Guotong Xie
CandidateDrug4Cancer: An Open Molecular Graph Learning Benchmark on Drug Discovery for Cancer
Mar 02, 2022
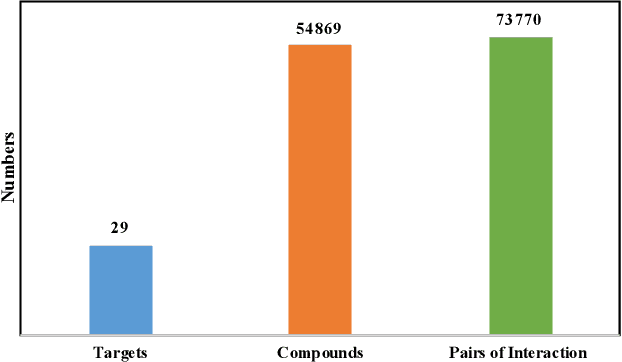
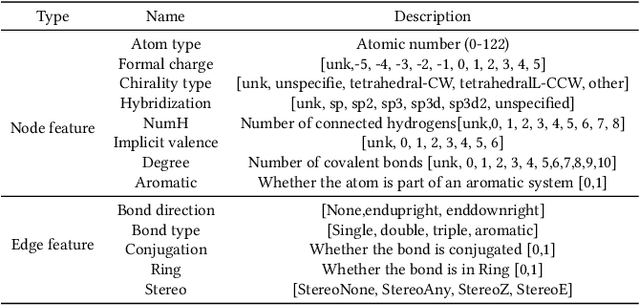
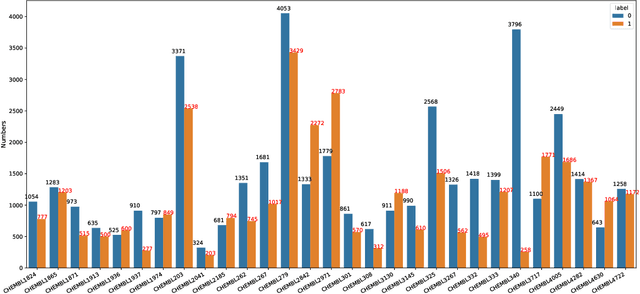
Abstract:Anti-cancer drug discoveries have been serendipitous, we sought to present the Open Molecular Graph Learning Benchmark, named CandidateDrug4Cancer, a challenging and realistic benchmark dataset to facilitate scalable, robust, and reproducible graph machine learning research for anti-cancer drug discovery. CandidateDrug4Cancer dataset encompasses multiple most-mentioned 29 targets for cancer, covering 54869 cancer-related drug molecules which are ranged from pre-clinical, clinical and FDA-approved. Besides building the datasets, we also perform benchmark experiments with effective Drug Target Interaction (DTI) prediction baselines using descriptors and expressive graph neural networks. Experimental results suggest that CandidateDrug4Cancer presents significant challenges for learning molecular graphs and targets in practical application, indicating opportunities for future researches on developing candidate drugs for treating cancers.
Superpixel-Based Building Damage Detection from Post-earthquake Very High Resolution Imagery Using Deep Neural Networks
Dec 22, 2021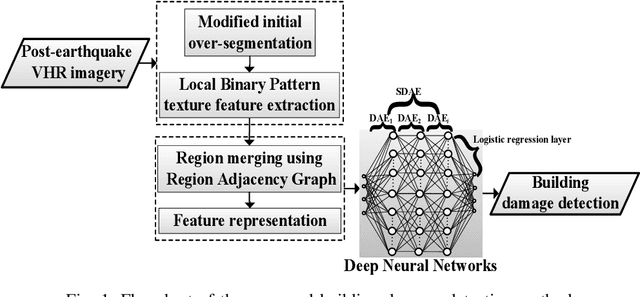
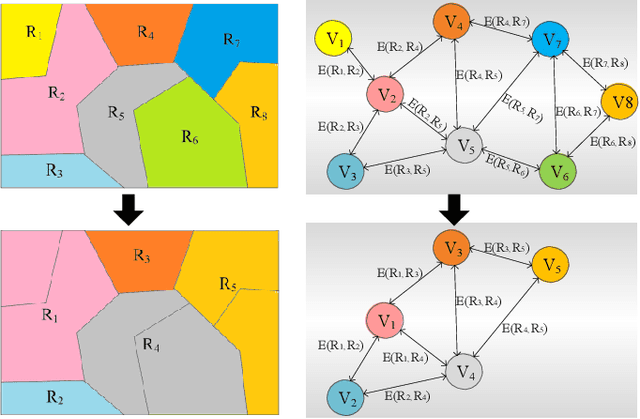
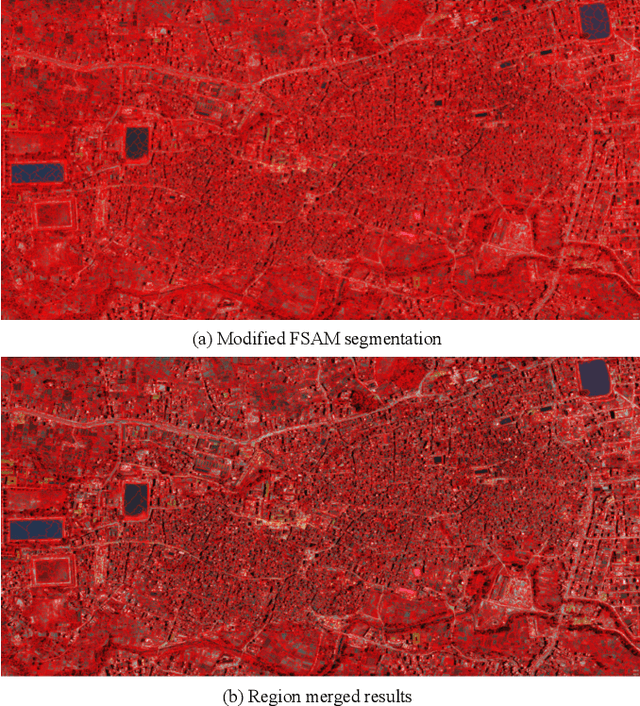
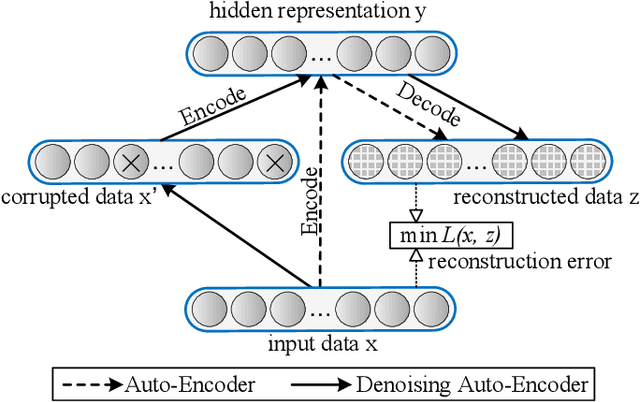
Abstract:Building damage detection after natural disasters like earthquakes is crucial for initiating effective emergency response actions. Remotely sensed very high spatial resolution (VHR) imagery can provide vital information due to their ability to map the affected buildings with high geometric precision. Many approaches have been developed to detect damaged buildings due to earthquakes. However, little attention has been paid to exploiting rich features represented in VHR images using Deep Neural Networks (DNN). This paper presents a novel superpixel based approach combining DNN and a modified segmentation method, to detect damaged buildings from VHR imagery. Firstly, a modified Fast Scanning and Adaptive Merging method is extended to create initial over-segmentation. Secondly, the segments are merged based on the Region Adjacent Graph (RAG), considered an improved semantic similarity criterion composed of Local Binary Patterns (LBP) texture, spectral, and shape features. Thirdly, a pre-trained DNN using Stacked Denoising Auto-Encoders called SDAE-DNN is presented, to exploit the rich semantic features for building damage detection. Deep-layer feature abstraction of SDAE-DNN could boost detection accuracy through learning more intrinsic and discriminative features, which outperformed other methods using state-of-the-art alternative classifiers. We demonstrate the feasibility and effectiveness of our method using a subset of WorldView-2 imagery, in the complex urban areas of Bhaktapur, Nepal, which was affected by the Nepal Earthquake of April 25, 2015.
Pairwise Half-graph Discrimination: A Simple Graph-level Self-supervised Strategy for Pre-training Graph Neural Networks
Oct 26, 2021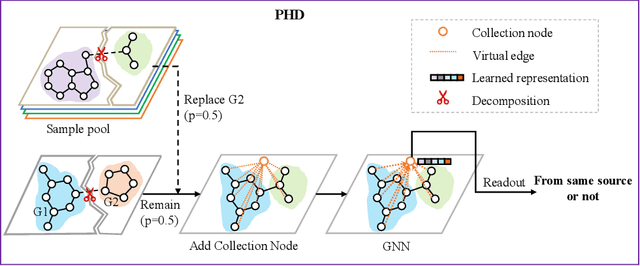
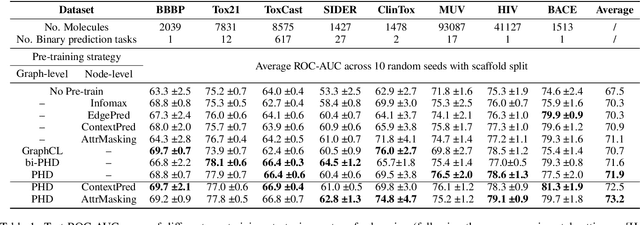
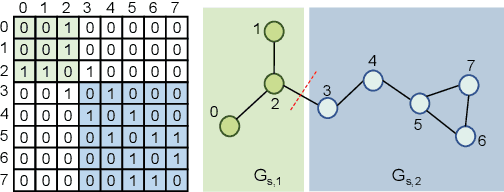
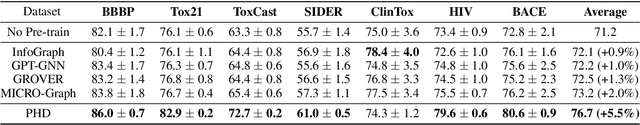
Abstract:Self-supervised learning has gradually emerged as a powerful technique for graph representation learning. However, transferable, generalizable, and robust representation learning on graph data still remains a challenge for pre-training graph neural networks. In this paper, we propose a simple and effective self-supervised pre-training strategy, named Pairwise Half-graph Discrimination (PHD), that explicitly pre-trains a graph neural network at graph-level. PHD is designed as a simple binary classification task to discriminate whether two half-graphs come from the same source. Experiments demonstrate that the PHD is an effective pre-training strategy that offers comparable or superior performance on 13 graph classification tasks compared with state-of-the-art strategies, and achieves notable improvements when combined with node-level strategies. Moreover, the visualization of learned representation revealed that PHD strategy indeed empowers the model to learn graph-level knowledge like the molecular scaffold. These results have established PHD as a powerful and effective self-supervised learning strategy in graph-level representation learning.
Multi-institutional Validation of Two-Streamed Deep Learning Method for Automated Delineation of Esophageal Gross Tumor Volume using planning-CT and FDG-PETCT
Oct 11, 2021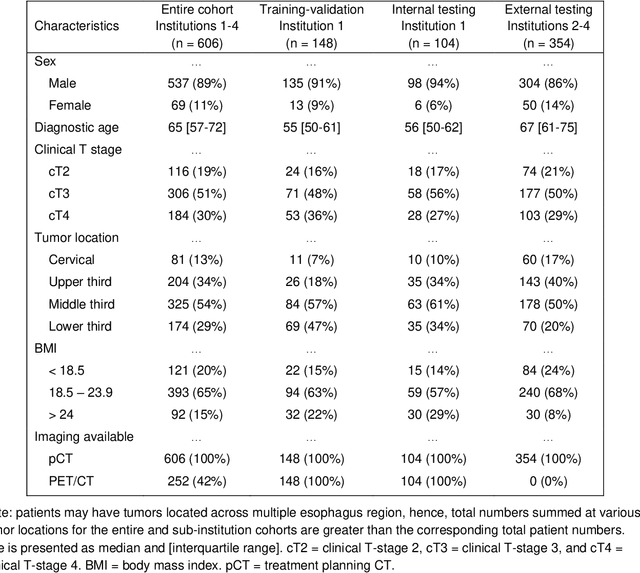
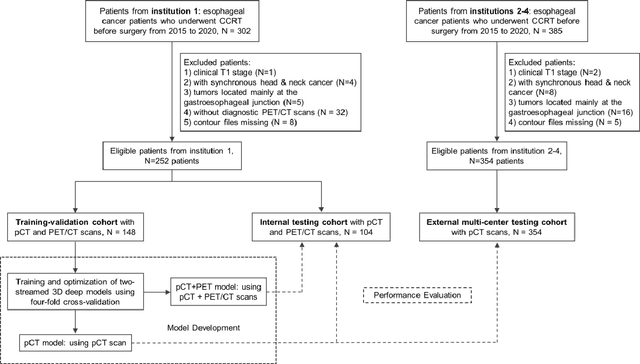
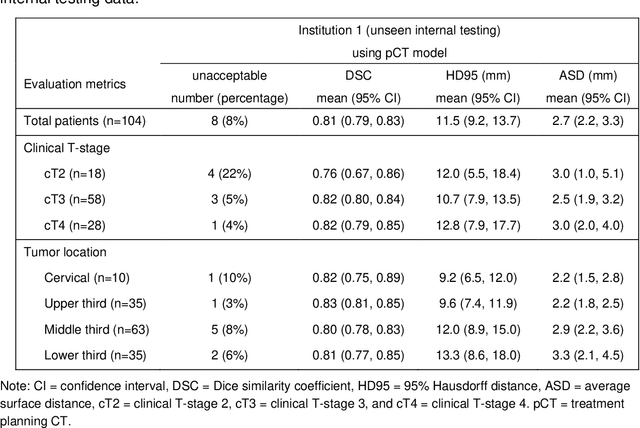
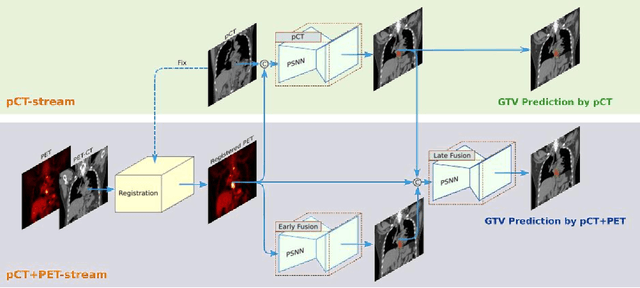
Abstract:Background: The current clinical workflow for esophageal gross tumor volume (GTV) contouring relies on manual delineation of high labor-costs and interuser variability. Purpose: To validate the clinical applicability of a deep learning (DL) multi-modality esophageal GTV contouring model, developed at 1 institution whereas tested at multiple ones. Methods and Materials: We collected 606 esophageal cancer patients from four institutions. 252 institution-1 patients had a treatment planning-CT (pCT) and a pair of diagnostic FDG-PETCT; 354 patients from other 3 institutions had only pCT. A two-streamed DL model for GTV segmentation was developed using pCT and PETCT scans of a 148 patient institution-1 subset. This built model had the flexibility of segmenting GTVs via only pCT or pCT+PETCT combined. For independent evaluation, the rest 104 institution-1 patients behaved as unseen internal testing, and 354 institutions 2-4 patients were used for external testing. We evaluated manual revision degrees by human experts to assess the contour-editing effort. The performance of the deep model was compared against 4 radiation oncologists in a multiuser study with 20 random external patients. Contouring accuracy and time were recorded for the pre-and post-DL assisted delineation process. Results: Our model achieved high segmentation accuracy in internal testing (mean Dice score: 0.81 using pCT and 0.83 using pCT+PET) and generalized well to external evaluation (mean DSC: 0.80). Expert assessment showed that the predicted contours of 88% patients need only minor or no revision. In multi-user evaluation, with the assistance of a deep model, inter-observer variation and required contouring time were reduced by 37.6% and 48.0%, respectively. Conclusions: Deep learning predicted GTV contours were in close agreement with the ground truth and could be adopted clinically with mostly minor or no changes.
SAME: Deformable Image Registration based on Self-supervised Anatomical Embeddings
Sep 23, 2021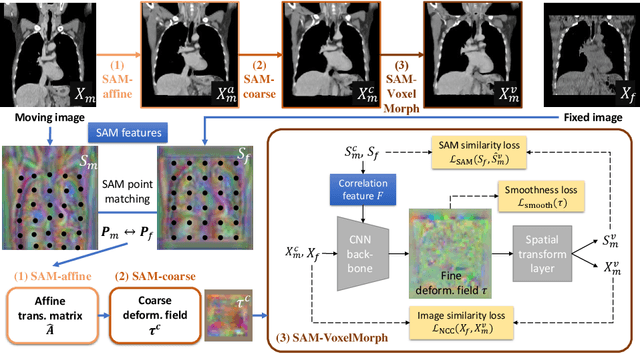
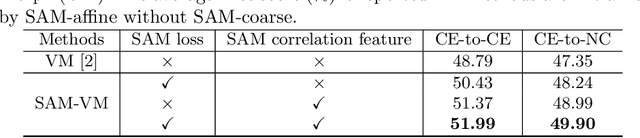
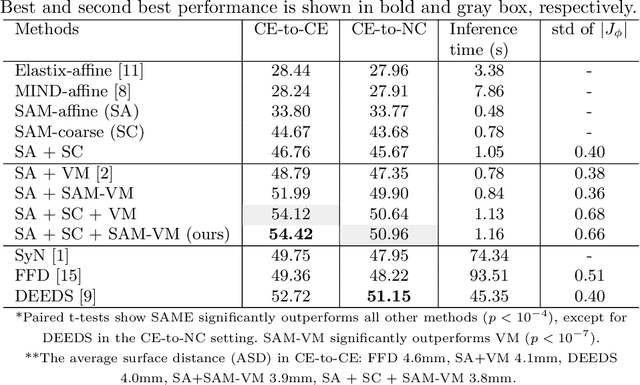

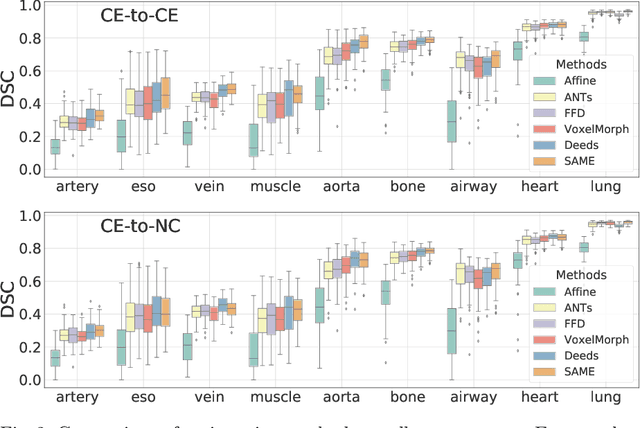
Abstract:In this work, we introduce a fast and accurate method for unsupervised 3D medical image registration. This work is built on top of a recent algorithm SAM, which is capable of computing dense anatomical/semantic correspondences between two images at the pixel level. Our method is named SAME, which breaks down image registration into three steps: affine transformation, coarse deformation, and deep deformable registration. Using SAM embeddings, we enhance these steps by finding more coherent correspondences, and providing features and a loss function with better semantic guidance. We collect a multi-phase chest computed tomography dataset with 35 annotated organs for each patient and conduct inter-subject registration for quantitative evaluation. Results show that SAME outperforms widely-used traditional registration techniques (Elastix FFD, ANTs SyN) and learning based VoxelMorph method by at least 4.7% and 2.7% in Dice scores for two separate tasks of within-contrast-phase and across-contrast-phase registration, respectively. SAME achieves the comparable performance to the best traditional registration method, DEEDS (from our evaluation), while being orders of magnitude faster (from 45 seconds to 1.2 seconds).
DeepStationing: Thoracic Lymph Node Station Parsing in CT Scans using Anatomical Context Encoding and Key Organ Auto-Search
Sep 20, 2021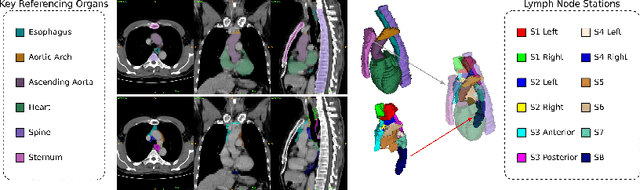
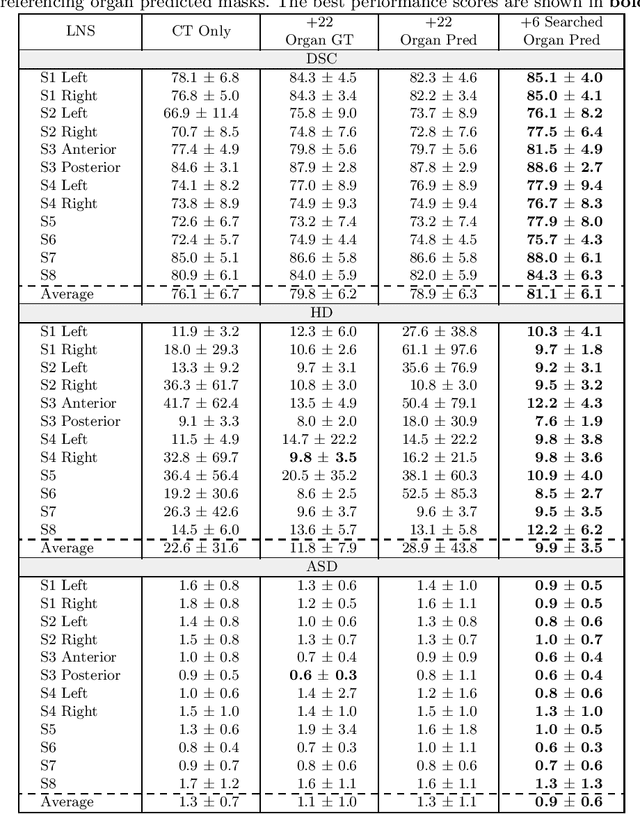
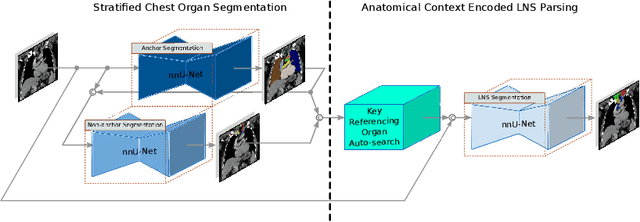
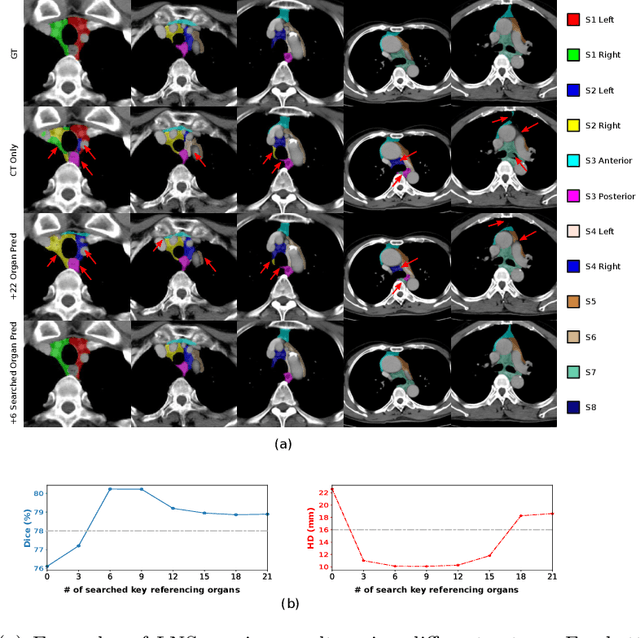
Abstract:Lymph node station (LNS) delineation from computed tomography (CT) scans is an indispensable step in radiation oncology workflow. High inter-user variabilities across oncologists and prohibitive laboring costs motivated the automated approach. Previous works exploit anatomical priors to infer LNS based on predefined ad-hoc margins. However, without voxel-level supervision, the performance is severely limited. LNS is highly context-dependent - LNS boundaries are constrained by anatomical organs - we formulate it as a deep spatial and contextual parsing problem via encoded anatomical organs. This permits the deep network to better learn from both CT appearance and organ context. We develop a stratified referencing organ segmentation protocol that divides the organs into anchor and non-anchor categories and uses the former's predictions to guide the later segmentation. We further develop an auto-search module to identify the key organs that opt for the optimal LNS parsing performance. Extensive four-fold cross-validation experiments on a dataset of 98 esophageal cancer patients (with the most comprehensive set of 12 LNSs + 22 organs in thoracic region to date) are conducted. Our LNS parsing model produces significant performance improvements, with an average Dice score of 81.1% +/- 6.1%, which is 5.0% and 19.2% higher over the pure CT-based deep model and the previous representative approach, respectively.
CBLUE: A Chinese Biomedical Language Understanding Evaluation Benchmark
Jul 06, 2021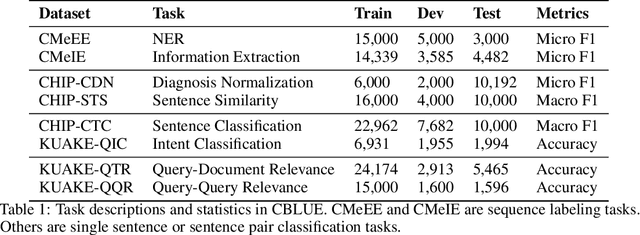
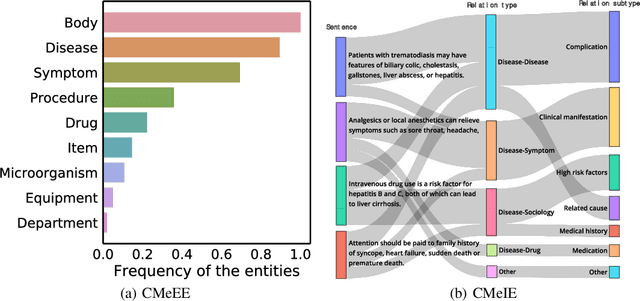
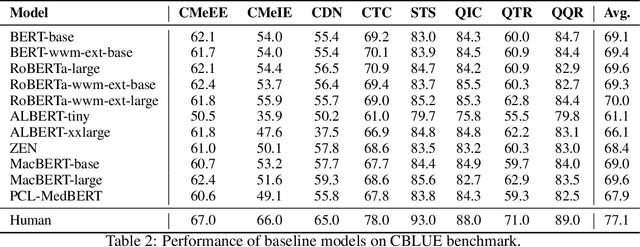
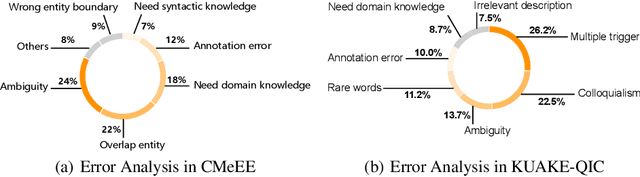
Abstract:Artificial Intelligence (AI), along with the recent progress in biomedical language understanding, is gradually changing medical practice. With the development of biomedical language understanding benchmarks, AI applications are widely used in the medical field. However, most benchmarks are limited to English, which makes it challenging to replicate many of the successes in English for other languages. To facilitate research in this direction, we collect real-world biomedical data and present the first Chinese Biomedical Language Understanding Evaluation (CBLUE) benchmark: a collection of natural language understanding tasks including named entity recognition, information extraction, clinical diagnosis normalization, single-sentence/sentence-pair classification, and an associated online platform for model evaluation, comparison, and analysis. To establish evaluation on these tasks, we report empirical results with the current 11 pre-trained Chinese models, and experimental results show that state-of-the-art neural models perform by far worse than the human ceiling. Our benchmark is released at \url{https://tianchi.aliyun.com/dataset/dataDetail?dataId=95414&lang=en-us}.
Winner Team Mia at TextVQA Challenge 2021: Vision-and-Language Representation Learning with Pre-trained Sequence-to-Sequence Model
Jun 24, 2021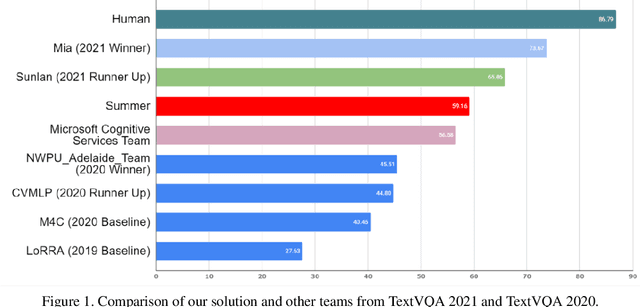
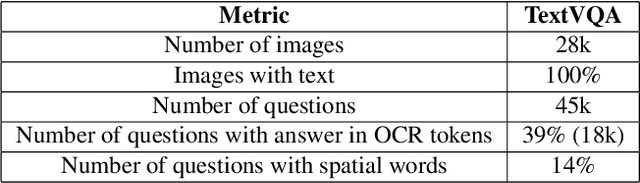
Abstract:TextVQA requires models to read and reason about text in images to answer questions about them. Specifically, models need to incorporate a new modality of text present in the images and reason over it to answer TextVQA questions. In this challenge, we use generative model T5 for TextVQA task. Based on pre-trained checkpoint T5-3B from HuggingFace repository, two other pre-training tasks including masked language modeling(MLM) and relative position prediction(RPP) are designed to better align object feature and scene text. In the stage of pre-training, encoder is dedicate to handle the fusion among multiple modalities: question text, object text labels, scene text labels, object visual features, scene visual features. After that decoder generates the text sequence step-by-step, cross entropy loss is required by default. We use a large-scale scene text dataset in pre-training and then fine-tune the T5-3B with the TextVQA dataset only.
Lesion Segmentation and RECIST Diameter Prediction via Click-driven Attention and Dual-path Connection
May 05, 2021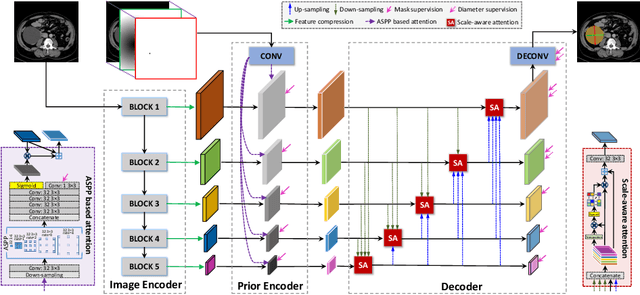
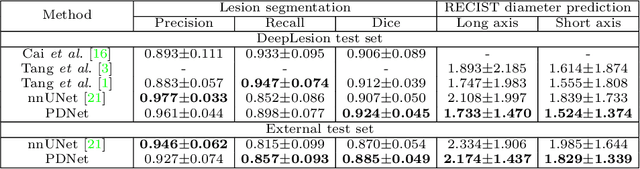
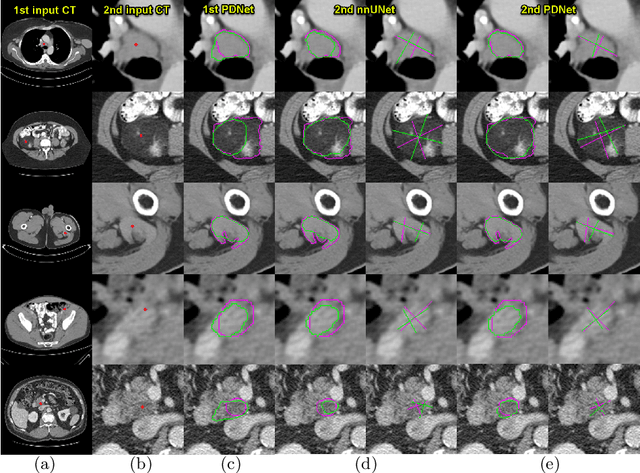
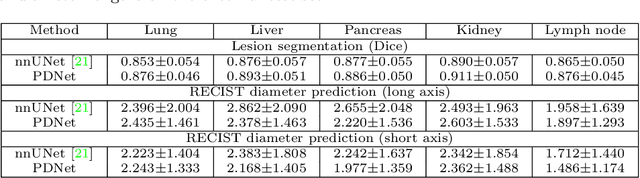
Abstract:Measuring lesion size is an important step to assess tumor growth and monitor disease progression and therapy response in oncology image analysis. Although it is tedious and highly time-consuming, radiologists have to work on this task by using RECIST criteria (Response Evaluation Criteria In Solid Tumors) routinely and manually. Even though lesion segmentation may be the more accurate and clinically more valuable means, physicians can not manually segment lesions as now since much more heavy laboring will be required. In this paper, we present a prior-guided dual-path network (PDNet) to segment common types of lesions throughout the whole body and predict their RECIST diameters accurately and automatically. Similar to [1], a click guidance from radiologists is the only requirement. There are two key characteristics in PDNet: 1) Learning lesion-specific attention matrices in parallel from the click prior information by the proposed prior encoder, named click-driven attention; 2) Aggregating the extracted multi-scale features comprehensively by introducing top-down and bottom-up connections in the proposed decoder, named dual-path connection. Experiments show the superiority of our proposed PDNet in lesion segmentation and RECIST diameter prediction using the DeepLesion dataset and an external test set. PDNet learns comprehensive and representative deep image features for our tasks and produces more accurate results on both lesion segmentation and RECIST diameter prediction.
Weakly-Supervised Universal Lesion Segmentation with Regional Level Set Loss
May 03, 2021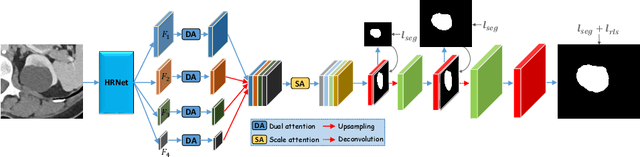
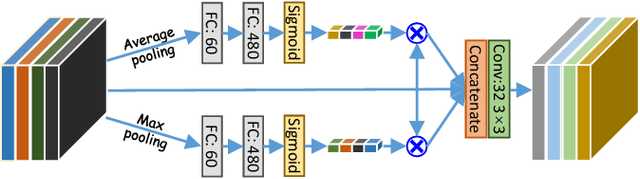
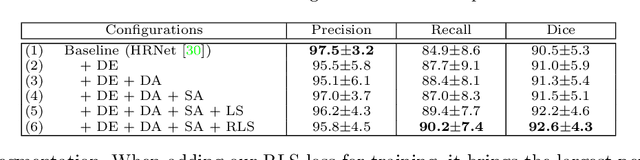
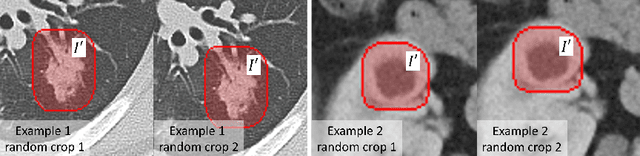
Abstract:Accurately segmenting a variety of clinically significant lesions from whole body computed tomography (CT) scans is a critical task on precision oncology imaging, denoted as universal lesion segmentation (ULS). Manual annotation is the current clinical practice, being highly time-consuming and inconsistent on tumor's longitudinal assessment. Effectively training an automatic segmentation model is desirable but relies heavily on a large number of pixel-wise labelled data. Existing weakly-supervised segmentation approaches often struggle with regions nearby the lesion boundaries. In this paper, we present a novel weakly-supervised universal lesion segmentation method by building an attention enhanced model based on the High-Resolution Network (HRNet), named AHRNet, and propose a regional level set (RLS) loss for optimizing lesion boundary delineation. AHRNet provides advanced high-resolution deep image features by involving a decoder, dual-attention and scale attention mechanisms, which are crucial to performing accurate lesion segmentation. RLS can optimize the model reliably and effectively in a weakly-supervised fashion, forcing the segmentation close to lesion boundary. Extensive experimental results demonstrate that our method achieves the best performance on the publicly large-scale DeepLesion dataset and a hold-out test set.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge