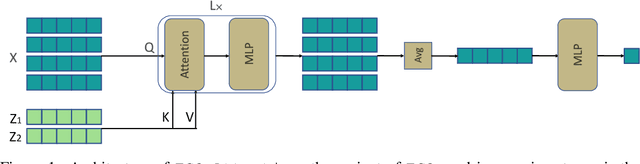
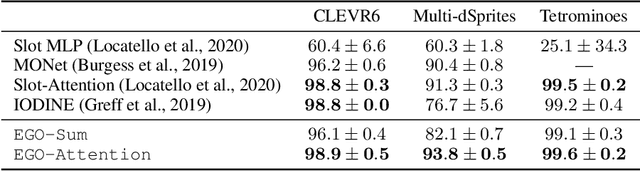
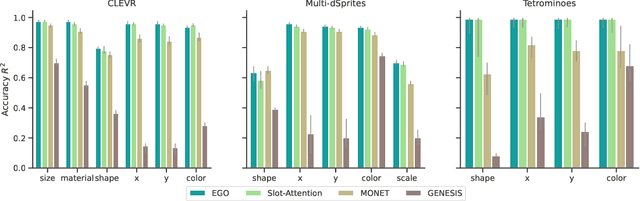
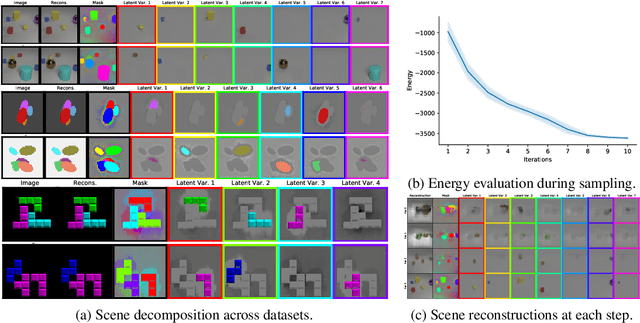
Abstract:Humans are remarkably good at understanding and reasoning about complex visual scenes. The capability to decompose low-level observations into discrete objects allows us to build a grounded abstract representation and identify the compositional structure of the world. Accordingly, it is a crucial step for machine learning models to be capable of inferring objects and their properties from visual scenes without explicit supervision. However, existing works on object-centric representation learning either rely on tailor-made neural network modules or strong probabilistic assumptions in the underlying generative and inference processes. In this work, we present \ours, a conceptually simple and general approach to learning object-centric representations through an energy-based model. By forming a permutation-invariant energy function using vanilla attention blocks readily available in Transformers, we can infer object-centric latent variables via gradient-based MCMC methods where permutation equivariance is automatically guaranteed. We show that \ours can be easily integrated into existing architectures and can effectively extract high-quality object-centric representations, leading to better segmentation accuracy and competitive downstream task performance. Further, empirical evaluations show that \ours's learned representations are robust against distribution shift. Finally, we demonstrate the effectiveness of \ours in systematic compositional generalization, by re-composing learned energy functions for novel scene generation and manipulation.
Interventional Causal Representation Learning
Oct 09, 2022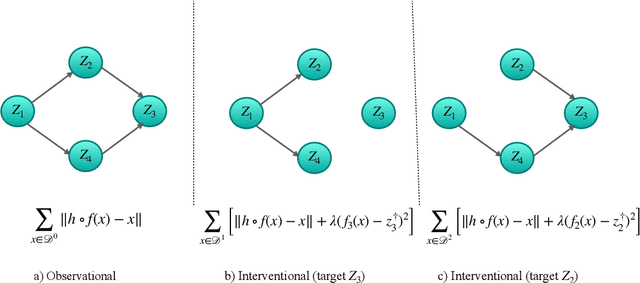
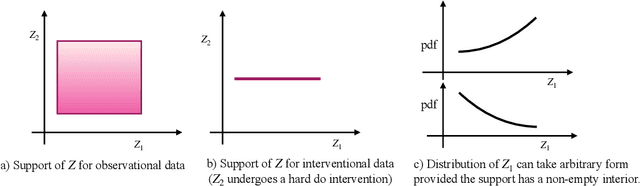
Abstract:The theory of identifiable representation learning aims to build general-purpose methods that extract high-level latent (causal) factors from low-level sensory data. Most existing works focus on identifiable representation learning with observational data, relying on distributional assumptions on latent (causal) factors. However, in practice, we often also have access to interventional data for representation learning. How can we leverage interventional data to help identify high-level latents? To this end, we explore the role of interventional data for identifiable representation learning in this work. We study the identifiability of latent causal factors with and without interventional data, under minimal distributional assumptions on the latents. We prove that, if the true latent variables map to the observed high-dimensional data via a polynomial function, then representation learning via minimizing the standard reconstruction loss of autoencoders identifies the true latents up to affine transformation. If we further have access to interventional data generated by hard $do$ interventions on some of the latents, then we can identify these intervened latents up to permutation, shift and scaling.
Stateful active facilitator: Coordination and Environmental Heterogeneity in Cooperative Multi-Agent Reinforcement Learning
Oct 07, 2022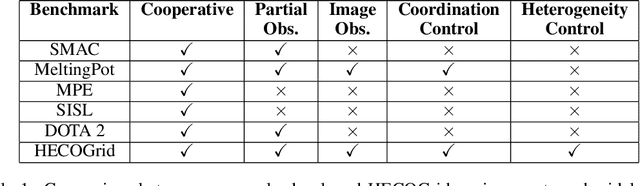
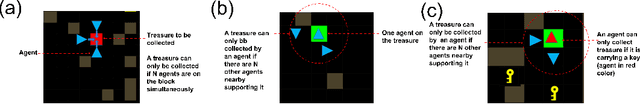
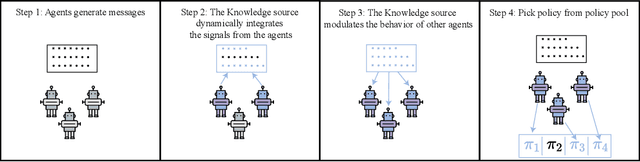
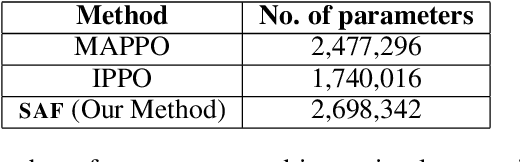
Abstract:In cooperative multi-agent reinforcement learning, a team of agents works together to achieve a common goal. Different environments or tasks may require varying degrees of coordination among agents in order to achieve the goal in an optimal way. The nature of coordination will depend on properties of the environment -- its spatial layout, distribution of obstacles, dynamics, etc. We term this variation of properties within an environment as heterogeneity. Existing literature has not sufficiently addressed the fact that different environments may have different levels of heterogeneity. We formalize the notions of coordination level and heterogeneity level of an environment and present HECOGrid, a suite of multi-agent RL environments that facilitates empirical evaluation of different MARL approaches across different levels of coordination and environmental heterogeneity by providing a quantitative control over coordination and heterogeneity levels of the environment. Further, we propose a Centralized Training Decentralized Execution learning approach called Stateful Active Facilitator (SAF) that enables agents to work efficiently in high-coordination and high-heterogeneity environments through a differentiable and shared knowledge source used during training and dynamic selection from a shared pool of policies. We evaluate SAF and compare its performance against baselines IPPO and MAPPO on HECOGrid. Our results show that SAF consistently outperforms the baselines across different tasks and different heterogeneity and coordination levels.
Generative Augmented Flow Networks
Oct 07, 2022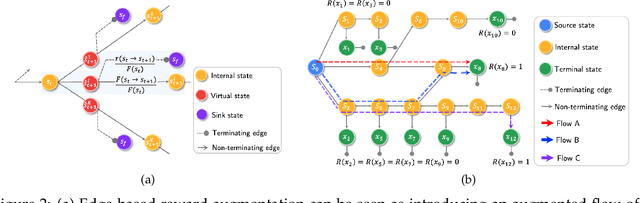
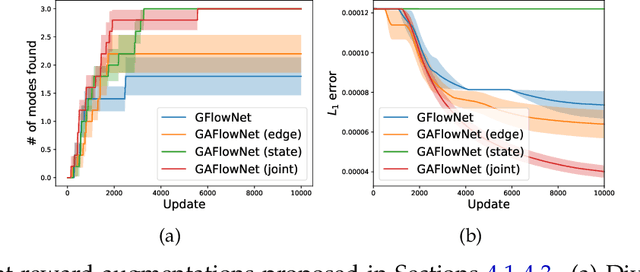
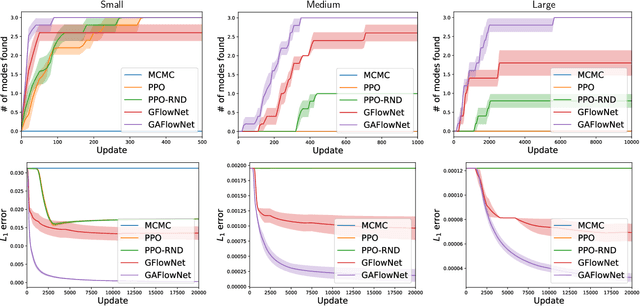
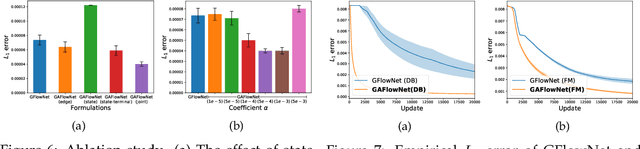
Abstract:The Generative Flow Network is a probabilistic framework where an agent learns a stochastic policy for object generation, such that the probability of generating an object is proportional to a given reward function. Its effectiveness has been shown in discovering high-quality and diverse solutions, compared to reward-maximizing reinforcement learning-based methods. Nonetheless, GFlowNets only learn from rewards of the terminal states, which can limit its applicability. Indeed, intermediate rewards play a critical role in learning, for example from intrinsic motivation to provide intermediate feedback even in particularly challenging sparse reward tasks. Inspired by this, we propose Generative Augmented Flow Networks (GAFlowNets), a novel learning framework to incorporate intermediate rewards into GFlowNets. We specify intermediate rewards by intrinsic motivation to tackle the exploration problem in sparse reward environments. GAFlowNets can leverage edge-based and state-based intrinsic rewards in a joint way to improve exploration. Based on extensive experiments on the GridWorld task, we demonstrate the effectiveness and efficiency of GAFlowNet in terms of convergence, performance, and diversity of solutions. We further show that GAFlowNet is scalable to a more complex and large-scale molecule generation domain, where it achieves consistent and significant performance improvement.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge