Wenting Chen
MedSAM-Agent: Empowering Interactive Medical Image Segmentation with Multi-turn Agentic Reinforcement Learning
Feb 03, 2026Abstract:Medical image segmentation is evolving from task-specific models toward generalizable frameworks. Recent research leverages Multi-modal Large Language Models (MLLMs) as autonomous agents, employing reinforcement learning with verifiable reward (RLVR) to orchestrate specialized tools like the Segment Anything Model (SAM). However, these approaches often rely on single-turn, rigid interaction strategies and lack process-level supervision during training, which hinders their ability to fully exploit the dynamic potential of interactive tools and leads to redundant actions. To bridge this gap, we propose MedSAM-Agent, a framework that reformulates interactive segmentation as a multi-step autonomous decision-making process. First, we introduce a hybrid prompting strategy for expert-curated trajectory generation, enabling the model to internalize human-like decision heuristics and adaptive refinement strategies. Furthermore, we develop a two-stage training pipeline that integrates multi-turn, end-to-end outcome verification with a clinical-fidelity process reward design to promote interaction parsimony and decision efficiency. Extensive experiments across 6 medical modalities and 21 datasets demonstrate that MedSAM-Agent achieves state-of-the-art performance, effectively unifying autonomous medical reasoning with robust, iterative optimization. Code is available \href{https://github.com/CUHK-AIM-Group/MedSAM-Agent}{here}.
MMedExpert-R1: Strengthening Multimodal Medical Reasoning via Domain-Specific Adaptation and Clinical Guideline Reinforcement
Jan 16, 2026Abstract:Medical Vision-Language Models (MedVLMs) excel at perception tasks but struggle with complex clinical reasoning required in real-world scenarios. While reinforcement learning (RL) has been explored to enhance reasoning capabilities, existing approaches face critical mismatches: the scarcity of deep reasoning data, cold-start limits multi-specialty alignment, and standard RL algorithms fail to model clinical reasoning diversity. We propose MMedExpert-R1, a novel reasoning MedVLM that addresses these challenges through domain-specific adaptation and clinical guideline reinforcement. We construct MMedExpert, a high-quality dataset of 10K samples across four specialties with step-by-step reasoning traces. Our Domain-Specific Adaptation (DSA) creates specialty-specific LoRA modules to provide diverse initialization, while Guideline-Based Advantages (GBA) explicitly models different clinical reasoning perspectives to align with real-world diagnostic strategies. Conflict-Aware Capability Integration then merges these specialized experts into a unified agent, ensuring robust multi-specialty alignment. Comprehensive experiments demonstrate state-of-the-art performance, with our 7B model achieving 27.50 on MedXpert-MM and 83.03 on OmniMedVQA, establishing a robust foundation for reliable multimodal medical reasoning systems.
Benchmarking Egocentric Clinical Intent Understanding Capability for Medical Multimodal Large Language Models
Jan 11, 2026Abstract:Medical Multimodal Large Language Models (Med-MLLMs) require egocentric clinical intent understanding for real-world deployment, yet existing benchmarks fail to evaluate this critical capability. To address these challenges, we introduce MedGaze-Bench, the first benchmark leveraging clinician gaze as a Cognitive Cursor to assess intent understanding across surgery, emergency simulation, and diagnostic interpretation. Our benchmark addresses three fundamental challenges: visual homogeneity of anatomical structures, strict temporal-causal dependencies in clinical workflows, and implicit adherence to safety protocols. We propose a Three-Dimensional Clinical Intent Framework evaluating: (1) Spatial Intent: discriminating precise targets amid visual noise, (2) Temporal Intent: inferring causal rationale through retrospective and prospective reasoning, and (3) Standard Intent: verifying protocol compliance through safety checks. Beyond accuracy metrics, we introduce Trap QA mechanisms to stress-test clinical reliability by penalizing hallucinations and cognitive sycophancy. Experiments reveal current MLLMs struggle with egocentric intent due to over-reliance on global features, leading to fabricated observations and uncritical acceptance of invalid instructions.
MedEinst: Benchmarking the Einstellung Effect in Medical LLMs through Counterfactual Differential Diagnosis
Jan 10, 2026Abstract:Despite achieving high accuracy on medical benchmarks, LLMs exhibit the Einstellung Effect in clinical diagnosis--relying on statistical shortcuts rather than patient-specific evidence, causing misdiagnosis in atypical cases. Existing benchmarks fail to detect this critical failure mode. We introduce MedEinst, a counterfactual benchmark with 5,383 paired clinical cases across 49 diseases. Each pair contains a control case and a "trap" case with altered discriminative evidence that flips the diagnosis. We measure susceptibility via Bias Trap Rate--probability of misdiagnosing traps despite correctly diagnosing controls. Extensive Evaluation of 17 LLMs shows frontier models achieve high baseline accuracy but severe bias trap rates. Thus, we propose ECR-Agent, aligning LLM reasoning with Evidence-Based Medicine standard via two components: (1) Dynamic Causal Inference (DCI) performs structured reasoning through dual-pathway perception, dynamic causal graph reasoning across three levels (association, intervention, counterfactual), and evidence audit for final diagnosis; (2) Critic-Driven Graph and Memory Evolution (CGME) iteratively refines the system by storing validated reasoning paths in an exemplar base and consolidating disease-specific knowledge into evolving illness graphs. Source code is to be released.
FaNe: Towards Fine-Grained Cross-Modal Contrast with False-Negative Reduction and Text-Conditioned Sparse Attention
Nov 15, 2025Abstract:Medical vision-language pre-training (VLP) offers significant potential for advancing medical image understanding by leveraging paired image-report data. However, existing methods are limited by Fa}lse Negatives (FaNe) induced by semantically similar texts and insufficient fine-grained cross-modal alignment. To address these limitations, we propose FaNe, a semantic-enhanced VLP framework. To mitigate false negatives, we introduce a semantic-aware positive pair mining strategy based on text-text similarity with adaptive normalization. Furthermore, we design a text-conditioned sparse attention pooling module to enable fine-grained image-text alignment through localized visual representations guided by textual cues. To strengthen intra-modal discrimination, we develop a hard-negative aware contrastive loss that adaptively reweights semantically similar negatives. Extensive experiments on five downstream medical imaging benchmarks demonstrate that FaNe achieves state-of-the-art performance across image classification, object detection, and semantic segmentation, validating the effectiveness of our framework.
Medical Reasoning in the Era of LLMs: A Systematic Review of Enhancement Techniques and Applications
Aug 01, 2025Abstract:The proliferation of Large Language Models (LLMs) in medicine has enabled impressive capabilities, yet a critical gap remains in their ability to perform systematic, transparent, and verifiable reasoning, a cornerstone of clinical practice. This has catalyzed a shift from single-step answer generation to the development of LLMs explicitly designed for medical reasoning. This paper provides the first systematic review of this emerging field. We propose a taxonomy of reasoning enhancement techniques, categorized into training-time strategies (e.g., supervised fine-tuning, reinforcement learning) and test-time mechanisms (e.g., prompt engineering, multi-agent systems). We analyze how these techniques are applied across different data modalities (text, image, code) and in key clinical applications such as diagnosis, education, and treatment planning. Furthermore, we survey the evolution of evaluation benchmarks from simple accuracy metrics to sophisticated assessments of reasoning quality and visual interpretability. Based on an analysis of 60 seminal studies from 2022-2025, we conclude by identifying critical challenges, including the faithfulness-plausibility gap and the need for native multimodal reasoning, and outlining future directions toward building efficient, robust, and sociotechnically responsible medical AI.
RadFabric: Agentic AI System with Reasoning Capability for Radiology
Jun 17, 2025Abstract:Chest X ray (CXR) imaging remains a critical diagnostic tool for thoracic conditions, but current automated systems face limitations in pathology coverage, diagnostic accuracy, and integration of visual and textual reasoning. To address these gaps, we propose RadFabric, a multi agent, multimodal reasoning framework that unifies visual and textual analysis for comprehensive CXR interpretation. RadFabric is built on the Model Context Protocol (MCP), enabling modularity, interoperability, and scalability for seamless integration of new diagnostic agents. The system employs specialized CXR agents for pathology detection, an Anatomical Interpretation Agent to map visual findings to precise anatomical structures, and a Reasoning Agent powered by large multimodal reasoning models to synthesize visual, anatomical, and clinical data into transparent and evidence based diagnoses. RadFabric achieves significant performance improvements, with near-perfect detection of challenging pathologies like fractures (1.000 accuracy) and superior overall diagnostic accuracy (0.799) compared to traditional systems (0.229 to 0.527). By integrating cross modal feature alignment and preference-driven reasoning, RadFabric advances AI-driven radiology toward transparent, anatomically precise, and clinically actionable CXR analysis.
TumorGen: Boundary-Aware Tumor-Mask Synthesis with Rectified Flow Matching
May 30, 2025



Abstract:Tumor data synthesis offers a promising solution to the shortage of annotated medical datasets. However, current approaches either limit tumor diversity by using predefined masks or employ computationally expensive two-stage processes with multiple denoising steps, causing computational inefficiency. Additionally, these methods typically rely on binary masks that fail to capture the gradual transitions characteristic of tumor boundaries. We present TumorGen, a novel Boundary-Aware Tumor-Mask Synthesis with Rectified Flow Matching for efficient 3D tumor synthesis with three key components: a Boundary-Aware Pseudo Mask Generation module that replaces strict binary masks with flexible bounding boxes; a Spatial-Constraint Vector Field Estimator that simultaneously synthesizes tumor latents and masks using rectified flow matching to ensure computational efficiency; and a VAE-guided mask refiner that enhances boundary realism. TumorGen significantly improves computational efficiency by requiring fewer sampling steps while maintaining pathological accuracy through coarse and fine-grained spatial constraints. Experimental results demonstrate TumorGen's superior performance over existing tumor synthesis methods in both efficiency and realism, offering a valuable contribution to AI-driven cancer diagnostics.
A Comprehensive Evaluation of Multi-Modal Large Language Models for Endoscopy Analysis
May 29, 2025
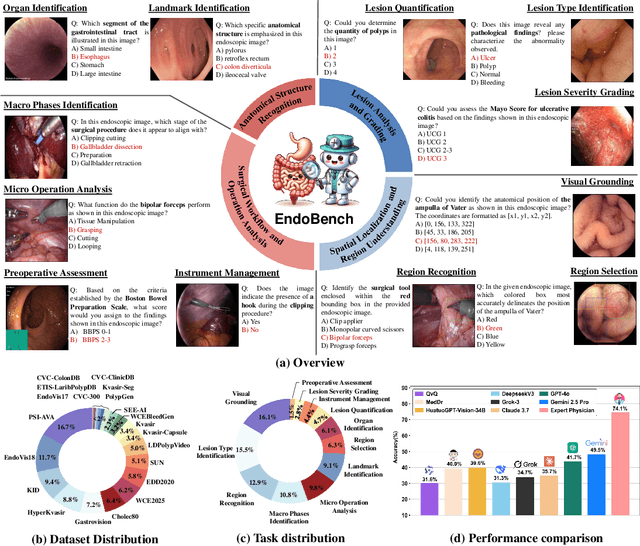


Abstract:Endoscopic procedures are essential for diagnosing and treating internal diseases, and multi-modal large language models (MLLMs) are increasingly applied to assist in endoscopy analysis. However, current benchmarks are limited, as they typically cover specific endoscopic scenarios and a small set of clinical tasks, failing to capture the real-world diversity of endoscopic scenarios and the full range of skills needed in clinical workflows. To address these issues, we introduce EndoBench, the first comprehensive benchmark specifically designed to assess MLLMs across the full spectrum of endoscopic practice with multi-dimensional capacities. EndoBench encompasses 4 distinct endoscopic scenarios, 12 specialized clinical tasks with 12 secondary subtasks, and 5 levels of visual prompting granularities, resulting in 6,832 rigorously validated VQA pairs from 21 diverse datasets. Our multi-dimensional evaluation framework mirrors the clinical workflow--spanning anatomical recognition, lesion analysis, spatial localization, and surgical operations--to holistically gauge the perceptual and diagnostic abilities of MLLMs in realistic scenarios. We benchmark 23 state-of-the-art models, including general-purpose, medical-specialized, and proprietary MLLMs, and establish human clinician performance as a reference standard. Our extensive experiments reveal: (1) proprietary MLLMs outperform open-source and medical-specialized models overall, but still trail human experts; (2) medical-domain supervised fine-tuning substantially boosts task-specific accuracy; and (3) model performance remains sensitive to prompt format and clinical task complexity. EndoBench establishes a new standard for evaluating and advancing MLLMs in endoscopy, highlighting both progress and persistent gaps between current models and expert clinical reasoning. We publicly release our benchmark and code.
FaceBench: A Multi-View Multi-Level Facial Attribute VQA Dataset for Benchmarking Face Perception MLLMs
Mar 27, 2025



Abstract:Multimodal large language models (MLLMs) have demonstrated remarkable capabilities in various tasks. However, effectively evaluating these MLLMs on face perception remains largely unexplored. To address this gap, we introduce FaceBench, a dataset featuring hierarchical multi-view and multi-level attributes specifically designed to assess the comprehensive face perception abilities of MLLMs. Initially, we construct a hierarchical facial attribute structure, which encompasses five views with up to three levels of attributes, totaling over 210 attributes and 700 attribute values. Based on the structure, the proposed FaceBench consists of 49,919 visual question-answering (VQA) pairs for evaluation and 23,841 pairs for fine-tuning. Moreover, we further develop a robust face perception MLLM baseline, Face-LLaVA, by training with our proposed face VQA data. Extensive experiments on various mainstream MLLMs and Face-LLaVA are conducted to test their face perception ability, with results also compared against human performance. The results reveal that, the existing MLLMs are far from satisfactory in understanding the fine-grained facial attributes, while our Face-LLaVA significantly outperforms existing open-source models with a small amount of training data and is comparable to commercial ones like GPT-4o and Gemini. The dataset will be released at https://github.com/CVI-SZU/FaceBench.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge