Quanzheng Li
CAMCA, Massachusetts General Hospital and Harvard Medical School
AdaFish: Fast low-rank parameter-efficient fine-tuning by using second-order information
Mar 19, 2024Abstract:Recent advancements in large-scale pretrained models have significantly improved performance across a variety of tasks in natural language processing and computer vision. However, the extensive number of parameters in these models necessitates substantial memory and computational resources for full training. To adapt these models for downstream tasks or specific application-oriented datasets, parameter-efficient fine-tuning methods leveraging pretrained parameters have gained considerable attention. However, it can still be time-consuming due to lots of parameters and epochs. In this work, we introduce AdaFish, an efficient algorithm of the second-order type designed to expedite the training process within low-rank decomposition-based fine-tuning frameworks. Our key observation is that the associated generalized Fisher information matrix is either low-rank or extremely small-scaled. Such a generalized Fisher information matrix is shown to be equivalent to the Hessian matrix. Moreover, we prove the global convergence of AdaFish, along with its iteration/oracle complexity. Numerical experiments show that our algorithm is quite competitive with the state-of-the-art AdamW method.
Cardiac Magnetic Resonance 2D+T Short- and Long-axis Segmentation via Spatio-temporal SAM Adaptation
Mar 15, 2024



Abstract:Accurate 2D+T myocardium segmentation in cine cardiac magnetic resonance (CMR) scans is essential to analyze LV motion throughout the cardiac cycle comprehensively. The Segment Anything Model (SAM), known for its accurate segmentation and zero-shot generalization, has not yet been tailored for CMR 2D+T segmentation. We therefore introduce CMR2D+T-SAM, a novel approach to adapt SAM for CMR 2D+T segmentation using spatio-temporal adaption. This approach also incorporates a U-Net framework for multi-scale feature extraction, as well as text prompts for accurate segmentation on both short-axis (SAX) and long-axis (LAX) views using a single model. CMR2D+T-SAM outperforms existing deep learning methods on the STACOM2011 dataset, achieving a myocardium Dice score of 0.885 and a Hausdorff distance (HD) of 2.900 pixels. It also demonstrates superior zero-shot generalization on the ACDC dataset with a Dice score of 0.840 and a HD of 4.076 pixels.
Conditional Score-Based Diffusion Model for Cortical Thickness Trajectory Prediction
Mar 11, 2024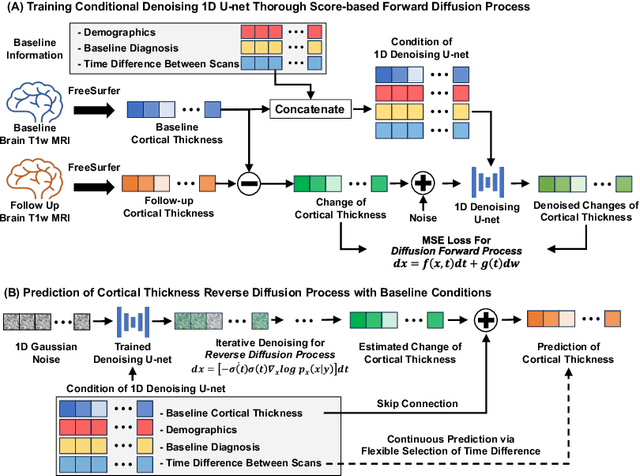
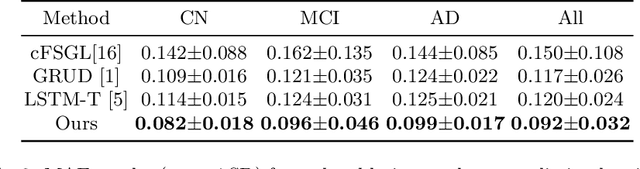
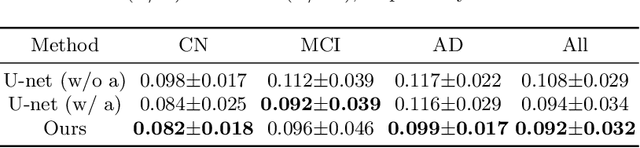
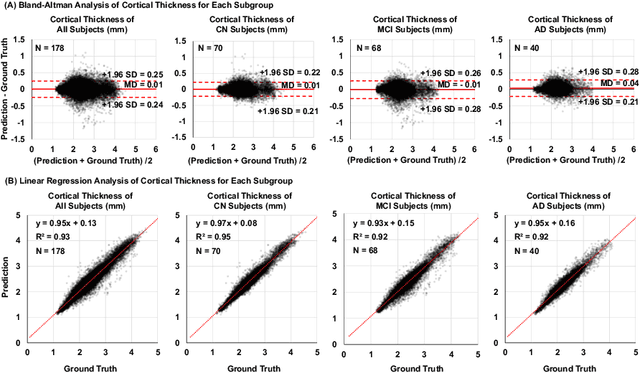
Abstract:Alzheimer's Disease (AD) is a neurodegenerative condition characterized by diverse progression rates among individuals, with changes in cortical thickness (CTh) closely linked to its progression. Accurately forecasting CTh trajectories can significantly enhance early diagnosis and intervention strategies, providing timely care. However, the longitudinal data essential for these studies often suffer from temporal sparsity and incompleteness, presenting substantial challenges in modeling the disease's progression accurately. Existing methods are limited, focusing primarily on datasets without missing entries or requiring predefined assumptions about CTh progression. To overcome these obstacles, we propose a conditional score-based diffusion model specifically designed to generate CTh trajectories with the given baseline information, such as age, sex, and initial diagnosis. Our conditional diffusion model utilizes all available data during the training phase to make predictions based solely on baseline information during inference without needing prior history about CTh progression. The prediction accuracy of the proposed CTh prediction pipeline using a conditional score-based model was compared for sub-groups consisting of cognitively normal, mild cognitive impairment, and AD subjects. The Bland-Altman analysis shows our diffusion-based prediction model has a near-zero bias with narrow 95% confidential interval compared to the ground-truth CTh in 6-36 months. In addition, our conditional diffusion model has a stochastic generative nature, therefore, we demonstrated an uncertainty analysis of patient-specific CTh prediction through multiple realizations.
Medical Image Synthesis via Fine-Grained Image-Text Alignment and Anatomy-Pathology Prompting
Mar 11, 2024



Abstract:Data scarcity and privacy concerns limit the availability of high-quality medical images for public use, which can be mitigated through medical image synthesis. However, current medical image synthesis methods often struggle to accurately capture the complexity of detailed anatomical structures and pathological conditions. To address these challenges, we propose a novel medical image synthesis model that leverages fine-grained image-text alignment and anatomy-pathology prompts to generate highly detailed and accurate synthetic medical images. Our method integrates advanced natural language processing techniques with image generative modeling, enabling precise alignment between descriptive text prompts and the synthesized images' anatomical and pathological details. The proposed approach consists of two key components: an anatomy-pathology prompting module and a fine-grained alignment-based synthesis module. The anatomy-pathology prompting module automatically generates descriptive prompts for high-quality medical images. To further synthesize high-quality medical images from the generated prompts, the fine-grained alignment-based synthesis module pre-defines a visual codebook for the radiology dataset and performs fine-grained alignment between the codebook and generated prompts to obtain key patches as visual clues, facilitating accurate image synthesis. We validate the superiority of our method through experiments on public chest X-ray datasets and demonstrate that our synthetic images preserve accurate semantic information, making them valuable for various medical applications.
Implicit Image-to-Image Schrodinger Bridge for CT Super-Resolution and Denoising
Mar 10, 2024Abstract:Conditional diffusion models have gained recognition for their effectiveness in image restoration tasks, yet their iterative denoising process, starting from Gaussian noise, often leads to slow inference speeds. As a promising alternative, the Image-to-Image Schr\"odinger Bridge (I2SB) initializes the generative process from corrupted images and integrates training techniques from conditional diffusion models. In this study, we extended the I2SB method by introducing the Implicit Image-to-Image Schrodinger Bridge (I3SB), transitioning its generative process to a non-Markovian process by incorporating corrupted images in each generative step. This enhancement empowers I3SB to generate images with better texture restoration using a small number of generative steps. The proposed method was validated on CT super-resolution and denoising tasks and outperformed existing methods, including the conditional denoising diffusion probabilistic model (cDDPM) and I2SB, in both visual quality and quantitative metrics. These findings underscore the potential of I3SB in improving medical image restoration by providing fast and accurate generative modeling.
The Radiation Oncology NLP Database
Jan 19, 2024



Abstract:We present the Radiation Oncology NLP Database (ROND), the first dedicated Natural Language Processing (NLP) dataset for radiation oncology, an important medical specialty that has received limited attention from the NLP community in the past. With the advent of Artificial General Intelligence (AGI), there is an increasing need for specialized datasets and benchmarks to facilitate research and development. ROND is specifically designed to address this gap in the domain of radiation oncology, a field that offers many opportunities for NLP exploration. It encompasses various NLP tasks including Logic Reasoning, Text Classification, Named Entity Recognition (NER), Question Answering (QA), Text Summarization, and Patient-Clinician Conversations, each with a distinct focus on radiation oncology concepts and application cases. In addition, we have developed an instruction-tuning dataset consisting of over 20k instruction pairs (based on ROND) and trained a large language model, CancerChat. This serves to demonstrate the potential of instruction-tuning large language models within a highly-specialized medical domain. The evaluation results in this study could serve as baseline results for future research. ROND aims to stimulate advancements in radiation oncology and clinical NLP by offering a platform for testing and improving algorithms and models in a domain-specific context. The ROND dataset is a joint effort of multiple U.S. health institutions. The data is available at https://github.com/zl-liu/Radiation-Oncology-NLP-Database.
APP-RUSS: Automated Path Planning for Robotic Ultrasound System
Oct 22, 2023Abstract:Autonomous robotic ultrasound System (RUSS) has been extensively studied. However, fully automated ultrasound image acquisition is still challenging, partly due to the lack of study in combining two phases of path planning: guiding the ultrasound probe to the scan target and covering the scan surface or volume. This paper presents a system of Automated Path Planning for RUSS (APP-RUSS). Our focus is on the first phase of automation, which emphasizes directing the ultrasound probe's path toward the target over extended distances. Specifically, our APP-RUSS system consists of a RealSense D405 RGB-D camera that is employed for visual guidance of the UR5e robotic arm and a cubic Bezier curve path planning model that is customized for delivering the probe to the recognized target. APP-RUSS can contribute to understanding the integration of the two phases of path planning in robotic ultrasound imaging, paving the way for its clinical adoption.
MediViSTA-SAM: Zero-shot Medical Video Analysis with Spatio-temporal SAM Adaptation
Sep 24, 2023Abstract:In recent years, the Segmentation Anything Model (SAM) has attracted considerable attention as a foundational model well-known for its robust generalization capabilities across various downstream tasks. However, SAM does not exhibit satisfactory performance in the realm of medical image analysis. In this study, we introduce the first study on adapting SAM on video segmentation, called MediViSTA-SAM, a novel approach designed for medical video segmentation. Given video data, MediViSTA, spatio-temporal adapter captures long and short range temporal attention with cross-frame attention mechanism effectively constraining it to consider the immediately preceding video frame as a reference, while also considering spatial information effectively. Additionally, it incorporates multi-scale fusion by employing a U-shaped encoder and a modified mask decoder to handle objects of varying sizes. To evaluate our approach, extensive experiments were conducted using state-of-the-art (SOTA) methods, assessing its generalization abilities on multi-vendor in-house echocardiography datasets. The results highlight the accuracy and effectiveness of our network in medical video segmentation.
RadOnc-GPT: A Large Language Model for Radiation Oncology
Sep 22, 2023



Abstract:This paper presents RadOnc-GPT, a large language model specialized for radiation oncology through advanced tuning methods. RadOnc-GPT was finetuned on a large dataset of radiation oncology patient records and clinical notes from the Mayo Clinic in Arizona. The model employs instruction tuning on three key tasks - generating radiotherapy treatment regimens, determining optimal radiation modalities, and providing diagnostic descriptions/ICD codes based on patient diagnostic details. Evaluations conducted by comparing RadOnc-GPT outputs to general large language model outputs showed that RadOnc-GPT generated outputs with significantly improved clarity, specificity, and clinical relevance. The study demonstrated the potential of using large language models fine-tuned using domain-specific knowledge like RadOnc-GPT to achieve transformational capabilities in highly specialized healthcare fields such as radiation oncology.
PolicyGPT: Automated Analysis of Privacy Policies with Large Language Models
Sep 19, 2023



Abstract:Privacy policies serve as the primary conduit through which online service providers inform users about their data collection and usage procedures. However, in a bid to be comprehensive and mitigate legal risks, these policy documents are often quite verbose. In practical use, users tend to click the Agree button directly rather than reading them carefully. This practice exposes users to risks of privacy leakage and legal issues. Recently, the advent of Large Language Models (LLM) such as ChatGPT and GPT-4 has opened new possibilities for text analysis, especially for lengthy documents like privacy policies. In this study, we investigate a privacy policy text analysis framework PolicyGPT based on the LLM. This framework was tested using two datasets. The first dataset comprises of privacy policies from 115 websites, which were meticulously annotated by legal experts, categorizing each segment into one of 10 classes. The second dataset consists of privacy policies from 304 popular mobile applications, with each sentence manually annotated and classified into one of another 10 categories. Under zero-shot learning conditions, PolicyGPT demonstrated robust performance. For the first dataset, it achieved an accuracy rate of 97%, while for the second dataset, it attained an 87% accuracy rate, surpassing that of the baseline machine learning and neural network models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge