Sekeun Kim
CAMCA, Massachusetts General Hospital and Harvard Medical School
Surgical Agent Orchestration Platform for Voice-directed Patient Data Interaction
Nov 11, 2025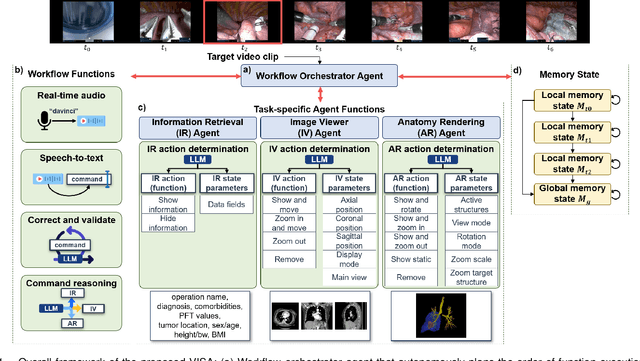
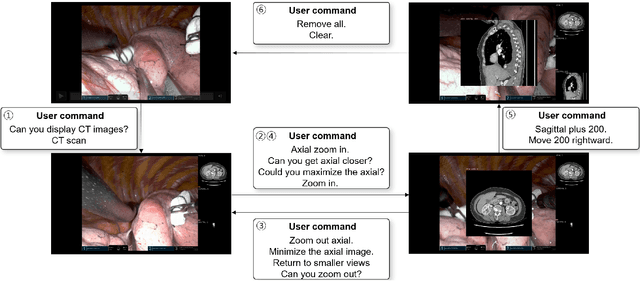
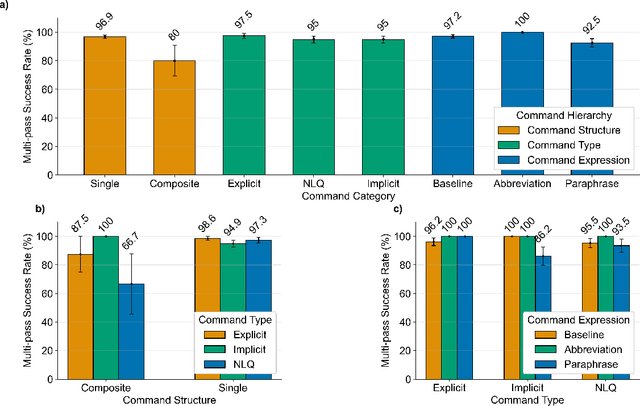
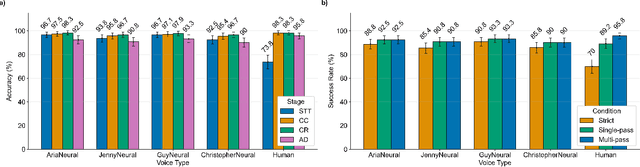

Abstract:In da Vinci robotic surgery, surgeons' hands and eyes are fully engaged in the procedure, making it difficult to access and manipulate multimodal patient data without interruption. We propose a voice-directed Surgical Agent Orchestrator Platform (SAOP) built on a hierarchical multi-agent framework, consisting of an orchestration agent and three task-specific agents driven by Large Language Models (LLMs). These LLM-based agents autonomously plan, refine, validate, and reason to map voice commands into specific tasks such as retrieving clinical information, manipulating CT scans, or navigating 3D anatomical models on the surgical video. We also introduce a Multi-level Orchestration Evaluation Metric (MOEM) to comprehensively assess the performance and robustness from command-level and category-level perspectives. The SAOP achieves high accuracy and success rates across 240 voice commands, while LLM-based agents improve robustness against speech recognition errors and diverse or ambiguous free-form commands, demonstrating strong potential to support minimally invasive da Vinci robotic surgery.
Cascaded 3D Diffusion Models for Whole-body 3D 18-F FDG PET/CT synthesis from Demographics
May 28, 2025Abstract:We propose a cascaded 3D diffusion model framework to synthesize high-fidelity 3D PET/CT volumes directly from demographic variables, addressing the growing need for realistic digital twins in oncologic imaging, virtual trials, and AI-driven data augmentation. Unlike deterministic phantoms, which rely on predefined anatomical and metabolic templates, our method employs a two-stage generative process. An initial score-based diffusion model synthesizes low-resolution PET/CT volumes from demographic variables alone, providing global anatomical structures and approximate metabolic activity. This is followed by a super-resolution residual diffusion model that refines spatial resolution. Our framework was trained on 18-F FDG PET/CT scans from the AutoPET dataset and evaluated using organ-wise volume and standardized uptake value (SUV) distributions, comparing synthetic and real data between demographic subgroups. The organ-wise comparison demonstrated strong concordance between synthetic and real images. In particular, most deviations in metabolic uptake values remained within 3-5% of the ground truth in subgroup analysis. These findings highlight the potential of cascaded 3D diffusion models to generate anatomically and metabolically accurate PET/CT images, offering a robust alternative to traditional phantoms and enabling scalable, population-informed synthetic imaging for clinical and research applications.
OWT: A Foundational Organ-Wise Tokenization Framework for Medical Imaging
May 08, 2025Abstract:Recent advances in representation learning often rely on holistic, black-box embeddings that entangle multiple semantic components, limiting interpretability and generalization. These issues are especially critical in medical imaging. To address these limitations, we propose an Organ-Wise Tokenization (OWT) framework with a Token Group-based Reconstruction (TGR) training paradigm. Unlike conventional approaches that produce holistic features, OWT explicitly disentangles an image into separable token groups, each corresponding to a distinct organ or semantic entity. Our design ensures each token group encapsulates organ-specific information, boosting interpretability, generalization, and efficiency while allowing fine-grained control in downstream tasks. Experiments on CT and MRI datasets demonstrate the effectiveness of OWT in not only achieving strong image reconstruction and segmentation performance, but also enabling novel semantic-level generation and retrieval applications that are out of reach for standard holistic embedding methods. These findings underscore the potential of OWT as a foundational framework for semantically disentangled representation learning, offering broad scalability and applicability to real-world medical imaging scenarios and beyond.
EchoFM: Foundation Model for Generalizable Echocardiogram Analysis
Oct 30, 2024



Abstract:Foundation models have recently gained significant attention because of their generalizability and adaptability across multiple tasks and data distributions. Although medical foundation models have emerged, solutions for cardiac imaging, especially echocardiography videos, are still unexplored. In this paper, we introduce EchoFM, a foundation model specifically designed to represent and analyze echocardiography videos. In EchoFM, we propose a self-supervised learning framework that captures both spatial and temporal variability patterns through a spatio-temporal consistent masking strategy and periodic-driven contrastive learning. This framework can effectively capture the spatio-temporal dynamics of echocardiography and learn the representative video features without any labels. We pre-train our model on an extensive dataset comprising over 290,000 echocardiography videos covering 26 scan views across different imaging modes, with up to 20 million frames of images. The pre-trained EchoFM can then be easily adapted and fine-tuned for a variety of downstream tasks, serving as a robust backbone model. Our evaluation was systemically designed for four downstream tasks after the echocardiography examination routine. Experiment results show that EchoFM surpasses state-of-the-art methods, including specialized echocardiography methods, self-supervised pre-training models, and general-purposed pre-trained foundation models, across all downstream tasks.
Retrieval Instead of Fine-tuning: A Retrieval-based Parameter Ensemble for Zero-shot Learning
Oct 13, 2024



Abstract:Foundation models have become a cornerstone in deep learning, with techniques like Low-Rank Adaptation (LoRA) offering efficient fine-tuning of large models. Similarly, methods such as Retrieval-Augmented Generation (RAG), which leverage vectorized databases, have further improved model performance by grounding outputs in external information. While these approaches have demonstrated notable success, they often require extensive training or labeled data, which can limit their adaptability in resource-constrained environments. To address these challenges, we introduce Retrieval-based Parameter Ensemble (RPE), a new method that creates a vectorized database of LoRAs, enabling efficient retrieval and application of model adaptations to new tasks. RPE minimizes the need for extensive training and eliminates the requirement for labeled data, making it particularly effective for zero-shot learning. Additionally, RPE is well-suited for privacy-sensitive domains like healthcare, as it modifies model parameters without accessing raw data. When applied to tasks such as medical report generation and image segmentation, RPE not only proved effective but also surpassed supervised fine-tuning methods in certain cases, highlighting its potential to enhance both computational efficiency and privacy in deep learning applications.
ECHOPulse: ECG controlled echocardio-grams video generation
Oct 04, 2024



Abstract:Echocardiography (ECHO) is essential for cardiac assessments, but its video quality and interpretation heavily relies on manual expertise, leading to inconsistent results from clinical and portable devices. ECHO video generation offers a solution by improving automated monitoring through synthetic data and generating high-quality videos from routine health data. However, existing models often face high computational costs, slow inference, and rely on complex conditional prompts that require experts' annotations. To address these challenges, we propose ECHOPULSE, an ECG-conditioned ECHO video generation model. ECHOPULSE introduces two key advancements: (1) it accelerates ECHO video generation by leveraging VQ-VAE tokenization and masked visual token modeling for fast decoding, and (2) it conditions on readily accessible ECG signals, which are highly coherent with ECHO videos, bypassing complex conditional prompts. To the best of our knowledge, this is the first work to use time-series prompts like ECG signals for ECHO video generation. ECHOPULSE not only enables controllable synthetic ECHO data generation but also provides updated cardiac function information for disease monitoring and prediction beyond ECG alone. Evaluations on three public and private datasets demonstrate state-of-the-art performance in ECHO video generation across both qualitative and quantitative measures. Additionally, ECHOPULSE can be easily generalized to other modality generation tasks, such as cardiac MRI, fMRI, and 3D CT generation. Demo can seen from \url{https://github.com/levyisthebest/ECHOPulse_Prelease}.
Autonomous Robotic Ultrasound System for Liver Follow-up Diagnosis: Pilot Phantom Study
May 09, 2024



Abstract:The paper introduces a novel autonomous robot ultrasound (US) system targeting liver follow-up scans for outpatients in local communities. Given a computed tomography (CT) image with specific target regions of interest, the proposed system carries out the autonomous follow-up scan in three steps: (i) initial robot contact to surface, (ii) coordinate mapping between CT image and robot, and (iii) target US scan. Utilizing 3D US-CT registration and deep learning-based segmentation networks, we can achieve precise imaging of 3D hepatic veins, facilitating accurate coordinate mapping between CT and the robot. This enables the automatic localization of follow-up targets within the CT image, allowing the robot to navigate precisely to the target's surface. Evaluation of the ultrasound phantom confirms the quality of the US-CT registration and shows the robot reliably locates the targets in repeated trials. The proposed framework holds the potential to significantly reduce time and costs for healthcare providers, clinicians, and follow-up patients, thereby addressing the increasing healthcare burden associated with chronic disease in local communities.
Prompt-driven Universal Model for View-Agnostic Echocardiography Analysis
Apr 09, 2024Abstract:Echocardiography segmentation for cardiac analysis is time-consuming and resource-intensive due to the variability in image quality and the necessity to process scans from various standard views. While current automated segmentation methods in echocardiography show promising performance, they are trained on specific scan views to analyze corresponding data. However, this solution has a limitation as the number of required models increases with the number of standard views. To address this, in this paper, we present a prompt-driven universal method for view-agnostic echocardiography analysis. Considering the domain shift between standard views, we first introduce a method called prompt matching, aimed at learning prompts specific to different views by matching prompts and querying input embeddings using a pre-trained vision model. Then, we utilized a pre-trained medical language model to align textual information with pixel data for accurate segmentation. Extensive experiments on three standard views showed that our approach significantly outperforms the state-of-the-art universal methods and achieves comparable or even better performances over the segmentation model trained and tested on same views.
Cardiac Magnetic Resonance 2D+T Short- and Long-axis Segmentation via Spatio-temporal SAM Adaptation
Mar 15, 2024



Abstract:Accurate 2D+T myocardium segmentation in cine cardiac magnetic resonance (CMR) scans is essential to analyze LV motion throughout the cardiac cycle comprehensively. The Segment Anything Model (SAM), known for its accurate segmentation and zero-shot generalization, has not yet been tailored for CMR 2D+T segmentation. We therefore introduce CMR2D+T-SAM, a novel approach to adapt SAM for CMR 2D+T segmentation using spatio-temporal adaption. This approach also incorporates a U-Net framework for multi-scale feature extraction, as well as text prompts for accurate segmentation on both short-axis (SAX) and long-axis (LAX) views using a single model. CMR2D+T-SAM outperforms existing deep learning methods on the STACOM2011 dataset, achieving a myocardium Dice score of 0.885 and a Hausdorff distance (HD) of 2.900 pixels. It also demonstrates superior zero-shot generalization on the ACDC dataset with a Dice score of 0.840 and a HD of 4.076 pixels.
MediViSTA-SAM: Zero-shot Medical Video Analysis with Spatio-temporal SAM Adaptation
Sep 24, 2023Abstract:In recent years, the Segmentation Anything Model (SAM) has attracted considerable attention as a foundational model well-known for its robust generalization capabilities across various downstream tasks. However, SAM does not exhibit satisfactory performance in the realm of medical image analysis. In this study, we introduce the first study on adapting SAM on video segmentation, called MediViSTA-SAM, a novel approach designed for medical video segmentation. Given video data, MediViSTA, spatio-temporal adapter captures long and short range temporal attention with cross-frame attention mechanism effectively constraining it to consider the immediately preceding video frame as a reference, while also considering spatial information effectively. Additionally, it incorporates multi-scale fusion by employing a U-shaped encoder and a modified mask decoder to handle objects of varying sizes. To evaluate our approach, extensive experiments were conducted using state-of-the-art (SOTA) methods, assessing its generalization abilities on multi-vendor in-house echocardiography datasets. The results highlight the accuracy and effectiveness of our network in medical video segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge