Le Song
MSAGPT: Neural Prompting Protein Structure Prediction via MSA Generative Pre-Training
Jun 11, 2024Abstract:Multiple Sequence Alignment (MSA) plays a pivotal role in unveiling the evolutionary trajectories of protein families. The accuracy of protein structure predictions is often compromised for protein sequences that lack sufficient homologous information to construct high quality MSA. Although various methods have been proposed to generate virtual MSA under these conditions, they fall short in comprehensively capturing the intricate coevolutionary patterns within MSA or require guidance from external oracle models. Here we introduce MSAGPT, a novel approach to prompt protein structure predictions via MSA generative pretraining in the low MSA regime. MSAGPT employs a simple yet effective 2D evolutionary positional encoding scheme to model complex evolutionary patterns. Endowed by this, its flexible 1D MSA decoding framework facilitates zero or few shot learning. Moreover, we demonstrate that leveraging the feedback from AlphaFold2 can further enhance the model capacity via Rejective Fine tuning (RFT) and Reinforcement Learning from AF2 Feedback (RLAF). Extensive experiments confirm the efficacy of MSAGPT in generating faithful virtual MSA to enhance the structure prediction accuracy. The transfer learning capabilities also highlight its great potential for facilitating other protein tasks.
Progress and Opportunities of Foundation Models in Bioinformatics
Feb 06, 2024



Abstract:Bioinformatics has witnessed a paradigm shift with the increasing integration of artificial intelligence (AI), particularly through the adoption of foundation models (FMs). These AI techniques have rapidly advanced, addressing historical challenges in bioinformatics such as the scarcity of annotated data and the presence of data noise. FMs are particularly adept at handling large-scale, unlabeled data, a common scenario in biological contexts due to the time-consuming and costly nature of experimentally determining labeled data. This characteristic has allowed FMs to excel and achieve notable results in various downstream validation tasks, demonstrating their ability to represent diverse biological entities effectively. Undoubtedly, FMs have ushered in a new era in computational biology, especially in the realm of deep learning. The primary goal of this survey is to conduct a systematic investigation and summary of FMs in bioinformatics, tracing their evolution, current research status, and the methodologies employed. Central to our focus is the application of FMs to specific biological problems, aiming to guide the research community in choosing appropriate FMs for their research needs. We delve into the specifics of the problem at hand including sequence analysis, structure prediction, function annotation, and multimodal integration, comparing the structures and advancements against traditional methods. Furthermore, the review analyses challenges and limitations faced by FMs in biology, such as data noise, model explainability, and potential biases. Finally, we outline potential development paths and strategies for FMs in future biological research, setting the stage for continued innovation and application in this rapidly evolving field. This comprehensive review serves not only as an academic resource but also as a roadmap for future explorations and applications of FMs in biology.
xTrimoPGLM: Unified 100B-Scale Pre-trained Transformer for Deciphering the Language of Protein
Jan 11, 2024Abstract:Protein language models have shown remarkable success in learning biological information from protein sequences. However, most existing models are limited by either autoencoding or autoregressive pre-training objectives, which makes them struggle to handle protein understanding and generation tasks concurrently. We propose a unified protein language model, xTrimoPGLM, to address these two types of tasks simultaneously through an innovative pre-training framework. Our key technical contribution is an exploration of the compatibility and the potential for joint optimization of the two types of objectives, which has led to a strategy for training xTrimoPGLM at an unprecedented scale of 100 billion parameters and 1 trillion training tokens. Our extensive experiments reveal that 1) xTrimoPGLM significantly outperforms other advanced baselines in 18 protein understanding benchmarks across four categories. The model also facilitates an atomic-resolution view of protein structures, leading to an advanced 3D structural prediction model that surpasses existing language model-based tools. 2) xTrimoPGLM not only can generate de novo protein sequences following the principles of natural ones, but also can perform programmable generation after supervised fine-tuning (SFT) on curated sequences. These results highlight the substantial capability and versatility of xTrimoPGLM in understanding and generating protein sequences, contributing to the evolving landscape of foundation models in protein science.
xTrimoGene: An Efficient and Scalable Representation Learner for Single-Cell RNA-Seq Data
Nov 26, 2023Abstract:Advances in high-throughput sequencing technology have led to significant progress in measuring gene expressions at the single-cell level. The amount of publicly available single-cell RNA-seq (scRNA-seq) data is already surpassing 50M records for humans with each record measuring 20,000 genes. This highlights the need for unsupervised representation learning to fully ingest these data, yet classical transformer architectures are prohibitive to train on such data in terms of both computation and memory. To address this challenge, we propose a novel asymmetric encoder-decoder transformer for scRNA-seq data, called xTrimoGene$^\alpha$ (or xTrimoGene for short), which leverages the sparse characteristic of the data to scale up the pre-training. This scalable design of xTrimoGene reduces FLOPs by one to two orders of magnitude compared to classical transformers while maintaining high accuracy, enabling us to train the largest transformer models over the largest scRNA-seq dataset today. Our experiments also show that the performance of xTrimoGene improves as we scale up the model sizes, and it also leads to SOTA performance over various downstream tasks, such as cell type annotation, perturb-seq effect prediction, and drug combination prediction. xTrimoGene model is now available for use as a service via the following link: https://api.biomap.com/xTrimoGene/apply.
Drug Synergistic Combinations Predictions via Large-Scale Pre-Training and Graph Structure Learning
Jan 14, 2023



Abstract:Drug combination therapy is a well-established strategy for disease treatment with better effectiveness and less safety degradation. However, identifying novel drug combinations through wet-lab experiments is resource intensive due to the vast combinatorial search space. Recently, computational approaches, specifically deep learning models have emerged as an efficient way to discover synergistic combinations. While previous methods reported fair performance, their models usually do not take advantage of multi-modal data and they are unable to handle new drugs or cell lines. In this study, we collected data from various datasets covering various drug-related aspects. Then, we take advantage of large-scale pre-training models to generate informative representations and features for drugs, proteins, and diseases. Based on that, a message-passing graph is built on top to propagate information together with graph structure learning flexibility. This is first introduced in the biological networks and enables us to generate pseudo-relations in the graph. Our framework achieves state-of-the-art results in comparison with other deep learning-based methods on synergistic prediction benchmark datasets. We are also capable of inferencing new drug combination data in a test on an independent set released by AstraZeneca, where 10% of improvement over previous methods is observed. In addition, we're robust against unseen drugs and surpass almost 15% AU ROC compared to the second-best model. We believe our framework contributes to both the future wet-lab discovery of novel drugs and the building of promising guidance for precise combination medicine.
xTrimoABFold: De novo Antibody Structure Prediction without MSA
Dec 11, 2022



Abstract:In the field of antibody engineering, an essential task is to design a novel antibody whose paratopes bind to a specific antigen with correct epitopes. Understanding antibody structure and its paratope can facilitate a mechanistic understanding of its function. Therefore, antibody structure prediction from its sequence alone has always been a highly valuable problem for de novo antibody design. AlphaFold2, a breakthrough in the field of structural biology, provides a solution to predict protein structure based on protein sequences and computationally expensive coevolutionary multiple sequence alignments (MSAs). However, the computational efficiency and undesirable prediction accuracy of antibodies, especially on the complementarity-determining regions (CDRs) of antibodies limit their applications in the industrially high-throughput drug design. To learn an informative representation of antibodies, we employed a deep antibody language model (ALM) on curated sequences from the observed antibody space database via a transformer model. We also developed a novel model named xTrimoABFold to predict antibody structure from antibody sequence based on the pretrained ALM as well as efficient evoformers and structural modules. The model was trained end-to-end on the antibody structures in PDB by minimizing the ensemble loss of domain-specific focal loss on CDR and the frame-aligned point loss. xTrimoABFold outperforms AlphaFold2 and other protein language model based SOTAs, e.g., OmegaFold, HelixFold-Single, and IgFold with a large significant margin (30+\% improvement on RMSD) while performing 151 times faster than AlphaFold2. To the best of our knowledge, xTrimoABFold achieved state-of-the-art antibody structure prediction. Its improvement in both accuracy and efficiency makes it a valuable tool for de novo antibody design and could make further improvements in immuno-theory.
ReSel: N-ary Relation Extraction from Scientific Text and Tables by Learning to Retrieve and Select
Oct 26, 2022Abstract:We study the problem of extracting N-ary relation tuples from scientific articles. This task is challenging because the target knowledge tuples can reside in multiple parts and modalities of the document. Our proposed method ReSel decomposes this task into a two-stage procedure that first retrieves the most relevant paragraph/table and then selects the target entity from the retrieved component. For the high-level retrieval stage, ReSel designs a simple and effective feature set, which captures multi-level lexical and semantic similarities between the query and components. For the low-level selection stage, ReSel designs a cross-modal entity correlation graph along with a multi-view architecture, which models both semantic and document-structural relations between entities. Our experiments on three scientific information extraction datasets show that ReSel outperforms state-of-the-art baselines significantly.
Uncovering the Structural Fairness in Graph Contrastive Learning
Oct 06, 2022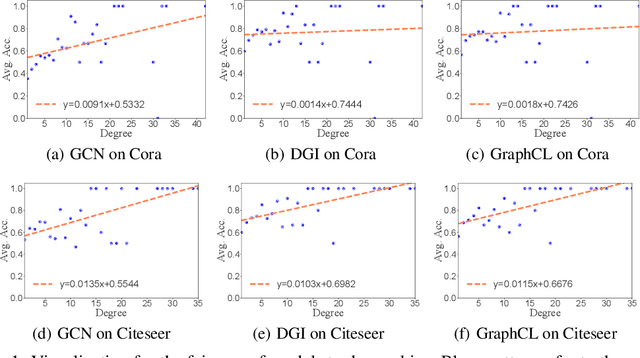
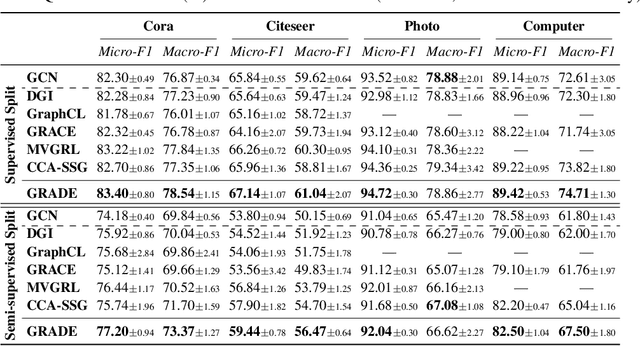
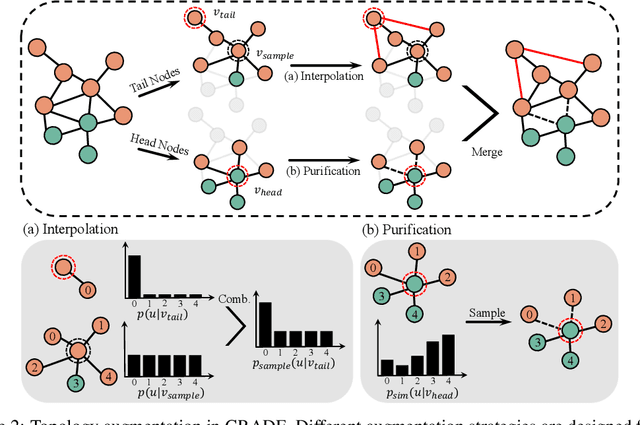
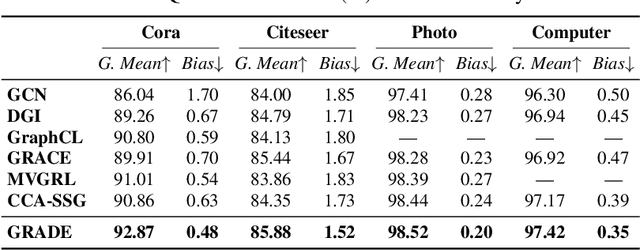
Abstract:Recent studies show that graph convolutional network (GCN) often performs worse for low-degree nodes, exhibiting the so-called structural unfairness for graphs with long-tailed degree distributions prevalent in the real world. Graph contrastive learning (GCL), which marries the power of GCN and contrastive learning, has emerged as a promising self-supervised approach for learning node representations. How does GCL behave in terms of structural fairness? Surprisingly, we find that representations obtained by GCL methods are already fairer to degree bias than those learned by GCN. We theoretically show that this fairness stems from intra-community concentration and inter-community scatter properties of GCL, resulting in a much clear community structure to drive low-degree nodes away from the community boundary. Based on our theoretical analysis, we further devise a novel graph augmentation method, called GRAph contrastive learning for DEgree bias (GRADE), which applies different strategies to low- and high-degree nodes. Extensive experiments on various benchmarks and evaluation protocols validate the effectiveness of the proposed method.
HelixFold-Single: MSA-free Protein Structure Prediction by Using Protein Language Model as an Alternative
Aug 09, 2022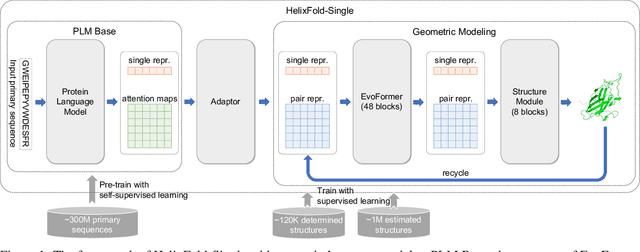

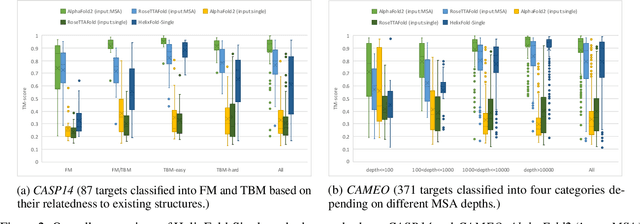
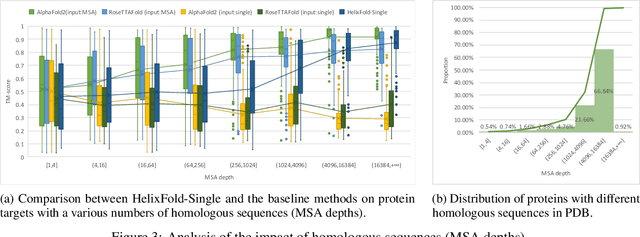
Abstract:AI-based protein structure prediction pipelines, such as AlphaFold2, have achieved near-experimental accuracy. These advanced pipelines mainly rely on Multiple Sequence Alignments (MSAs) as inputs to learn the co-evolution information from the homologous sequences. Nonetheless, searching MSAs from protein databases is time-consuming, usually taking dozens of minutes. Consequently, we attempt to explore the limits of fast protein structure prediction by using only primary sequences of proteins. HelixFold-Single is proposed to combine a large-scale protein language model with the superior geometric learning capability of AlphaFold2. Our proposed method, HelixFold-Single, first pre-trains a large-scale protein language model (PLM) with thousands of millions of primary sequences utilizing the self-supervised learning paradigm, which will be used as an alternative to MSAs for learning the co-evolution information. Then, by combining the pre-trained PLM and the essential components of AlphaFold2, we obtain an end-to-end differentiable model to predict the 3D coordinates of atoms from only the primary sequence. HelixFold-Single is validated in datasets CASP14 and CAMEO, achieving competitive accuracy with the MSA-based methods on the targets with large homologous families. Furthermore, HelixFold-Single consumes much less time than the mainstream pipelines for protein structure prediction, demonstrating its potential in tasks requiring many predictions. The code of HelixFold-Single is available at https://github.com/PaddlePaddle/PaddleHelix/tree/dev/apps/protein_folding/helixfold-single, and we also provide stable web services on https://paddlehelix.baidu.com/app/drug/protein-single/forecast.
Graph Condensation via Receptive Field Distribution Matching
Jun 28, 2022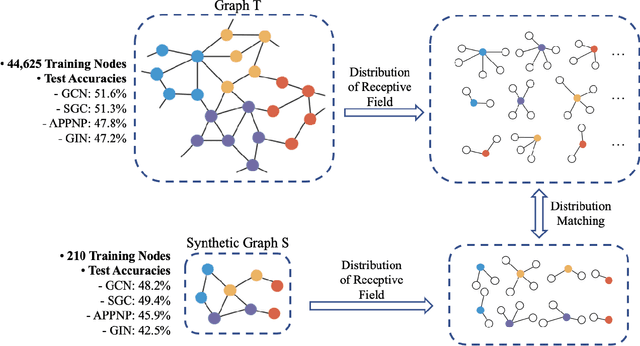

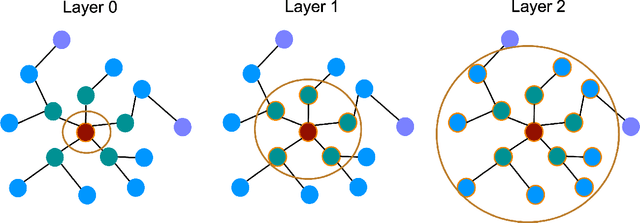

Abstract:Graph neural networks (GNNs) enable the analysis of graphs using deep learning, with promising results in capturing structured information in graphs. This paper focuses on creating a small graph to represent the original graph, so that GNNs trained on the size-reduced graph can make accurate predictions. We view the original graph as a distribution of receptive fields and aim to synthesize a small graph whose receptive fields share a similar distribution. Thus, we propose Graph Condesation via Receptive Field Distribution Matching (GCDM), which is accomplished by optimizing the synthetic graph through the use of a distribution matching loss quantified by maximum mean discrepancy (MMD). Additionally, we demonstrate that the synthetic graph generated by GCDM is highly generalizable to a variety of models in evaluation phase and that the condensing speed is significantly improved using this framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge