Dinggang Shen
Fellow, IEEE
DVasMesh: Deep Structured Mesh Reconstruction from Vascular Images for Dynamics Modeling of Vessels
Dec 01, 2024



Abstract:Vessel dynamics simulation is vital in studying the relationship between geometry and vascular disease progression. Reliable dynamics simulation relies on high-quality vascular meshes. Most of the existing mesh generation methods highly depend on manual annotation, which is time-consuming and laborious, usually facing challenges such as branch merging and vessel disconnection. This will hinder vessel dynamics simulation, especially for the population study. To address this issue, we propose a deep learning-based method, dubbed as DVasMesh to directly generate structured hexahedral vascular meshes from vascular images. Our contributions are threefold. First, we propose to formally formulate each vertex of the vascular graph by a four-element vector, including coordinates of the centerline point and the radius. Second, a vectorized graph template is employed to guide DVasMesh to estimate the vascular graph. Specifically, we introduce a sampling operator, which samples the extracted features of the vascular image (by a segmentation network) according to the vertices in the template graph. Third, we employ a graph convolution network (GCN) and take the sampled features as nodes to estimate the deformation between vertices of the template graph and target graph, and the deformed graph template is used to build the mesh. Taking advantage of end-to-end learning and discarding direct dependency on annotated labels, our DVasMesh demonstrates outstanding performance in generating structured vascular meshes on cardiac and cerebral vascular images. It shows great potential for clinical applications by reducing mesh generation time from 2 hours (manual) to 30 seconds (automatic).
Evaluation of OpenAI o1: Opportunities and Challenges of AGI
Sep 27, 2024



Abstract:This comprehensive study evaluates the performance of OpenAI's o1-preview large language model across a diverse array of complex reasoning tasks, spanning multiple domains, including computer science, mathematics, natural sciences, medicine, linguistics, and social sciences. Through rigorous testing, o1-preview demonstrated remarkable capabilities, often achieving human-level or superior performance in areas ranging from coding challenges to scientific reasoning and from language processing to creative problem-solving. Key findings include: -83.3% success rate in solving complex competitive programming problems, surpassing many human experts. -Superior ability in generating coherent and accurate radiology reports, outperforming other evaluated models. -100% accuracy in high school-level mathematical reasoning tasks, providing detailed step-by-step solutions. -Advanced natural language inference capabilities across general and specialized domains like medicine. -Impressive performance in chip design tasks, outperforming specialized models in areas such as EDA script generation and bug analysis. -Remarkable proficiency in anthropology and geology, demonstrating deep understanding and reasoning in these specialized fields. -Strong capabilities in quantitative investing. O1 has comprehensive financial knowledge and statistical modeling skills. -Effective performance in social media analysis, including sentiment analysis and emotion recognition. The model excelled particularly in tasks requiring intricate reasoning and knowledge integration across various fields. While some limitations were observed, including occasional errors on simpler problems and challenges with certain highly specialized concepts, the overall results indicate significant progress towards artificial general intelligence.
Towards General Text-guided Image Synthesis for Customized Multimodal Brain MRI Generation
Sep 25, 2024



Abstract:Multimodal brain magnetic resonance (MR) imaging is indispensable in neuroscience and neurology. However, due to the accessibility of MRI scanners and their lengthy acquisition time, multimodal MR images are not commonly available. Current MR image synthesis approaches are typically trained on independent datasets for specific tasks, leading to suboptimal performance when applied to novel datasets and tasks. Here, we present TUMSyn, a Text-guided Universal MR image Synthesis generalist model, which can flexibly generate brain MR images with demanded imaging metadata from routinely acquired scans guided by text prompts. To ensure TUMSyn's image synthesis precision, versatility, and generalizability, we first construct a brain MR database comprising 31,407 3D images with 7 MRI modalities from 13 centers. We then pre-train an MRI-specific text encoder using contrastive learning to effectively control MR image synthesis based on text prompts. Extensive experiments on diverse datasets and physician assessments indicate that TUMSyn can generate clinically meaningful MR images with specified imaging metadata in supervised and zero-shot scenarios. Therefore, TUMSyn can be utilized along with acquired MR scan(s) to facilitate large-scale MRI-based screening and diagnosis of brain diseases.
SDF-Net: A Hybrid Detection Network for Mediastinal Lymph Node Detection on Contrast CT Images
Sep 10, 2024



Abstract:Accurate lymph node detection and quantification are crucial for cancer diagnosis and staging on contrast-enhanced CT images, as they impact treatment planning and prognosis. However, detecting lymph nodes in the mediastinal area poses challenges due to their low contrast, irregular shapes and dispersed distribution. In this paper, we propose a Swin-Det Fusion Network (SDF-Net) to effectively detect lymph nodes. SDF-Net integrates features from both segmentation and detection to enhance the detection capability of lymph nodes with various shapes and sizes. Specifically, an auto-fusion module is designed to merge the feature maps of segmentation and detection networks at different levels. To facilitate effective learning without mask annotations, we introduce a shape-adaptive Gaussian kernel to represent lymph node in the training stage and provide more anatomical information for effective learning. Comparative results demonstrate promising performance in addressing the complex lymph node detection problem.
Prototype Learning Guided Hybrid Network for Breast Tumor Segmentation in DCE-MRI
Aug 11, 2024



Abstract:Automated breast tumor segmentation on the basis of dynamic contrast-enhancement magnetic resonance imaging (DCE-MRI) has shown great promise in clinical practice, particularly for identifying the presence of breast disease. However, accurate segmentation of breast tumor is a challenging task, often necessitating the development of complex networks. To strike an optimal trade-off between computational costs and segmentation performance, we propose a hybrid network via the combination of convolution neural network (CNN) and transformer layers. Specifically, the hybrid network consists of a encoder-decoder architecture by stacking convolution and decovolution layers. Effective 3D transformer layers are then implemented after the encoder subnetworks, to capture global dependencies between the bottleneck features. To improve the efficiency of hybrid network, two parallel encoder subnetworks are designed for the decoder and the transformer layers, respectively. To further enhance the discriminative capability of hybrid network, a prototype learning guided prediction module is proposed, where the category-specified prototypical features are calculated through on-line clustering. All learned prototypical features are finally combined with the features from decoder for tumor mask prediction. The experimental results on private and public DCE-MRI datasets demonstrate that the proposed hybrid network achieves superior performance than the state-of-the-art (SOTA) methods, while maintaining balance between segmentation accuracy and computation cost. Moreover, we demonstrate that automatically generated tumor masks can be effectively applied to identify HER2-positive subtype from HER2-negative subtype with the similar accuracy to the analysis based on manual tumor segmentation. The source code is available at https://github.com/ZhouL-lab/PLHN.
CIResDiff: A Clinically-Informed Residual Diffusion Model for Predicting Idiopathic Pulmonary Fibrosis Progression
Aug 05, 2024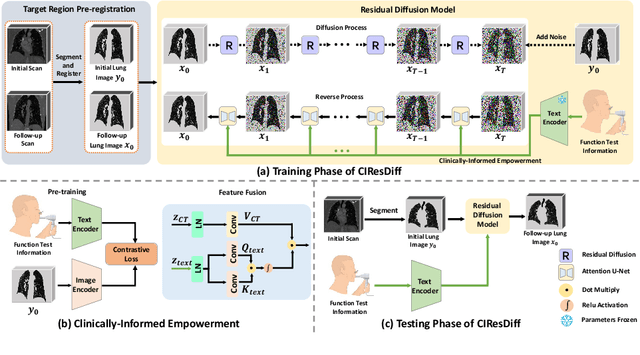
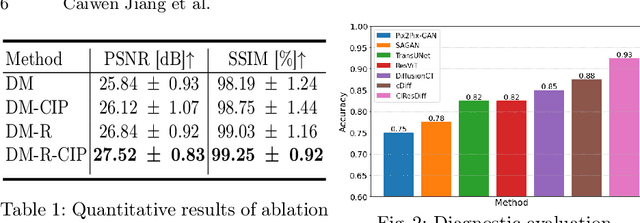
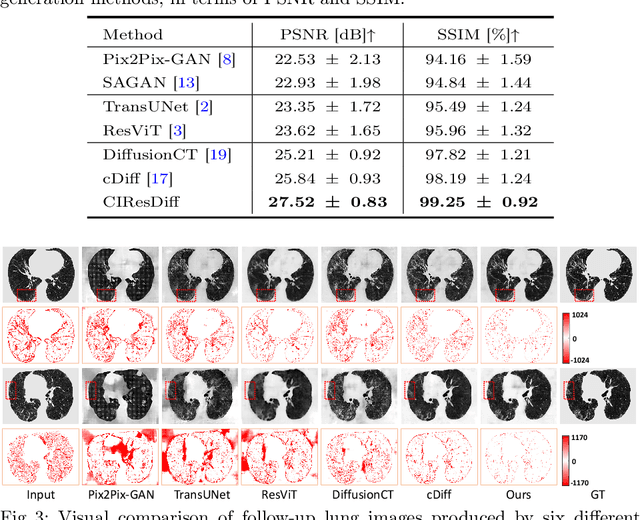
Abstract:The progression of Idiopathic Pulmonary Fibrosis (IPF) significantly correlates with higher patient mortality rates. Early detection of IPF progression is critical for initiating timely treatment, which can effectively slow down the advancement of the disease. However, the current clinical criteria define disease progression requiring two CT scans with a one-year interval, presenting a dilemma: a disease progression is identified only after the disease has already progressed. To this end, in this paper, we develop a novel diffusion model to accurately predict the progression of IPF by generating patient's follow-up CT scan from the initial CT scan. Specifically, from the clinical prior knowledge, we tailor improvements to the traditional diffusion model and propose a Clinically-Informed Residual Diffusion model, called CIResDiff. The key innovations of CIResDiff include 1) performing the target region pre-registration to align the lung regions of two CT scans at different time points for reducing the generation difficulty, 2) adopting the residual diffusion instead of traditional diffusion to enable the model focus more on differences (i.e., lesions) between the two CT scans rather than the largely identical anatomical content, and 3) designing the clinically-informed process based on CLIP technology to integrate lung function information which is highly relevant to diagnosis into the reverse process for assisting generation. Extensive experiments on clinical data demonstrate that our approach can outperform state-of-the-art methods and effectively predict the progression of IPF.
A Comprehensive Review of Multimodal Large Language Models: Performance and Challenges Across Different Tasks
Aug 02, 2024
Abstract:In an era defined by the explosive growth of data and rapid technological advancements, Multimodal Large Language Models (MLLMs) stand at the forefront of artificial intelligence (AI) systems. Designed to seamlessly integrate diverse data types-including text, images, videos, audio, and physiological sequences-MLLMs address the complexities of real-world applications far beyond the capabilities of single-modality systems. In this paper, we systematically sort out the applications of MLLM in multimodal tasks such as natural language, vision, and audio. We also provide a comparative analysis of the focus of different MLLMs in the tasks, and provide insights into the shortcomings of current MLLMs, and suggest potential directions for future research. Through these discussions, this paper hopes to provide valuable insights for the further development and application of MLLM.
A dual-task mutual learning framework for predicting post-thrombectomy cerebral hemorrhage
Aug 01, 2024Abstract:Ischemic stroke is a severe condition caused by the blockage of brain blood vessels, and can lead to the death of brain tissue due to oxygen deprivation. Thrombectomy has become a common treatment choice for ischemic stroke due to its immediate effectiveness. But, it carries the risk of postoperative cerebral hemorrhage. Clinically, multiple CT scans within 0-72 hours post-surgery are used to monitor for hemorrhage. However, this approach exposes radiation dose to patients, and may delay the detection of cerebral hemorrhage. To address this dilemma, we propose a novel prediction framework for measuring postoperative cerebral hemorrhage using only the patient's initial CT scan. Specifically, we introduce a dual-task mutual learning framework to takes the initial CT scan as input and simultaneously estimates both the follow-up CT scan and prognostic label to predict the occurrence of postoperative cerebral hemorrhage. Our proposed framework incorporates two attention mechanisms, i.e., self-attention and interactive attention. Specifically, the self-attention mechanism allows the model to focus more on high-density areas in the image, which are critical for diagnosis (i.e., potential hemorrhage areas). The interactive attention mechanism further models the dependencies between the interrelated generation and classification tasks, enabling both tasks to perform better than the case when conducted individually. Validated on clinical data, our method can generate follow-up CT scans better than state-of-the-art methods, and achieves an accuracy of 86.37% in predicting follow-up prognostic labels. Thus, our work thus contributes to the timely screening of post-thrombectomy cerebral hemorrhage, and could significantly reform the clinical process of thrombectomy and other similar operations related to stroke.
Knowledge-Guided Prompt Learning for Lifespan Brain MR Image Segmentation
Jul 31, 2024



Abstract:Automatic and accurate segmentation of brain MR images throughout the human lifespan into tissue and structure is crucial for understanding brain development and diagnosing diseases. However, challenges arise from the intricate variations in brain appearance due to rapid early brain development, aging, and disorders, compounded by the limited availability of manually-labeled datasets. In response, we present a two-step segmentation framework employing Knowledge-Guided Prompt Learning (KGPL) for brain MRI. Specifically, we first pre-train segmentation models on large-scale datasets with sub-optimal labels, followed by the incorporation of knowledge-driven embeddings learned from image-text alignment into the models. The introduction of knowledge-wise prompts captures semantic relationships between anatomical variability and biological processes, enabling models to learn structural feature embeddings across diverse age groups. Experimental findings demonstrate the superiority and robustness of our proposed method, particularly noticeable when employing Swin UNETR as the backbone. Our approach achieves average DSC values of 95.17% and 94.19% for brain tissue and structure segmentation, respectively. Our code is available at https://github.com/TL9792/KGPL.
A Progressive Single-Modality to Multi-Modality Classification Framework for Alzheimer's Disease Sub-type Diagnosis
Jul 26, 2024



Abstract:The current clinical diagnosis framework of Alzheimer's disease (AD) involves multiple modalities acquired from multiple diagnosis stages, each with distinct usage and cost. Previous AD diagnosis research has predominantly focused on how to directly fuse multiple modalities for an end-to-end one-stage diagnosis, which practically requires a high cost in data acquisition. Moreover, a significant part of these methods diagnose AD without considering clinical guideline and cannot offer accurate sub-type diagnosis. In this paper, by exploring inter-correlation among multiple modalities, we propose a novel progressive AD sub-type diagnosis framework, aiming to give diagnosis results based on easier-to-access modalities in earlier low-cost stages, instead of modalities from all stages. Specifically, first, we design 1) a text disentanglement network for better processing tabular data collected in the initial stage, and 2) a modality fusion module for fusing multi-modality features separately. Second, we align features from modalities acquired in earlier low-cost stage(s) with later high-cost stage(s) to give accurate diagnosis without actual modality acquisition in later-stage(s) for saving cost. Furthermore, we follow the clinical guideline to align features at each stage for achieving sub-type diagnosis. Third, we leverage a progressive classifier that can progressively include additional acquired modalities (if needed) for diagnosis, to achieve the balance between diagnosis cost and diagnosis performance. We evaluate our proposed framework on large diverse public and in-home datasets (8280 in total) and achieve superior performance over state-of-the-art methods. Our codes will be released after the acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge